Abstract
A novel method was developed for the screening and detection of fucoidanase activity in bacterial colonies using fucoidan–agarose plates and staining with hexadecyltrimethylammonium bromide (Cetavlon). Colonies with fucoidanase activity were indicated by a transparent halo. The medium containing undegraded fucoidan showed as a milk-colored background, but areas with degraded fucoidan under the colonies were visualized as a transparent halo.
Keywords
Bacteria; Fucoidanase; Fucoidan; Plate assay; Fucoidanase activity assay; Carbohydrate polyacrylamide gel electrophoresis
Introduction
Fucoidans are matrix polysaccharides from brown marine seaweeds, localized in the extracellular matrix. Fucoidans belong to a family of sulfated homo- and heteropolysaccharides including polysaccharides built as a highly sulfated homopolymer of fucose and polysaccharides with a high content of glucuronic acid and a low content of fucose and sulfates. The main backbone of the homopolymers consists of α-1,3-linked and/or α-1,4-linked sulfated fucose residues. These polysaccharides have attracted a great deal of attention in recent years due to the many health benefits they have demonstrated (Holtkamp et al., 2009).
Enzymes able to degrade fucoidan are important tools to establish the structural characteristics and biological functions of these polysaccharides. Fucoidanases have remained elusive and poorly understood enzymes despite the growing interest in them. Investigation into these enzymes is partly hampered by the absence of promising sources and the complexity of the methods used for the determination of fucoidanases activity.
Until now, enzymes degrading fucoidans have been found only in marine organisms such as microbes (Bakunina et al., 2002, Ivanova et al., 2003, Descamps et al., 2006 and Bakunina et al., 2000) and invertebrates (Burtseva et al., 2000 and Kusaykin et al., 2003). However, the fucoidanase activities produced by these organisms are extremely low. In fact, there is no standard fucoidanase assay. This greatly complicates the comparison of the activities and specificities of different enzymes. Fucoidanase activity can be detected by a viscometric assay specific for endodepolymerases (Kitamura et al., 1992 and Furukawa et al., 1992), by the measurement of an increase in the content of reducing end groups (Ivanova et al., 2003 and Bakunina et al., 2000), or by a carbohydrate–polyacrylamide gel electrophoresis (C–PAGE) assay based on the release of anionic oligosaccharides from sulfated fucan (Colin et al., 2006). None of these methods are expression methods and there are many difficulties in using these methods for the rapid screening of a large number of bacterial strains.
Considering these facts, this study was undertaken to develop a method for fucoidanase screening which must be cheap, easy to perform, allow simultaneous analysis of large numbers of samples and give rapid results.
Materials and Methods
Organisms and Cultivation
The bacteria used were Formosa algae KMM 3553T, Formosa sp. MF 2-3, Coheasibacter sp. SF 2-8, Cytophaga sp. ZBS 33F, and unidentified strain MF 4-5 from the Collection of Marine Organisms (KMM) of G.B. Elyakov Pacific Institute of Bioorganic Chemistry. These strains were cultivated on marine agar 2216 (BD). Strains Coheasibacter sp. SF 2-8, and Cytophaga sp. ZBS 33F were negative for fucoidan degradation.
Carbohydrate–Polyacrylamide Gel Electrophoresis (C–PAGE)
The activity of the fucoidanase was monitored by C–PAGE. The fucoidanase activity was detected by the occurrence of charged oligosaccharide bands in the gel. For investigation of extracellular enzymes a piece of agar gel was cut out from the halo area and homogenized with 50 μl 0.5 M Tris–HCl buffer, pH 6.8 and 10 μl of the loading buffer.
The samples were electrophoresed through a 5% (w/v) stacking gel with Tris–HCl buffer pH 6.8 and a 27% (w/v) resolving polyacrylamide gel with Tris–HCl buffer pH 8.8. The gel was 1 mm thick. Gel staining was performed with 0.01% O-toluidine blue in a mixture with EtOH, AcOH and H2O, at a volume ratio of 2:1:1.
Results and Discussion
The activity of enzymes hydrolyzing polysaccharides is usually measured using the release of mono-and oligosaccharides produced during hydrolysis by a conventional reducing sugar assay (Ivanova et al., 2003 and Bakunina et al., 2000) or by changing the physical properties of the substrate, such as the viscosity (Kitamura et al., 1992 and Furukawa et al., 1992).
The most common method for monitoring the activity of polysaccharide hydrolases, by increasing the amount of reducing sugars (Ivanova et al., 2003 and Bakunina et al., 2000), was almost unusable for fucoidanases. Even in the case of fucoidan, which was efficiently cleaved as clearly shown by electrophoresis, the reducing ability of the solution measured by the Nelson method was close to zero. It seems that this is associated with the features of the chemical structure of the hydrolysis products of fucoidan.
A method for the identification of enzymatic hydrolysis products of fucoidan using electrophoresis in PAAG is most representative, although it is expensive, time-consuming and not very suitable for screening a large number of samples.
Here, we propose a simple method for the determination of fucoidanase activity and fucoidan utilization by bacteria growing on agar medium. The method is based on the ability of the cationic detergent hexadecyltrimethylammonium bromide (Cetavlon) to form a water-insoluble salt with a white color in the presence of fucoidan. The fucoidan that is enzymatically degraded has a low charge density and is not precipitated with Cetavlon.
To demonstrate our method, we selected strains producing fucoidanases (Formosa algae KMM 3553T and Formosa sp. MF 2-3) and strains not producing fucoidanases (Coheasibacter sp. SF 2-8 and Cytophaga sp. ZBS 33F, negative control). The strains were cultivated for 3 days at 28 °C on marine agar 2216 (BD) containing 0.5% fucoidan from Fucus evanescens purified as previously described ( Silchenko et al., 2013). Bacterial cells were removed from the agar surface and a 1% solution of Cetavlon in distilled water was added to the Petri dishes. After incubation for 30 min at 25 °C, the dishes were washed with water several times. Transparent areas appeared under colonies possessing fucoidanase activity (Fig. 1).
|
|
|
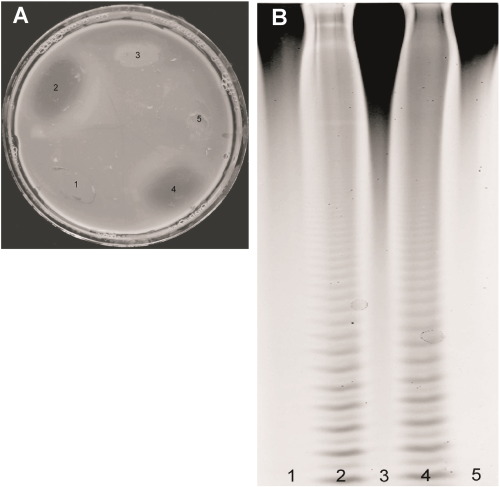
Fig. 1. Solid medium with 0.5% fucoidans treated with a 1% aqueous solution of Cetavlon (A) and an electropherogram of the same samples of bacteria after incubation with fucoidans (B). 1 – strain MF 4-5, 5 – Coheasibacter sp. SF 2-8 and 3 – Cytophaga sp. ZBS 33F (strains not producing fucoidanase); 2 – Formosa algae KMM 3553T; 4 – Formosa sp. MF 2-3 (fucoidanase-producing strains).
|
It should be noted that the concentration of fucoidan in the medium (0.5%) was optimal. Fucoidan at lower concentrations (from 0.1% to 0.4%) give a poor precipitate with Cetavlon. Staining plates with dyes specific to the anionic groups of polymers was unsuccessful because native and degraded fucoidan stained with the same intensity.
To confirm our results, we used carbohydrate polyacrylamide gel electrophoresis. The samples, after enzymatic hydrolysis of fucoidans by selected strains producing fucoidanases (Formosa algae KMM 3553T and Formosa sp. MF 2-3) and strains not producing fucoidanases (Coheasibacter sp. SF 2-8 and Cytophaga sp. ZBS 33F, negative control), were assessed by C-PAAG. Samples 2 and 4 contained fucoidanases ( Fig. 1 A) that were able to cleave the polysaccharide to low molecular weight fucooligosaccharides (Fig. 1 B, lines 2, and 4). It can be seen (Fig. 1) that there was a complete coincidence between the results obtained by the express method in Petri dishes and by C-PAAG.
In conclusion, the method has significant advantages over previously used methods and provides an assessment of fucoidanase activity simultaneously in a large numbers of bacterial strains.
Acknowledgments
This work was supported by the Russian Foundation for Basic Research (RFBR), project nos. 15-04-01004 and 16-54-540004, and an international collaboration project between Vietnam Academy of Science and Technology and Far-Eastern Branch, the Russian Academy of Science “Biocatalysis for brown algae polysaccharides modification as a basis for obtaining and studying fragments responsible for their biological activity” (VAST.HTQT.NGA.10/14-15, VAST.HTQT.NGA. 06/16-17).
References
- Bakunina et al., 2000 I. Bakunina, L.S. Shevchenko, O.I. Nedashkovskaia, N.M. Shevchenko, S.A. Alekseeva, V.V. Mikhailov, T.N. Zviagintseva; Screening of marine bacteria for fucoidan hydrolases; Mikrobiologiia, 69 (2000), pp. 370–376
- Bakunina et al., 2002 I. Bakunina, O.I. Nedashkovskaia, S.A. Alekseeva, E.P. Ivanova, L.A. Romanenko, N.M. Gorshkova, V.V. Isakov, T.N. Zviagintseva, V.V. Mikhailov; Degradation of fucoidan by the marine proteobacterium Pseudoalteromonas citrea; Mikrobiologiia, 71 (2002), pp. 49–55
- Burtseva et al., 2000 Y.V. Burtseva, M.I. Kusajkin, V.V. Sova, N.M. Shevchenko, A.S. Skobun, T.N. Zvyagintseva; The distribution of fucoidanases and some glycosidases in marine invertebrates; Biologia Morya., 26 (2000), pp. 429–432
- Colin et al., 2006 S. Colin, E. Deniaud, M. Jam, V. Descamps, Y. Chevolot, N. Kervarec, J.C. Yvin, T. Barbeyron, G. Michel, B. Kloareg; Cloning and biochemical characterization of the fucanase FcnA: definition of a novel glycoside hydrolase family specific for sulfated fucans; Glycobiology, 16 (2006), pp. 1021–1032
- Descamps et al., 2006 V. Descamps, S. Colin, M. Lahaye, M. Jam, C. Richard, P. Potin, T. Barbeyron, J.C. Yvin, B. Kloareg; Isolation and culture of a marine bacterium degrading the sulfated fucans from marine brown algae; Mar. Biotechnol. (NY)., 8 (2006), pp. 27–39
- Furukawa et al., 1992 S. Furukawa, T. Fujikawa, D. Koga, A. Ide; Purification and Some Properties of Exo-Type Fucoidanases from Vibrio sp. N-5; Biosci. Biotechnol. Biochem., 56 (1992), pp. 1829–1834
- Holtkamp et al., 2009 A.D. Holtkamp, S. Kelly, R. Ulber, S. Lang; Fucoidans and fucoidanases—focus on techniques for molecular structure elucidation and modification of marine polysaccharides; Appl. Microbiol. Biotechnol., 82 (2009), pp. 1–11
- Ivanova et al., 2003 E.P. Ivanova, I.Y. Bakunina, O.I. Nedashkovskaya, N.M. Gorshkova, Y.V. Alexeeva, E.A. Zelepuga, T.N. Zvaygintseva, D.V. Nicolau, V.V. Mikhailov; Ecophysiological variabilities in ectohydrolytic enzyme activities of some Pseudoalteromonas species, P. citrea, P. issachenkonii, and P. nigrifaciens; Curr. Microbiol., 46 (2003), pp. 6–10
- Kitamura et al., 1992 K. Kitamura, M. Matsuo, T. Yasui; Enzymatic degradation of fucoidan by fucoidanase from the hepatopancreas of Patinopecten yessoensis; Biosci. Biotechnol. Biochem., 56 (1992), pp. 490–494
- Kusaykin et al., 2003 M.I. Kusaykin, Y.V. Burtseva, T.G. Svetasheva, V.V. Sova, T.N. Zvyagintseva; Distribution of O-glycosylhydrolases in marine invertebrates. Enzymes of the marine mollusk Littorina kurila that catalyze fucoidan transformation; Biochemistry (Mosc), 68 (2003), pp. 317–324
- Silchenko et al., 2013 A.S. Silchenko, M.I. Kusaykin, V.V. Kurilenko, A.M. Zakharenko, V.V. Isakov, T.S. Zaporozhets, A.K. Gazha, T.N. Zvyagintseva; Hydrolysis of fucoidan by fucoidanase; (2013)
Document information
Published on 31/08/16
Licence: Other
Share this document
Keywords
claim authorship
Are you one of the authors of this document?