Inherited arrhythmic disorders are linked with cardiac channelopathies [1]. Inherited long QT syndrome (LQTS) is associated with an increased risk for ventricular arrhythmia and sudden cardiac death (SCD) in a structurally normal heart. The estimated incidence for LQTS is 1:5000 [2].
In contemporary practice, genetic testing not only helps find a culprit mutation in approximately 75% of individuals with a clinical diagnosis of LQTS, but also has a significant role in stratifying the underlying risk for sudden cardiac death SCD [2]. However, approximately 20% of LQTS cases remain genetically elusive [3]. The most common LQTS channelopathy mutations were found in potassium (K) channels in LQT1 and LQT2 and sodium channelopathies in LQT3 (SCN5A) and represent the majority of LQT phenotypes (i.e. around 75%), but some of them are due to affected calcium channels [4].
Pathogenic mutations in CACNA1C and KCNE1 have been reported in association with LQTS. LQT8 is caused by the mutations in CACNA1C gene, which encodes an alpha-1 subunit of a voltage-dependent calcium channel (L-type cardiac calcium channel) [5]. CACNA1C mutations can mediate calcium ions' influx into the cell characterized by an increase in calcium sparkles, likely resulting in QT prolongation [3] ; [6]. Pathogenic mutations in CACNA1C have been reported to also manifest as Timothy syndrome and Brugada syndrome type 3 [3].
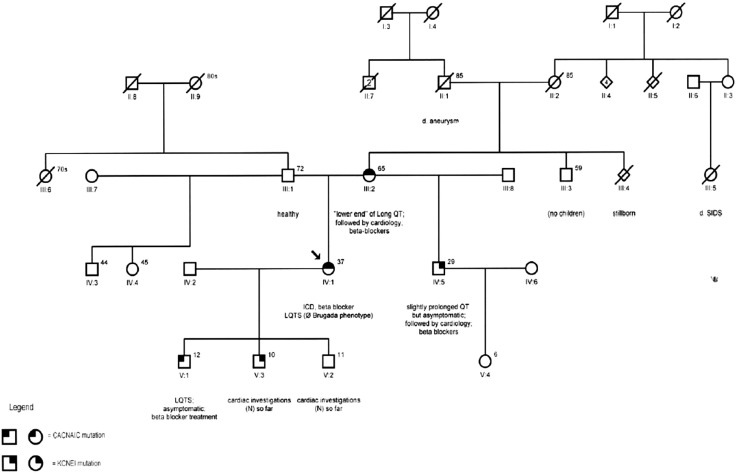
We report here a Caucasian family presented with aborted SCD in a family member, this member was subsequently found to have novel variants in two genes CACNA1C and KCNE1. A 38-year-old woman (proband, case IV.1) presented with aborted VF arrest ( Fig. 1). At presentation, the patients electrocardiograms revealed a long QT interval of 627 ms. She underwent risk assessment investigations including coronary angiogram, echocardiography and genetic testing. Eventually she underwent insertion of an implantable cardioverter defibrillator for secondary prevention purpose. A genetic testing panel including sequencing and deletion/duplication analysis of 12 genes associated with LQTS was performed which revealed two variants of unknown significance – one in the CACNA1C gene and the other in the KCNE1 gene. We obtained a targeted 4-generation pedigree ( Fig. 1), which revealed that her asymptomatic mother (case III.2) have a slightly lengthened QT interval (457 ms), and a maternal first cousin-who died of Sudden Infant Death Syndrome (III.4). There was also a remote report of SCD in her maternal grandmothers sister before the age of 40 (II.5).
|
|
|
Fig. 1. Family pedigree showing carriers of CACNA1C gene variant (left upper quarter shadow) and KCNE1 (right upper quadrant shadow). |
In order to further characterize the 2 genetic variants, familial gene testing was conducted in conjunction with a general clinical cardiac evaluation. Case (III.2, age 65) was found to have both the CACNA1C and KCNE1 variants. One of the 3 sons of the proband (case V.1, age 12) was asymptomatic and was found to have the CACNA1C variant alone but treated by B-blocker. The probands second son (case V.2, age 11) was found to carry neither of these variants with underlying normal ECG. The third sibling (asymptomatic, V.3, age 10) was found to have the KCNE1 variant and was also asymptomatic.
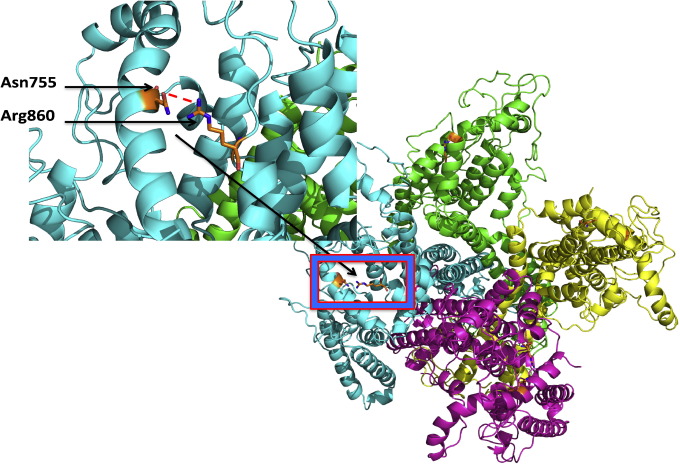
Genomic DNA was isolated from peripheral blood leukocytes using standard protocols. To investigate the isolated R860Q mutation of CACNA1C gene, a model of the CACNA1C monomer (residues 741 – 1262) was built using the Phyre2 protein fold recognition server [7]. A tetramer was built using the SSM superpose feature in Coot “molecular-graphics application for model building and validation of biological molecules” to align the CACNA1C model to the 4 molecules of the biological assembly of the NavAb voltage-gated sodium channel (PDB ID: 3RVZ). Residue R860 is not predicted to be involved with the tetramer formation. It is, however, located near residue N755 that it could form a hydrogen-bonding interaction, stabilizing the structure of the domain. A R860Q mutation results in a short side chain and would break this critical interaction, creating instability in the domain structure (Fig. 2).
|
|
|
Fig. 2. This crystallographic model of the CACNA1C monomer (residues 741 – 1262) was built using the Phyre2 protein fold recognition server. This depicts R860 Q variant mutation where residue R860 is located near residue N755 that is supposed to form a hydrogen-bonding interaction, stabilizing the structure of the domain. |
Our major findings included in the proband revealed perturbed CACNA1C gene is most likely associated with aborted SCD with prominent QT prolongation at presentation. This variant presented with positive epinephrine provocation test and EST in another two family members. There was no history of Timothy syndrome or BrS in the family. Functional analysis was reported as “likely disease causing”. This is to document that R860Q semi-conservative amino acid substitution in CACNA1C gene is clinically associated with high risk for SCD. While the proband has received intracardiac defibrillator (ICD) the other two members have started on B-blocker only being asymptomatic. In addition to CACNA1C gene mutation, a novel variant of unknown clinical significance (S105L) in KCNE1 gene is also found in the proband and other two family members. The lab reported the variant to be ‘likely benign’. However, the proband has both gene mutations and presented with aborted SCD.
In this report, we found a novel gene variant mutation (R860Q mutation of CACNA1C gene) in the CACNA1C gene represents a mutation of the traditional LQT8 subtype and is limited to a single pedigree. This variant is expected to be a disease-causing mutation with abnormally prolonged QTc durations [8] and predispose to SCD. To the best of our knowledge, the reported mutation R860Q has not been previously recognized as a disease causing mutation or as a benign polymorphism. The R860Q variant is a semi-conservative amino acid substitution and this residue shares the same properties, but it differs in size, charge, and other properties that may impact secondary structures. In silico analysis the mutation R860Q lead to damage to the protein structure/function hence current augmentation and disease expression. This is supported by an evidence in mutations in nearby residues (P857R, R858H) that have been reported in “human gene mutation database” HGMD in association with LQTS [9]. Hence the functional importance of this region of the protein is appreciated.
Most recent reports suggest that the voltage-sensing domain (VSD) movement may be altered by a KCNE1 gene mutation. It is dynamic and has a benign nature on its own. However, this would modulate the gating of the KCNQ1 channel and subsequently lead to LQTS [10]. KCNE1 (Voltage-gated potassium channel) and KCNE1 variant mutation (S105L) gene variant was found in our cohort is in TCG > TTG, (at C.314) C > T in Exon 3 of KCNE1. This is a translocation of Cytosin by Thiamine in location 314 on Exon 3 of KCNE1 gene in chromosome 21. In silico analysis of S105L heterozygous variant is reported unlikely to be a disease-causing variant.
In conclusion we report a novel variant (R860Q) in the CACNA1C gene (LQT8) associated with LQTS and aborted SCD. The crystallographic model depicts R860Q variant mutation where residue R860 is located near residue N755 and helps in forming a hydrogen-bond interaction, thus over stabilizing the structure of the domain causing voltage augmentation, thus causing symptoms. We anticipate this report to help other healthcare providers to guide their practice in managing patients with this variant in CACNA1C gene. The KCNE1 variant, though reported as a “variant of uncertain significance”, may be associated with late-onset predisposition for long QT and needs further clarification.
No conflict of interest to declare.
References
- [1] R.J. Myerburg; Physiological variations, environmental factors, and genetic modifications in inherited LQT syndromes; J. Am. Coll. Cardiol., 65 (4) (2015), pp. 375–377
- [2] I. Goldenberg, A.J. Moss; Long QT syndrome; J. Am. Coll. Cardiol., 51 (24) (2008), pp. 2291–2300
- [3] N.J. Boczek, J.M. Best, D.J. Tester, J.R. Giudicessi, S. Middha, J.M. Evans, et al.; Exome sequencing and systems biology converge to identify novel mutations in the L-type calcium channel, CACNA1C, linked to autosomal dominant long QT syndrome; Circ. Cardiovasc. Genet., 6 (3) (2013), pp. 279–289
- [4] D.J. Tester, A.J. Benton, L. Train, B. Deal, L.M. Baudhuin, M.J. Ackerman; Prevalence and spectrum of large deletions or duplications in the major long QT syndrome-susceptibility genes and implications for long QT syndrome genetic testing; Am. J. Cardiol., 106 (8) (2010), pp. 1124–1128
- [5] L.O. Goodwin, D.S. Karabinus, R.G. Pergolizzi, S. Benoff; L-type voltage-dependent calcium channel alpha-1C subunit mRNA is present in ejaculated human spermatozoa; Mol. Hum. Reprod., 6 (2) (2000), pp. 127–136
- [6] I. Splawski, K.W. Timothy, L.M. Sharpe, N. Decher, P. Kumar, R. Bloise, et al.; Ca(V)1.2 calcium channel dysfunction causes a multisystem disorder including arrhythmia and autism; Cell, 119 (1) (2004), pp. 19–31
- [7] L.A. Kelley, S. Mezulis, C.M. Yates, M.N. Wass, M.J. Sternberg; The Phyre2 web portal for protein modeling, prediction and analysis; Nat. Protoc., 10 (6) (2015), pp. 845–858
- [8] A.S. Amin, Y.M. Pinto, A.A. Wilde; Long QT syndrome: beyond the causal mutation; J. Physiol., 591 (Pt 17) (2013), pp. 4125–4139
- [9] P.D. Stenson, E.V. Ball, K. Howells, A.D. Phillips, M. Mort, D.N. Cooper; The human gene mutation database: providing a comprehensive central mutation database for molecular diagnostics and personalized genomics; Hum. Genomics, 4 (2) (2009), pp. 69–72
- [10] T.T. Koopmann, M. Alders, R.J. Jongbloed, S. Guerrero, M.M. Mannens, A.A. Wilde, et al.; Long QT syndrome caused by a large duplication in the KCNH2 (HERG) gene undetectable by current polymerase chain reaction-based exon-scanning methodologies; Heart Rhythm., 3 (1) (2006), pp. 52–55
Document information
Published on 19/05/17
Submitted on 19/05/17
Licence: Other
Share this document
Keywords
claim authorship
Are you one of the authors of this document?