Highlights
- Melanoma cells with acquired resistance frequently express high amounts of β-catenin.
- In these cells a hyper-activation of Stat3 co-occurs leading to an interaction with β-catenin.
- Such BRAFi resistant melanoma cells are sensitive to knockdown of β-catenin and Stat3 or pharmacologic Stat3 inhibition.
Treatment with BRAF inhibitors is the basis of the standard therapy for melanoma patients with BRAFV600 mutated metastases. Although this therapy achieves impressive short-term benefit, many patients suffer from relapse after several months of treatment even if the therapy is combined with MEK inhibitors. The development of therapy resistant tumor cells is a multifactorial transformation and several cellular mechanisms are already described. Here, we found an interaction of two well-known tumor-associated proteins, namely β-catenin and Stat3 in resistant metastatic melanoma cells. Patient data support that these proteins are involved in the resistance to BRAF inhibitor therapy.
Abstract
Acquired resistance to second generation BRAF inhibitors (BRAFis), like vemurafenib is limiting the benefits of long term targeted therapy for patients with malignant melanomas that harbor BRAF V600 mutations. Since many resistance mechanisms have been described, most of them causing a hyperactivation of the MAPK- or PI3K/AKT signaling pathways, one potential strategy to overcome BRAFi resistance in melanoma cells would be to target important common signaling nodes. Known factors that cause secondary resistance include the overexpression of receptor tyrosine kinases (RTKs), alternative splicing of BRAF or the occurrence of novel mutations in MEK1 or NRAS.
In this study we show that β-catenin is stabilized and translocated to the nucleus in approximately half of the melanomas that were analyzed and which developed secondary resistance towards BRAFi. We further demonstrate that β-catenin is involved in the mediation of resistance towards vemurafenib in vitro and in vivo . Unexpectedly, β-catenin acts mainly independent of the TCF/LEF dependent canonical Wnt-signaling pathway in resistance development, which partly explains previous contradictory results about the role of β-catenin in melanoma progression and therapy resistance. We further demonstrate that β-catenin interacts with Stat3 after chronic vemurafenib treatment and both together cooperate in the acquisition and maintenance of resistance towards BRAFi.
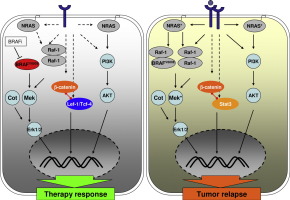
Graphical abstract
Scheme of the change of interacting proteins with β-catenin during the acquisition of resistance to BRAFi from TCF/LEF family members to novel interactants like Stat3. Binding of ligands to different receptor tyrosine kinases like c -met and Egfr family members activate the MAPK signaling pathway and PI3K signaling pathway independently of mutated BRAF. Additional MAPK signaling reactivating mechanisms like the acquisition of novel mutations (asterisk), the formation of Raf heterodimers to the BRAFi. In this environment a switch in the interaction partners of β-catenin to Stat3 occurs further enhancing resistance and contributing to the relapse of the tumor. Dotted lines indicated weak activating potential or indirect mechanisms.
Keywords
β-catenin ; Melanoma ; BRAF ; Stat3 ; Resistance ; Vemurafenib
1. Introduction
Approximately 50% of human melanomas harbor activating V600E/K mutations in the serine/threonine kinase BRAF resulting in constitutive activation of the RAF/MEK/ERK mitogen-activated protein kinase (MAPK) pathway (Davies et al., 2002 , Kumar et al., 2003 and Long et al., 2011 ). Clinical trials with BRAF inhibitors (BRAFis) such as vemurafenib and dabrafenib that specifically target these mutated forms and cause an effective inhibition of the MAPK pathway have shown impressive response rates of > 50% in patients with metastatic melanoma harboring the BRAF V600E mutation (Chapman et al., 2011 , Flaherty et al., 2010 and Hauschild et al., 2012 ) which were confirmed in daily routine in the clinics. However, the acquisition of secondary resistance leading to relapse within 7–12 months was observed in a majority of patients (Carlino et al., 2013 , Chapman et al., 2012 , Menzies and Long, 2014 and Wagle et al., 2011 ). Analyses of the underlying resistance mechanisms in these patients indicated that some vemurafenib-resistant melanoma cells have reactivated the MAPK signaling pathway by alternative mechanisms (Holderfield et al., 2014 ) which promoted successful clinical trials using combinations of BRAF inhibitors with MEK inhibitors (Johnson et al., 2014 , Long et al., 2014 and Robert et al., 2014 ). However, although the median progression-free survival in the group receiving the BRAF and MEK inhibitor was nearly doubled, compared to the mono-therapy group, resistance still developed. Therefore, additional factors independent of MAPK reactivation must play a role in resistance acquisition to BRAFi as well. Indeed, resistance to BRAF or MEK inhibitors correlated with the activation of PI3K/AKT signaling pathway which is achieved by up-regulation of receptor tyrosine kinases (RTKs) such as platelet-derived growth factor receptor (PDGFR) beta, vascular endothelial growth factor receptor (VEGFR), epidermal growth factor receptor (EGFR), insulin-growth factor receptor 1 (IGF1R) and the hepatocyte growth factor (HGF) receptor c-met (Girotti et al., 2013 , Nazarian et al., 2010 , Straussman et al., 2012 , Sun et al., 2014 , Villanueva et al., 2010 and Wilson et al., 2012 ). However, the available data suggest that multiple RTKs are up-regulated or hyper-activated in therapy-resistant cells and that targeting of only one RTK is not sufficient to overcome therapy resistance in general. Since RTKs activate both, the MAPK and PI3K/AKT signaling pathways, a potential strategy to overcome resistance to BRAFi in melanoma cells would be to target common important signaling nodes rather than attempting to target a specific RTK.
One key nodal point is β-catenin, which acts as a transcriptional activator in many signaling pathways, including the Wnt, EGF (Ji et al., 2009 ), HGF (Purcell et al., 2011 ), IGF (Desbois-Mouthon et al., 2001 ), TGF (Zhou, 2011 ) and cadherin pathways (Benham-Pyle et al., 2015 and Zhang et al., 2013 ). Beta-catenin regulates the expression of several genes involved in tumor progression and is part of at least three major functional distinct complexes in a cell. First, β-catenin is a component of adherens junctions where it links cadherins to the cytoskeleton. Second, a cytoplasmic complex regulates its degradation by the interaction with casein kinase 1α (CK1α), glycogen synthase kinase 3 (GSK3), adenomatous polyposis coli (APC) and axin-1. Third, β-catenin accumulates upon activation and translocates to the nucleus, where it classically transactivates transcription of β-catenin/T cell factor/lymphocyte enhancer binding factor (TCF/LEF)-responsive genes together with TCF/LEF family members (Watanabe and Dai, 2011 ).
The role of the β-catenin signaling pathway in therapy resistance has been sparsely analyzed so far. Beta-catenin signaling is involved in AKT activation in prostate cancer cells, whereas inhibition down-regulates AKT activity and induces chemosensitivity in PTEN-mutated prostate cancer cells. Activation of β-catenin signaling mediates chemoresistance in several solid tumors (Saifo et al., 2010 ). It is further speculated that increased chemoresistance is partially linked to β-catenin-mediated up-regulation of drug efflux transporters like MDR-1 (Lim et al., 2009 ). In therapy-resistant colorectal cancer cells, expression of the transcription factor TCF4 was found to be up-regulated and silencing of TCF4 sensitized the cells to chemo- and radiotherapy (Kendziorra et al., 2011 ). Our own data indicate that β-catenin plays a critical role in chemoresistance of melanoma cells (Sinnberg et al., 2011 ). In contrast, it was shown that vemurafenib treated A375 melanoma cells have activated β-catenin signaling which in turn synergized with vemurafenib to induce apoptosis (Biechele et al., 2012 ). Moreover the studies of Biechele and Conrad revealed an additional important role of β-catenin in the induction of apoptosis after MEK inhibition provoking a pro-apoptotic role of β-catenin in the course of MAPK pathway inhibition (Conrad et al., 2012 ). However, the same group recently published that elevated nuclear β-catenin levels correlated with a worse prognosis for BRAFi treated melanoma patients, which developed secondary resistance in their tumors (Chien et al., 2014 ). These contradictory results indicate an urgent need to unravel the role of β-catenin in the mechanisms mediating acquired therapy resistance towards BRAFi.
2. Materials and Methods
2.1. Cell Culture of Melanoma Cell Lines
The human metastatic BRAFV600E mutated melanoma cell lines 451Lu, Mel1617, A375, and SKMel19 were cultured in RPMI 1640 Medium (Gibco life technologies) which was supplemented with 10% fetal calf serum (FCS) (Biochrom/Merck Millipore,) and 1% Penicillin and Streptomycin (Gibco/life technologies). The vemurafenib resistant cell lines were generated by continuous treatment with increasing concentration of PLX4032 up to 2 μM for several months. The culture medium of the resistant cell lines was changed to a PLX4032 free medium 24 h before they were subjected into experiments.
The 451Lu cells stably expressing tetracycline-inducible shRNA against β-catenin (451Lu TetOn-shCTNNB1 ) were generated as previously described ( Sinnberg et al., 2011 ). The shRNA against β-catenin was induced by the application of 1 μg/ml doxycycline to the culture medium for 24 h to 96 h before they were used in the respective experiments.
2.2. Chemicals
The BRAFV600E/K kinase inhibitor PLX4032 vemurafenib (LC Laboratories), Stat3 inhibitors Stattic and S3I-201 (both Selleck) as well as the β-catenin/TCF complex inhibitor PKF-115-584 (Novartis) were used for the specific inhibition of signaling pathways. For Tet-inducible shRNA induction doxycycline (Applichem) was used. Wnt3A and Wnt5A (StemRD) were used for the activation of Wnt signaling.
2.3. Viability assays
4-Methylumbelliferyl heptanoate (MUH) assay was performed for the analysis of the cell viability. 2.5 × 103 cells were seeded into 96 well plate cavities in quintuplicates or sixtuplicates 24 h before treatment. According to the analysis, cells were pre-stimulated with PKF-115-564 (50 ng/ml), with lithium chloride (7.5 mM), with Wnt3A or Wnt5A (up to 100 ng/ml) for 6 h as well as with Stattic (up to 6 μM) or S31–201 (up to 160 μM) for 3 h. Additionally the cells were treated with PLX4032 (up to 20 μM) for 72 h. Viability analysis was performed after a washing step using PBS and subsequent incubation of the cells in 100 μg/ml 4-methylumbelliferyl heptanoate diluted in PBS for 1 h at 37 °C. The fluorescence (ex355nm/em460nm) was detected in a Tristar fluorescence microplate reader (Berthold).
Alamar blue viability assay was used to analyze the viability of transfected cells for normalizing reporter assay signals to cellular viability when no Renilla-CMV was used. Briefly 1 mg/ml alamar blue stock solution was pre-diluted in culture medium (1:10) and 10 μl of this solution was added to 100 μl culture medium of each 96 well cavity. After incubation for 1 h at 37 °C the fluorescence of resofurin was measured in fluorescence microplate reader (Berthold, Germany) at ex540nm/em640nm. Background subtracted sample values were used for normalization of reporter signals.
2.4. Luciferase Reporter Assays
SuperTopFlash, pcDNA3.1-GLuci-CRE/-AP1/-NFAT as well as pLuc TKS3 (Stat3) reporter were transiently transfected for the analysis of specific pathway activity. 2.5 × 105 melanoma cells were seeded into 6 well plate cavities 24 h prior to transfection. Reporter plasmids (2 μg/well) and CMV renilla plasmids (120 ng/well) were co-transfected using ScreenFect A Transfection Kits (Genaxxon bioscience) or Lipofectamine 3000 Transfection Reagent (Life Technologies) according to the manufacturers protocols for 24 h. Then 1 × 104 transfected cells were seeded into 96 well plate cavities at minimum in quintuplicates and optionally pre-treated with 15 mM lithium chloride for 6 h. Additional treatments were PLX4032 (up to 10 μM), Stattic (up to 2 μM) or Wnt5a and Wnt3a (100 ng/ml) for further 24 h. Cells were lysed in 50 μl passive lysis buffer (Promega). 10 μl of lysates or 10 μl of supernatant of the culture medium (Gaussia reporters) was used for the measurement of the firefly and renilla luciferase activity in a Tristar luminometer (Berthold). Luciferase activity was measured using D-luciferin or renilla and gaussia substrate coelentarazine as previously described (Albert and Silvia, 2012 ).
2.5. Cell Cycle Assay
2 × 105 melanoma cells per 6 well cavity were seeded 24 h prior to treatment. DMSO (0.02–0.05%) treated control cells and PLX4032 (2 or 5 μM) treated cells were analyzed in triplicates. Cells were commonly treated for 72 h. After permeabilization of the cells in 70% ice-cold ethanol for at least 1 h, they were re-suspended in PBS with 100 μg/ml RNAseA (Applichem) and 50 μg/ml propidium iodide (Sigma) and stained for 30 min. FACS analysis for the detection of the distribution of the cells in the each cell cycle phases was performed with a LSRII FACS (BD) using FACSDiva software (BD).
2.6. SA-Beta-Galactosidase Staining
The Senescence Cells Histochemical Staining Kit (Sigma-Aldrich) was used according to the manufacturers recommendations. Briefly, 1.0 × 105 cells were seeded into 12 well plates and treated with the indicated concentrations of vemurafenib for 72 h. In case of co-treatment with doxycycline for the induction of shRNA, a pre-treatment with 1 μg/ml doxycycline for 24 h was performed. After fixation and washing of the cells they were stained using the x-gal substrate solution for approx. 3 h at 37 °C without CO2 . The percentage of senescent cells was calculated using the ratio of blue cells to total cell number per microscopic picture. Three independent samples were analyzed in quadruplicates.
2.7. Immunohistochemistry
For immunofluorescence staining of cultivated cells, 2.5 × 104 cells were seeded per chamber on 8 well chamber slides (BD Falcon) and grown for 24 h. The cells were treated with 0.02% DMSO or 2 μM PLX4032 for 24 h, washed with PBS and fixed in 4% formaldehyde for 15 min. They were blocked in PBS/1%BSA/0.3% Triton-X100 and incubated overnight with the primary antibody against β-catenin (1:100 dilutions, Cell Signaling #9562). For the detection, the samples were incubated with the secondary antibody Alexa Fluor® 647-conjugated AffiniPure Fab Fragment Donkey anti-Rabbit IgG(H + L) (Jackson ImmunoReserarch) for 2 h. The nuclear staining was performed by monomeric cyanine nucleic acid stain (YO-PRO-1) (Invitrogen). Immunohistochemistry staining of clinical FFPE specimens was performed with a β-catenin specific antibody (Cell Signaling #9562) diluted 1:50 in PBS containing 0.3% Triton-X100 and 1% BSA. Briefly 5 μm FFPE tissue sections were de-paraffinized and antigen retrieval was performed in citrate buffer pH 6 in a pressure cooker for 2 min under pressure before a slow cooling down of the samples in the hot buffer. Afterwards tissue sections were stained according to the manufacturers protocol (Thermo Scientific Lab Vision UltraVision LP Detection System: AP Polymer) using FastRed (Thermo Scientific Lab Vision Liquid Fast-Red Substrate System) as substrate.
2.8. Western Blotting
Melanoma cells were harvested with Trypsin/EDTA (Biochrom/Merck Millipore,), washed two times with cold PBS and lysed in RIPA Lysis Buffer (Thermo Scientific). For preparation of nuclear and cytoplasmic protein extracts NE-PER Nuclear and Cytoplasmic Extraction Reagents (Thermo Scientific) were used according to the manufacturers protocol. 15 μg of whole cell lysate protein samples or cytoplasmic protein samples as well as 5 μg of nuclear fraction protein samples were used for the SDS-PAGE followed by blotting onto PVDF membranes (Roche). After blocking for 60 min in PBS-T (0.1% Tween-20) with 5% dry milk the following primary antibodies were applied on a roller mixer overnight: phospho-Stat3 (Ser727), phospho-Stat3 (Tyr705), Stat3 (79D7), Stat3 (124H6), β-actin (D6A8), β-catenin (D10A8), phospho-p42/44 MAPK (ERK1/2), p42/44 MAPK (ERK1/2), LEF1 (C12A5), TCF4 (C48H11), phospho-AKT (Ser473), AKT, phospho-GSK-3β (Ser9) (D85E12), GSK-3β (D5C5Z), Wnt5a/b (C27E8), GAPDH (14C10) (all Cell Signaling Technology); β-catenin E-5 (sc-7963), lamin-B (sc-6216) (Santa Cruz Biotechnology); MITF (C5) (Cat.No. OP126L) (Calbiochem.). Immunodetection was performed using anti-rabbit or anti-mouse secondary peroxidase-conjugated antibodies (Cell Signaling Technology). Visualization was done with using ECL (Thermo) or ECL prime (GE Helathcare Lifesciences). Anti-rabbit (Cell Signaling Technology) and anti-goat (Novus) AP conjugated antibodies were used as well. In these cases detection was performed with CDP-Star (Roche) according to the manual.
2.9. Miniaturized GST-Pulldown-Assay for the Quantification of “Free” β-Catenin
The quantity of the “free” β-catenin pool which is involved in the Wnt/β-catenin signaling was determined using a miniaturized bead-based GST-pulldown assay as described previously (Luckert et al., 2012 ). Total β-catenin was measured using a bead-based sandwich immunoassay for normalization. The analyses were performed with 20 μg protein extract derived from sensitive and resistant melanoma cell cultures. Results have been normalized by forming the average of total β-catenin and “free” β-catenin.
2.10. Co-immunoprecipitation
Cell pellets were homogenized in 200 μl lysis buffer (10 mM Tris/Cl pH7.5, 150 mM NaCl, 0.5% NP40, 1 μg DNaseI, 2 mM MgCl2 , 2 mM PMSF, 1 × phosSTOP phosphatase inhibitor (Roche), 1 × protease inhibitor mix M (Serva) by repeated pipetting for 40 min on ice. After a centrifugation step (10 min at 18,000 × g ) the soluble protein fraction was incubated with 50 μl of the agarose-coupled BC2-nanobody (Traenkle et al., 2015 ) for 12 h on an end-over-end rotor at 4 °C. The bead pellet was washed two times in 0.5 ml dilution buffer (10 mM Tris/Cl pH 7.5, 150 mM NaCl, 2 mM PMSF). After the last washing step the beads were transferred to a new cup, resuspended in 2 × SDS-containing sample buffer and boiled for 10 min at 95 °C. Samples (5% input, 5% non-bound and 10% bound) were analyzed by SDS-PAGE followed by western blotting. Immunoblots were probed with antibodies directed against β-catenin, GAPDH, MITF, and LEF-1.
For the detection of the interaction with Stat3, proteins were cross-linked using 0.4% formaldehyde/PBS solution before the precipitation. 5 × 106 cells were resuspended in 2 ml fixing solution and incubated at room temperature for 4 min on a roller mixer before centrifugation at 320 × g for 3 min. The supernatants were removed and the cells were incubated in 1 ml ice-cold 1.25 M glycine/PBS for 1 min to quench the formalin reaction and were centrifuged for 3 min. The cells were washed in 5 ml PBS and lysed in 250 μl RIPA-Buffer (Thermo Scientific) for 60 min. The precipitation was performed according to the co-immunoprecipitation protocol (Cell Signaling). The proteins were precipitated using β-catenin (D10A) XP Rabbit (Cell Signaling Technology), β-catenin (E-5) sc-7963 mouse antibody (Santa Cruz Technology), Stat3 (D3Z2G) Rabbit mAb and Stat3 (124H6) Mouse mAb (both Cell Signaling Biotechnology) and the bound proteins were detected.
2.11. siRNA Transfection
20 nM siRNA (all synthesized by biomers.net Germany) against Stat3 (sense: aacuucagacccgucaacaaa-dTdT;; antisense: uuuguugacgggucugaag-dTdT) and β-catenin (sense: gguggugguuaauaaggcu-dTdT; antisense: agccuuauuaaccaccacc-dTdG) were reversely transfected using the riboxx FECT (riboxx Life Sciences) on 96 well plates. Therefore, the riboxxFECT (1:25) and siRNA were separately diluted using Opti-MEM (Life technologies) and the solutions were gently mixed in a 1:1 ratio. The solution was incubated for 15 min and subsequently, 50 μl of the solution was transferred to each 96-well cavities. Additionally, 2.5 × 103 cells resuspended in 50 μl culture medium were added to each well and incubated for 24 h at 37 °C. The transfected cells were subsequently treated with up to 20 μM PLX4032 for 72 h and the viability was measured via MUH assay.
2.12. Lentiviral Gene Transfer
Stat3 overexpression lentivirus was produced using HEK 293T cells (Biocat, Germany) transfected with human STAT3 cloned into pLX304 (HsCD00443857 DNASU plasmid repository (Seiler et al., 2013 )) as well as second-generation packaging and envelope plasmids pCMVΔR8.2 and pMD2.G. Melanoma cells were transduced with lentivirus containing supernatants in the presence of 8 μg/ml polybrene (Sigma) and cultured in 5 μg/ml blasticidin (Merck Millipore) containing selection medium.
2.13. Xenograft Melanoma Model
For the in vivo tumor growth assays, 1 × 106 vemurafenib resistant 451Lu cells stably expressing tetracycline-inducible shRNA against β-catenin were subcutaneously injected into SCID mice. All mice were daily treated with 25 mg/kg vemurafenib (LC Laboratories) i.p. for 25 days post injection, until they developed approx. 100 mm3 tumor nodules. The mice were randomized into four groups (n = 5). The first group was subsequently fed with 1 mg/ml doxycycline (Applichem) in the drinking water, the second group received daily injections of 25 mg/kg vemurafenib i.p, the third group received both treatments and the fourth group served as untreated control group. Drinking water was generally supplemented with 1% sucrose to reduce the bitter taste due to doxycycline. The Tumor size was monitored for 40 days post injection by calliper measurements of tumor length and width. The tumor volume was calculated using the following formula: V = 0.4 × length × width2 . All animal experiments were approved by the regional council (Regierungspraesidium Tuebingen Az35/9185.81-2).
2.14. PET Imaging
Animals were fasted for 6 h prior to the FDG injection and doxycycline treatment was interrupted due to the necessary sucrose addition to the drinking water. ~ 13 MBq FDG in a max. volume of 100 μl were injected i.v. into the tail vein under 1.5% isoflurane narcosis evaporated in oxygen at a flow rate of 0.5 l/min (Abbott, Wiesbaden, Germany) and the animals were kept under anesthesia for 55 min. post-injection in a heated box. Blood glucose and body weight measurements were performed immediately before FDG injection. Subsequently animals were placed on a carbon bed and scanned for 10 min in an Inveon small animal PET scanner (Siemens Preclinical Solutions, Knoxville, TN, USA). Body temperature was maintained at 37 °C by a heating pad and a rectal temperature sensor. Image reconstruction was performed using Inveon Acquisition Workplace 1.5.0.28 (Siemens Preclinical Solutions, Knoxville, TN, USA) with an iterative ordered-subset expectation maximization algorithm (OSEM2D) with four iterations. No attenuation and scatter correction was applied, according to our standard protocol for PET imaging with mice. The reconstructed voxel size was 0.776 × 0.776 × 0.796 mm. Images were analyzed in Inveon Research Workplace (Siemens Preclinical Solutions, Knoxville, TN, USA).
2.15. Statistical Analysis
Microsoft Excel and Graphpad Prism 6.0 were used for the statistical analyses of the data. Graphs show mean values with their SD unless differently mentioned. Statistic used for p-value calculations and significance determinations are given in the corresponding figure legends. p-values < 0.05 were considered statistically significant (with * for p < 0.05, ** for p < 0.01 and *** for p < 0.001). Dose response curves were fitted using Graphpad Prism 6.0 for IC50 calculations (using mostly the nonlinear regression curve fit log(inhibitor) vs. response with variable slope).
3. Results
3.1. A Subset of Vemurafenib Resistant Melanoma Cells Exhibit Stabilized Nuclear β-Catenin In Vivo and In Vitro
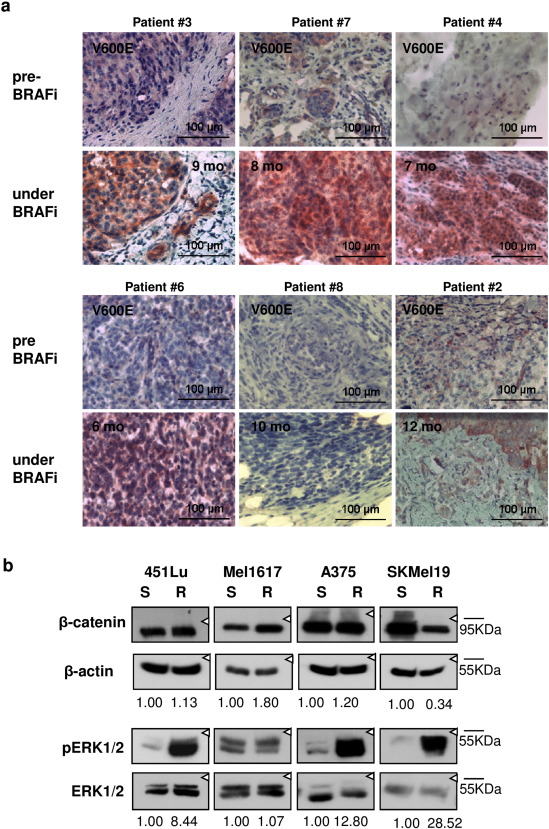
In our previous work we found elevated levels of nuclear β-catenin in several metastatic melanoma biopsies and cell lines compared to cell lines of radial growth phase origin (Sinnberg et al., 2010 and Sinnberg et al., 2011 ). To investigate whether β-catenin levels are also altered during resistance development to BRAFi treatment, we collected tumor biopsies from eight stage IV melanoma patients before initiation of vemurafenib treatment and after development of drug resistance for analysis of β-catenin protein expression by immunohistochemistry. Time to resistance development in the presented cases was 5 to 12 months (Fig. 1 a). We observed accumulation of β-catenin in 4 out of 8 resistance-acquired tumors (50%) in comparison to the β-catenin levels in excised lesions collected before the start of the treatment (Fig. 1 a). In one sample pair (12.5%) a diverse expression pattern was seen including areas with increased β-catenin levels, whereas in three samples no increase in β-catenin expression was detected (37.5%). We used a tissue microarray to compare the observed frequencies of strong β-catenin staining with BRAFi naïve melanomas (Supplementary Fig. 1 ). Among the 270 primary melanomas 39 showed strong β-catenin expression (14.5%) and 203 a moderate staining (60.7%). None of the 13 metastatic melanomas had a strong signal intensity (0%) and seven biopsies showed a moderate (53.8%) staining pattern. This clearly shows that the high expression level of β-catenin in 50% of the BRAFi resistant tumor biopsies is extraordinary.
|
|
|
Fig. 1. Beta-catenin expression levels increase in BRAFi resistant melanomas cells compared to the sensitive parentals. a) Immunohistochemical staining for β-catenin of clinical specimens before and after the acquisition of resistance to BRAFi. Beta-catenin expression levels are shown in red (Fast Red substrate) with hematoxylin counter staining (scale bar is 100 μm). b) Immunoblots of whole cell lysates from sensitive and resistant pairs of the melanoma cell lines 451Lu, Mel1617, A375 and SKMel19 showing the expression levels of β-catenin and (phospho) Erk1/2. Semiquantitative analysis was performed by building the ratios of (β-catenin:β-actin) and (p-ERk1/2:Erk1/2) and normalization to the sensitive cells. |
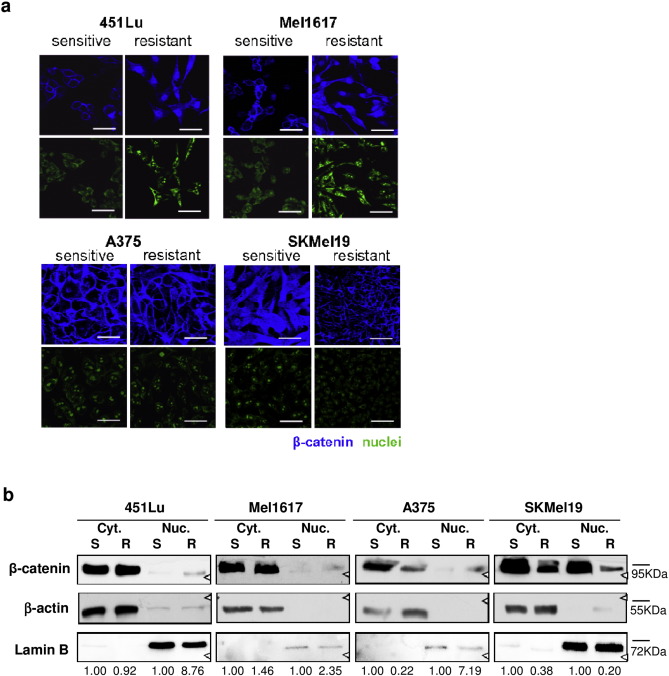
To unravel the functional relevance of β-catenin expression on BRAFi resistance development we generated BRAFi resistant melanoma cell lines (Mel1617-R, 451Lu-R, A375-R, and SKMel19-R) by continuous treatment with vemurafenib. The chronically treated cells were significantly more resistant to BRAF inhibition with less affected cellular viability and reduced apoptosis induction compared to the respective parental sensitive cell lines (Mel1617-S, 451Lu-S, A375-S, and SKMel19-S) (Supplementary Fig. 2a, b ). On a molecular level the resistant cell lines showed a strong MAPK pathway activity based on high phosphorylation of Erk1/2 (Fig. 1 b). Phospho-AKT was moderately increased in resistant Mel1617 and A375 cells (Supplementary Fig. 1f ). We tested for different resistance mechanisms that lead to the re- and hyperactivation of the MAPK signaling pathway. No novel mutations in NRAS (codon 61) or MEK1 (exons 2, 3 and 6) could be detected by Sanger sequencing in the resistant cell lines (Supplementary Fig. 1c ). Interestingly A375-R and SKMel19-R cells showed signs of highly enriched truncated BRAF (t-BRAF) splice products which are known to be associated with BRAFi adaptation and resistance mediation (Supplementary Fig. 1d ). We further analyzed the expression of several RTKs (KIT, PDGFRB, MET, IGF1R, EGFR), the alternative MEK activator COT and BRAF (Supplementary Fig. 1e ) on a transcriptional level. PDGFRB and EGFR mRNA levels were upregulated especially in 451Lu-R cells whereas SKMel19-R had higher levels of mRNA coding for MET. Total β-catenin protein levels were at similar levels or only modestly increased in Mel1617-R and 451Lu-R cells and even decreased in SKMel19-R cells. We found no consistent change in the expression of components of other signaling pathways like PTEN in cellular lysates of resistant cells compared to their sensitive counterparts (Supplementary Fig. 1f ) pointing to the diversity of resistance mechanisms in our cell lines. Immunofluorescence staining and confocal microscopy indicated a prominent nuclear localization of β-catenin in vemurafenib resistant melanoma Mel1617-R, 451Lu-R and A375-R cells in comparison to the sensitive parental cells (Fig. 2 a). Further western blot analyses of fractionated cell lysates confirmed that the resistance-acquired cell lines 451Lu-R, Mel1617-R and A375-R exhibit increased nuclear β-catenin levels (Fig. 2 b). We further analyzed the protein expression of important β-catenin regulating proteins (Axin1, APC, sFRP1) in order to find the trigger of β-catenin up-regulation and nuclear translocation. No consistent changes were detected and resistant cells expressed rather high levels of APC, a component of the cytoplasmic degradation complex of β-catenin and SKMel19-R cells produced increased amounts of sFRP1 (Supplementary Fig. 1g ). Taken together, these results indicate that a major subset of vemurafenib resistant melanoma cells with high MAPK signaling activity shows increased nuclear β-catenin levels in vitro and in vivo .
|
|
|
Fig. 2. Nuclear localized beta-catenin in BRAFi resistant melanomas cells compared to the sensitive parentals. a) Immunofluorescence staining for β-catenin (blue) and nuclear YOPRO-1 staining (green) with confocal microscopy for expression and localization analysis of β-catenin (white scale bars represent 50 μm). b) Immunoblot of cytosolic and nuclear extracts showing the increased nuclear localization of β-catenin in the resistant cells of the cell lines 451Lu, Mel1617 and A375. Semiquantitation was performed densitometrically by using the ratios of [β-catenin:β-actin] for cytosolic and [β-catenin:LaminB] for nuclear fractions of the sensitive and resistant cell line pairs. All ratios were normalized to the sensitive parental cell line to compare the sensitive to the corresponding resistant cell line. |
3.2. β-Catenin Knockdown Sensitizes Melanoma Cells Towards Vemurafenib Treatment In Vitro and In Vivo
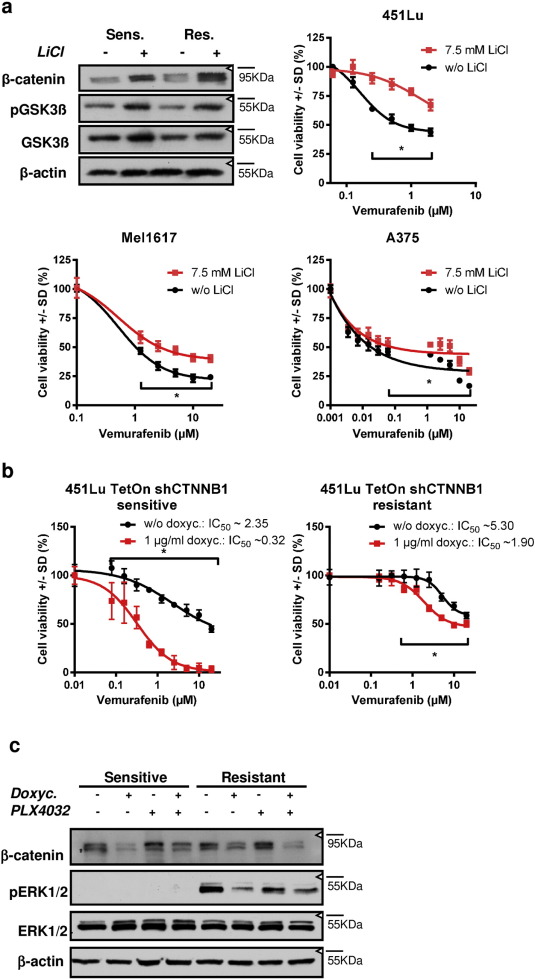
In the next step, we analyzed whether β-catenin is functionally involved in the resistance mechanisms to BRAF inhibitors. Therefore, we modulated cellular β-catenin levels in different ways in order to test any β-catenin dependent influence on the resistance towards vemurafenib. At first, we stabilized β-catenin by the inhibition of GSK3β using lithium chloride in 451Lu-S, Mel1617-S and A375-S cells. As expected lithium treatment resulted in an increased phosphorylation of GSK3β at serine 9 and accumulation of β-catenin. This went along with an attenuation of the growth inhibitory effect of vemurafenib in BRAFi-sensitive cells in a mild but significant manner (Fig. 3 a).
|
|
|
Fig. 3. Beta-catenin is directly involved in the efficacy of the response to BRAFi and suppresses growth inhibition a) Cell viability assay (MUH) of melanoma cell lines pre-treated with 7.5 mM LiCl for 24 h for stabilization of β-catenin. Immunoblots show the stabilization of β-catenin after treatment of 451Lu cells with LiCl. After the pre-treatment, cells were treated with increasing concentrations of vemurafenib for 72 h before the assessment of cell viability. Black symbols represent sensitive control cells and red symbols represent LiCl pre-treated cells. Signals were normalized to the control cells without vemurafenib treatment. Mean +/ SD values of six replicates are shown. Multiple t-tests with Holm–Šídák correction were used to compare data points of the two curves and p < 0.05 was considered as significant (asterisk). b) Cell viability assays (MUH) after Tet-inducible knockdown of β-catenin in 451LuTet-S (left diagram) or 451LuTet-R cells (right diagram). Knockdown was induced by pre-treatment with 1 μg/ml doxycycline for 24 h (red symbols) and compared to cells without doxycycline treatment (black symbols). Multiple t-tests with Holm–Šídák correction were used to compare data points of the two curves and p < 0.05 was considered as significant (asterisk). F-test revealed a significant different IC50 of the fitted curves. c) Immunoblot analysis for β-catenin and (phospho-) Erk1/2 shows protein level changes after induction of shCTNNB1 in combination with vemurafenib treatment 24 h after treatment in comparison to vehicle control treated cells. |
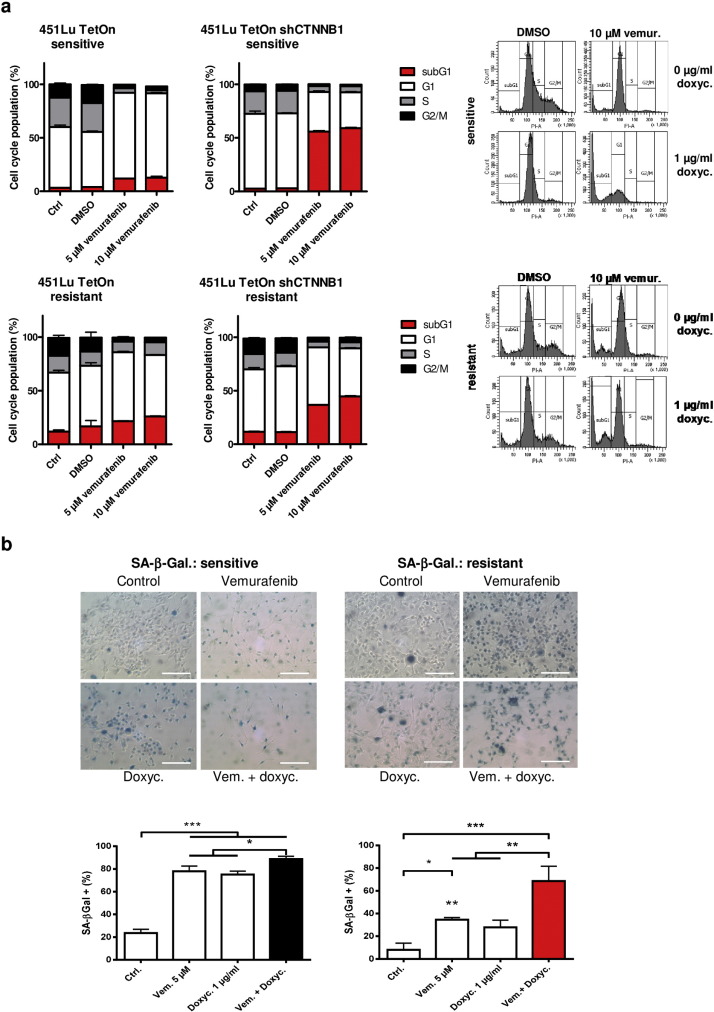
To directly address the effects of β-catenin on resistance to BRAF inhibition, we generated vemurafenib resistant 451Lu cells that had been stably transfected with a doxycycline inducible and β-catenin (CTNNB1 ) specific shRNA vector pTER-shCTNNB1 ( Sinnberg et al., 2011 and van de Wetering et al., 2003 ) before initiation of long-term treatment with vemurafenib for the development of resistance to BRAFi. The time until vemurafenib-dependent growth inhibition was overridden by resistance mechanisms was doubled to 8 weeks (compared to almost 4 weeks without induction β-catenin specific shRNA) if the chronic BRAFi therapy was combined with permanent knockdown of β-catenin by the addition of 1 μg/ml doxycycline (Supplementary Fig. 3a ). Treating of the cells with doxycycline resulted in a marked reduced β-catenin protein level in the parental sensitive as well as in the 451Lu Tet-inducible cells with acquired resistance to vemurafenib. Interestingly, the induced β-catenin knockdown significantly increased the susceptibility of the parental sensitive 451Lu cells towards vemurafenib in the MUH viability assay and cell cycle analysis (Figs. 3 b, 4 a). The decreased viability went along with an enhanced caspase 3/7 activity 48 h after combined knockdown with vemurafenib treatment. Apoptosis induction was inhibited by pre-treatment of the cells with 7.5 mM LiCl as β-catenin stabilizing agent (Supplementary Fig. 3b ).
|
|
|
Fig. 4. Knockdown of β-catenin enhances the BRAFi mediated induction of apoptosis and senescence. a) Cell cycle analysis of Tet-inducible 451LuTet-S (upper panel) and 451LuTet-R (lower panel) cells after treatment with 5 or 10 μM vemurafenib for 72 h. Doxycycline pre-treatment for 24 h (right bar diagrams) was used to induce β-catenin specific shRNA. Representative FACS histograms of the cell cycle analysis comparing vehicle (DMSO) treated cells with vemurafenib (10 μM) are shown. Cell cycle distributions represent the result of three biologic replicates (mean +/− SD). b) SA-associated β-galactosidase staining after 72 h of vemurafenib treatment of the above mentioned Tet-inducible cells (scale bars represent 200 μm). Clear blue cells were counted and normalized to the total number of cells using four different microscopic pictures (top row) per treatment. Mean numbers +/− SD are shown from three independent biological replicates (bottom row). ANOVA analyis was performed using Tukeys multiple comparisons test (asterisks). |
Furthermore, the doxycycline-induced knockdown of β-catenin even partially re-sensitized the previously insusceptible 451Lu resistant melanoma cells in terms of reduction of viability and induction of apoptosis (Fig. 3 b, 4 a). Diminished phospho-Erk1/2 levels co-occurred with the knockdown of β-catenin (Fig. 3 c) resulting in markedly reduced cell proliferation and growth arrest, which was mainly mediated by a cell cycle arrest in G1 and the induction of senescence as determined by increased SA-β-galactosidase activity (Fig. 4 a, b). In order to exclude any sensitizing effect of doxycycline on the susceptibility of BRAFV600E melanoma cells to BRAFi we combined the vemurafenib treatment with 2 μg/ml doxycycline for three consecutive days and could not see any significant difference between the mono-therapy using vemurafenib in ascending concentrations and its combination with the antibiotic in the viability assay (Supplementary Fig. 3c ). These data indicate that β-catenin is involved in the development as well as the maintenance of resistance towards vemurafenib.
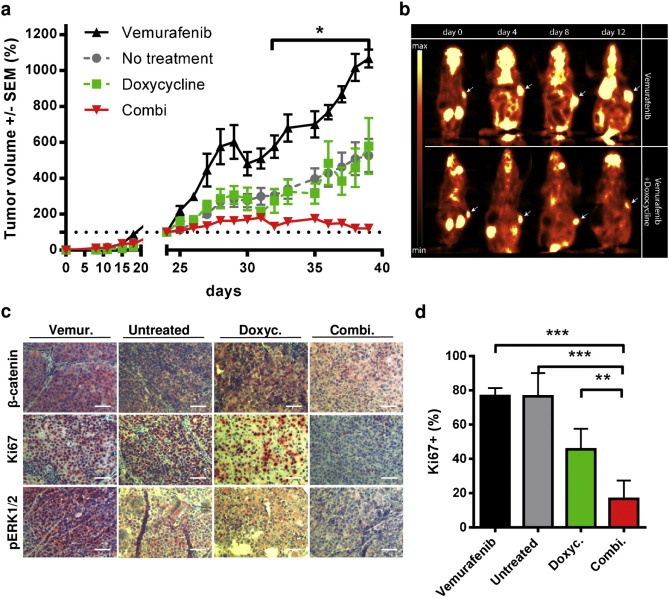
Subsequently, we evaluated whether the observed impact of β-catenin on the efficacy of BRAFis in terms of growth inhibition in vitro is also a relevant phenomenon under in vivo conditions. For this purpose resistant 451Lu cells harboring the Tet-inducible CTNNB1 -specific shRNA were subcutaneously injected into SHO mice ( Fig. 5 ). The mice received daily intraperitoneal (i.p.) injections of vemurafenib (25 mg/kg) intraperitoneal (i.p.) for 24 days until a median tumor volume of approximately 100 mm3 was reached. Mice were randomized into four different therapy groups: the 1st group was continued with daily injections of vemurafenib (i.p. 25 mg/kg), the 2nd group received drinking water supplemented with doxycycline (1 mg/ml), the 3rd group was treated with a combination of vemurafenib and doxycycline and the 4th group did not receive any further therapy. The therapy group of mice treated with vemurafenib alone exhibited rapid tumor growth as detected by caliper measurements and PET imaging (Fig. 5 a, b). In comparison, a doxycycline mediated down-regulation of β-catenin significantly decelerated tumor growth compared to continuous BRAFi therapy. A reduced tumor growth after knockdown of β-catenin could be also observed in the group that did not receive further vemurafenib injections (Fig. 5 a). Immunohistochemistry staining of isolated tumors from double treated mice indicated a clearly decreased protein expression of the proliferation marker Ki67 and phospho-ERK1/2 in comparison to the other therapy groups, confirming the reduced tumor cell growth (Fig. 5 c, d) and the previous in vitro data. Analysis of the time points when the abort criteria were reached showed a survival benefit for mice receiving combinatorial therapy of BRAFi and β-catenin knockdown. None of these double-treated mice reached the maximum tolerable tumor size (15 mm in diameter) within the observation time of 40 days, in contrast to the mice of the control and single treated groups (Supplementary Fig. 3d ).
|
|
|
Fig. 5. Knockdown of β-catenin synergizes with vemurafenib for the inhibition of tumor growth of BRAFi resistant melanomas. a) Xenograft experiment using Tet-inducible 451Lu-R cells for s.c. injection (1 × 106 cells) into SHO mice. After formation of 100 mm3 tumor nodules upon daily i.p. injections with vemurafenib (25 mg/kg) the mice were randomized into four treatment groups: i) vemurafenib treatment (black), ii) untreated (grey), iii) shRNA induction by doxycycline (1 mg/ml) in the drinking water ad libitum and (green) iv) combination (red) (n = 5 per group). The tumor growth was daily monitored by caliper measurements and normalized to the size at day 0 of treatment. Multiple t tests were used to determine the data points of the combination group with p < 0.05 (asterisk). b) Positron emission tomography using F18-FDG was performed on days 4, 8 and 12 post-start of therapy. c) Immunistochemistry of removed tumors from SHO mice (from 2F). Tumors of every group were stained for β-catenin expression, phospho-Erk1/2 and Ki67 using Fast Red as substrate. Representative microscopic pictures are shown of every group (scale bars indicate 100 μm). d) Ki67 positive cells were counted and normalized to total cell numbers as indicator for cell proliferation. Three different tumors per group were used for data analysis. Mean percentages +/− SD are shown. ANOVA analysis was performed using Tukeys multiple comparisons test (asterisks). |
3.3. β-Catenin Acts Independent of the TCF/LEF Dependent Canonical Wnt-Signaling in the Resistance Mechanism Towards Vemurafenib Treatment
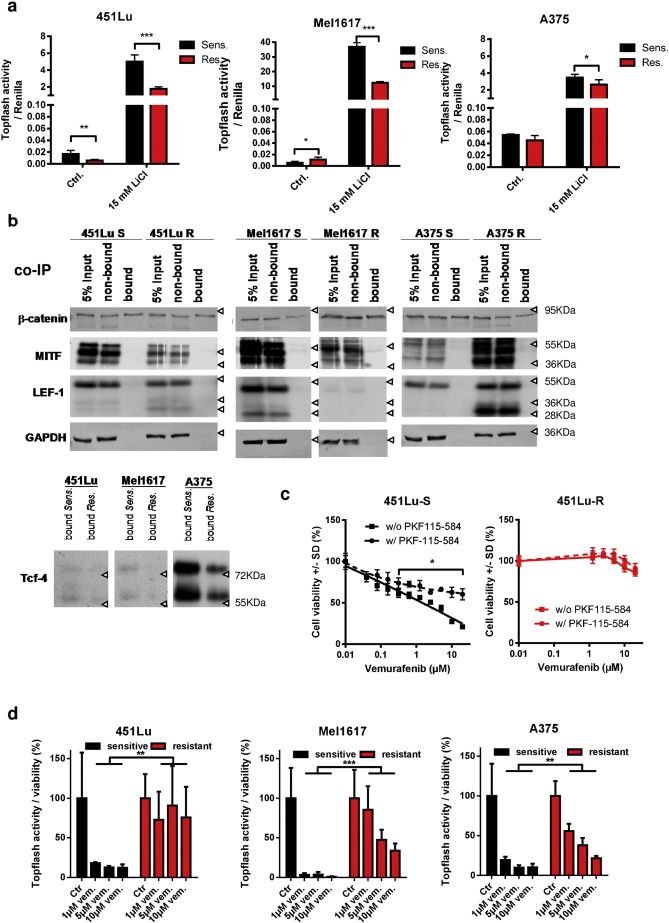
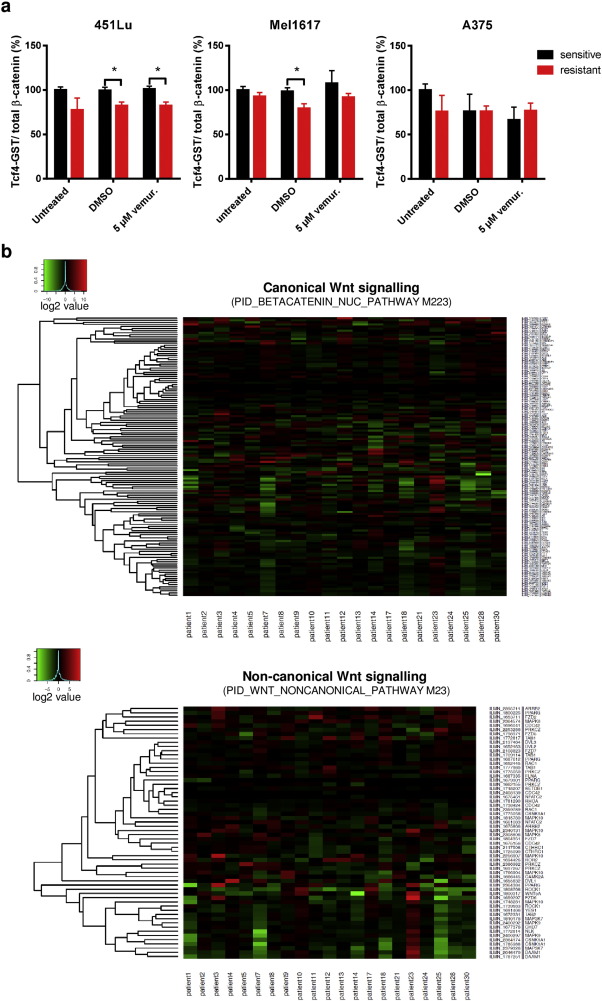
Since nuclear accumulation of β-catenin is strongly indicative for its transcriptional activity in the canonical Wnt signaling pathway, we analyzed the impact of nuclear β-catenin on TCF/LEF dependent gene transcription in BRAFi resistant cells. Therefore, TCF/LEF dependent transcription was determined using a luciferase reporter assay (Super8xTOPFlash) in vemurafenib sensitive and resistant Mel1617, 451Lu and A375 melanoma cell lines (Fig. 6 a). Since the three resistant cell lines exhibit increased nuclear accumulation of β-catenin, we expected to find an increase in luciferase activity in resistant clones versus their sensitive counterparts. Surprisingly, the basal level of the TCF/LEF dependent signal was not clearly induced in resistant cells and even reduced in the 451Lu-R cells compared to the sensitive parentals (Fig. 6 a). Stabilization of β-catenin using lithium chloride (15 mM) resulted in a markedly increased TOPFlash luciferase signal in resistant and sensitive cells but revealed a significantly diminished induction factor of TCF/LEF dependent gene transcription in the resistant cells. A lower TOPFlash luciferase signal is suggestive for reduced interaction of β-catenin with TCF/LEF. Therefore we performed co-immunoprecipitation experiments of β-catenin in 451Lu, Mel1617 and A375 cells with subsequent immunoblotting for microphtalmia-associated transcription factor (Mitf), Lymphoid enhancer-binding factor 1 (LEF-1) or T -Cell-Specific Transcription Factors (TCF). All three are known to be important interactants of nuclear β-catenin in melanocytic cells. In the BRAFi resistant melanoma cell lines the interaction with LEF-1, TCF-4 and Mitf was at very weak levels, confirming that the additional amount of nuclear β-catenin is not mainly involved in the canonical Wnt signaling pathway in these cells. Of note the expression of TCF-4 and LEF-1 as well as of Mitf was diminished in the resistant cell lines Mel1617-R and 451Lu-R compared to the sensitive parentals ( Fig. 6 b). In a GST-pulldown assay employing GST-tagged TCF-4 we also found a rather significantly reduced interaction capacity of β-catenin with TCF-4 in resistant versus sensitive melanoma cells (451Lu and Mel1617) confirming that the increased nuclear β-catenin levels in the resistant cells do not increase the canonical Wnt signaling capacity (Supplementary Fig. 4a ). This was in line with the analysis of previously published transcriptome data on fifty-nine melanomas from twenty-one different patients before chronic therapy with a BRAFi and after resistance had developed (Rizos et al., 2014 ). Transcriptome analysis using gene sets related to the regulation of nuclear β-catenin signaling and target gene transcription did not reveal a differential expression of the β-catenin interaction partners LEF1 or TCF7L2 (TCF-4) (Supplementary Fig. 4b ).
|
|
|
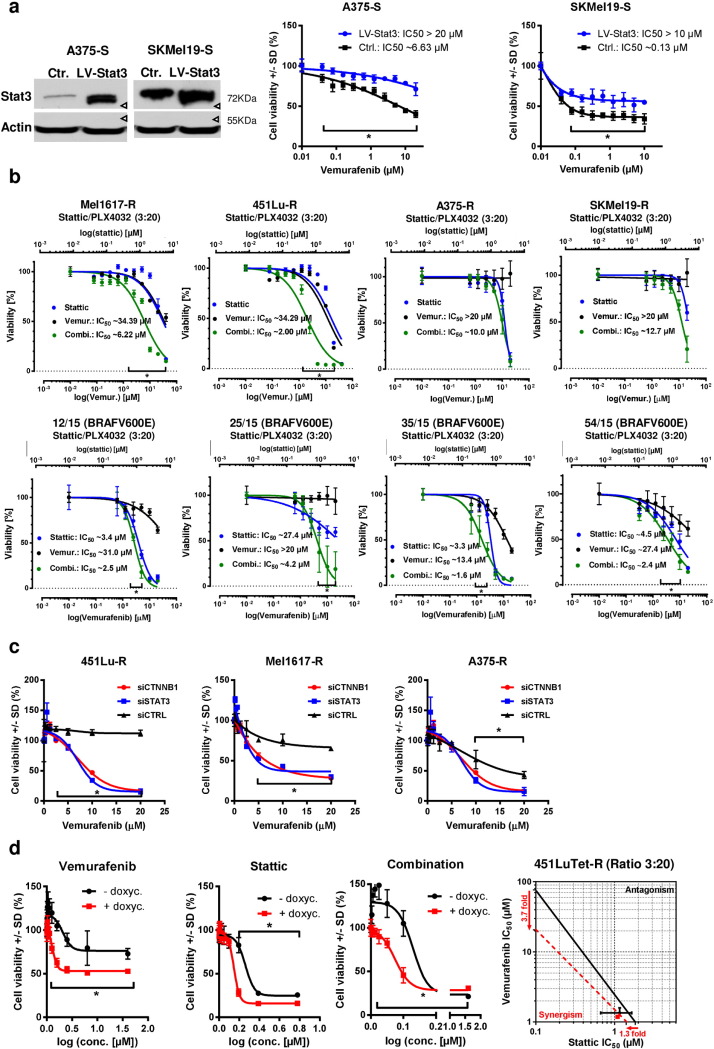
Fig. 6. Accumulated β-catenin in BRAFi resistant melanoma cell lines acts independent of the canonical Wnt signaling pathway and the TCF/LEF factors. a) TOPflash luciferase reporter assays were done to measure the transcriptional activity of TCF/LEF complexes. Sensitive (black bars) and resistant (red bars) melanoma cell lines were transfected with the reporter construct plus CMV-renilla luciferase as a normalization control and treated for 24 h with the indicated concentrations of vemurafenib. For the induction of the full signaling activity a pre-treatment with 15 mM LiCl was performed. Firefly luciferase activity was normalized to renilla activity. Mean values and standard deviations of six samples are presented. Multiple t-tests with Holm–Šídák correction were used to compare data of sensitive and resistant samples and p < 0.05 was considered as significant (asterisk). b) Co-immunoprecipitation experiments for the detection of interactions of β-catenin with TCF4, LEF1 and Mitf. Soluble lysates were prepared from the indicated sensitive and resistant melanoma cells and incubated with an immobilized β-catenin specific nanobody. Input, non-bound and bound fractions were separated on a SDS-PAGE followed by immunoblot analysis for the transcription factors TCF4, LEF1 and MITF. c) Cell viability assay (MUH) for testing the effects of released β-catenin from the complexes with TCF/LEF by 50 nM PKF115–584 (circles with dashed curve). The inhibitor was pre-incubated for 6 h before the treatment with increasing concentrations of vemurafenib for 72 h. The assay was measured in quintuplicates. Mean values +/− SD are shown. Multiple t-tests with Holm–Šídák correction were used to compare data points of the two curves and p < 0.05 was considered as significant (asterisk). d) TOPFlash assay was used to investigate the influence of vemurafenib on β-catenin /TCF/LEF dependent transcription. Reporter transfected cells were treated for 24 h with the indicated concentrations of vemurafenib before assaying the luciferase activity. Firefly values were normalized to renilla signals and to the corresponding untreated controls. Mean values and standard deviations of sixtuplicates are presented. Multiple t-tests with Holm–Šídák correction were used to compare data of sensitive and resistant samples and p < 0.05 was considered as significant (asterisk). |
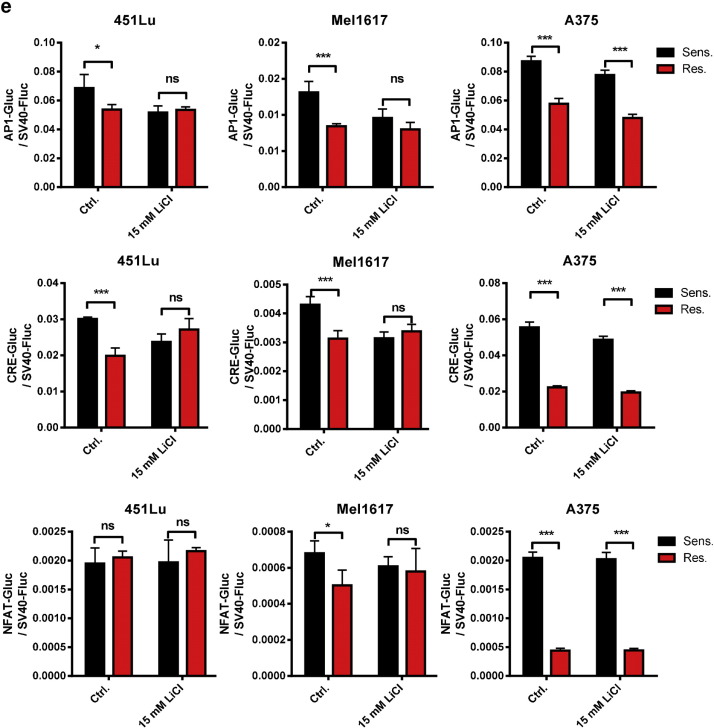
Furthermore, we inhibited protein interactions of β-catenin with TCF/LEF by the treatment with sub-toxic concentrations of the β-catenin antagonist PKF115–584 (Sinnberg et al., 2011 ). Surprisingly, similar to the β-catenin stabilization by lithium chloride, the inhibition of the interaction between β-catenin and TCF/LEF factors enhanced the resistance towards vemurafenib in the sensitive 451Lu cell line which indicates a β-catenin mediated TCF/LEF independent resistance mechanism by the release of β-catenin from this complex. The degree of resistance remained unaltered in the corresponding resistance-acquired cell line using PKF115–584 (Fig. 6 c). We further used the Super8xTopFlash reporter to analyze the effect of BRAFi on the TCF-4/LEF-1 dependent transcriptional activity of β-catenin. We therefore treated sensitive and resistant cells that had been transfected with the reporter construct with increasing concentrations of vemurafenib for 24 h using clinically relevant doses of 1–10 μM. BRAFi treatment reduced the luciferase signal in the sensitive cells in a significant and concentration-dependent manner. This effect was less pronounced in the resistant cell lines (Fig. 6 d). This was in agreement with a rather low binding capacity of β-catenin to TCF-4 after treatment with vemurafenib in a pulldown assay (Supplementary Fig. 4a ). Finally, to evaluate a potential direct effect of Wnt ligands that activate either the canonical Wnt or non-canonical Wnt/Ca2 + signaling pathways on vemurafenib resistance, we treated the sensitive melanoma cell lines 30 min before the addition of vemurafenib with Wnt3a or Wnt5a. Both pre-treatments did not change sensitivity towards the BRAFi in the cells (Supplementary Fig. 4c, d ). In order to further illuminate a possible role of the non-canonical Wnt pathways in our resistant cell lines we used luciferase reporter assays (Ring et al., 2014 ) detecting transcriptional activity of protein kinase A (PKA) signaling (CRE/cAMP responsive element binding sites), Ca2 + /protein kinase C (PKC) signaling (NFAT/nuclear factor of activated T cells binding sites) and jun N-terminal kinase (JNK) signaling (AP-1/activator protein 1 binding sites). No significant correlation between β-catenin expression and reporter activity was seen. Interestingly, resistant cells showed a lower basal non-canonical Wnt activity than sensitive parental cells. This activity was reduced after stabilization of β-catenin with LiCl in sensitive 451Lu and Mel1617 cells but not in resistant cells (Supplementary Fig. 4e ). Together, these data strongly support the hypothesis that stabilized nuclear β-catenin acts at least partially independent of the common TCF-4/LEF-1 dependent canonical Wnt/β-catenin signaling in the mediation of vemurafenib resistance.
3.4. Stat3 Signaling Is Critical for the β-Catenin Mediated Resistance Towards Vemurafenib
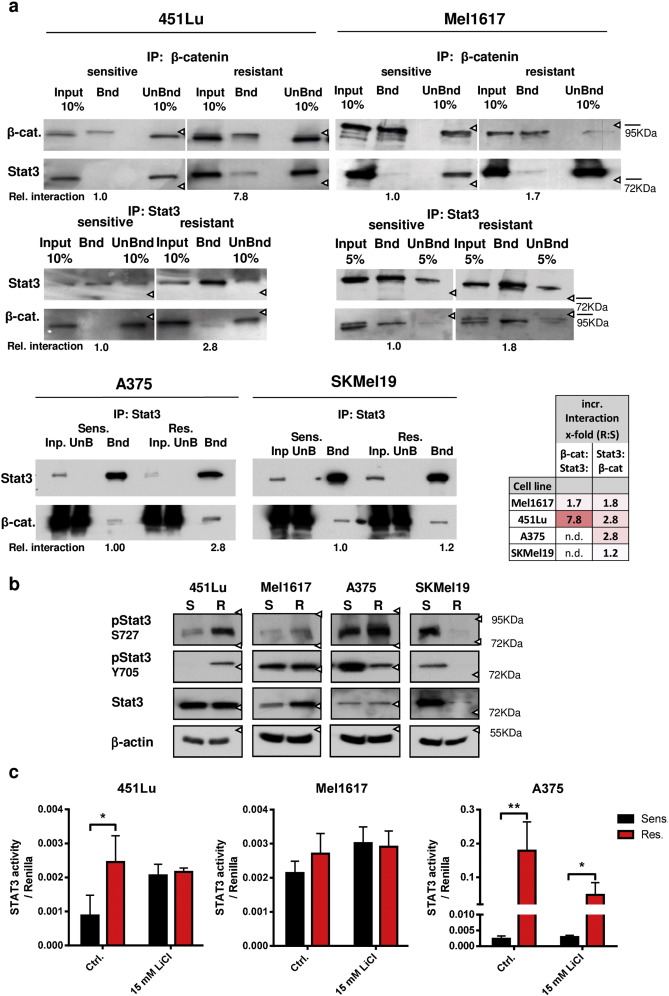
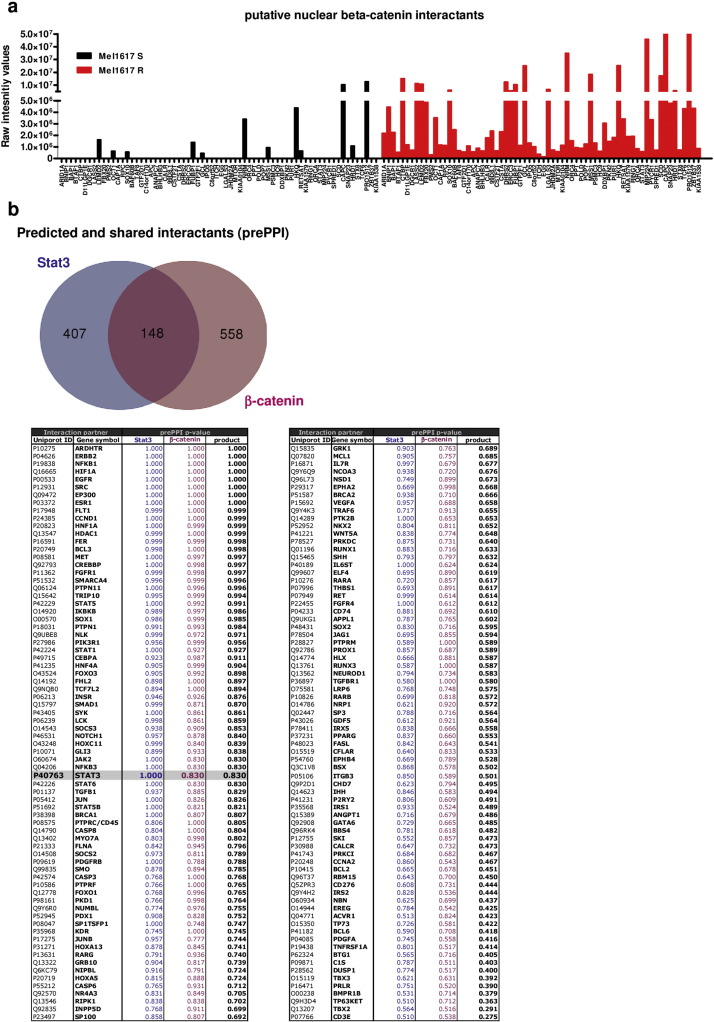
Since we found that resistance mediation of β-catenin to BRAFi is largely independent of the canonical Wnt signaling in melanoma cells we investigated the involvement of other factors. Therefore, we performed co-immunoprecipitation of β-catenin in nuclear extracts of sensitive and resistant Mel1617 cells followed by LC-MS/MS analyses of the interaction partners (Supplementary Table 1 ). Among the top candidates of identified nuclear proteins enriched in the Mel1617-R sample, Stat3 was suspicious as a critical β-catenin interaction with a significantly increased interaction in resistant compared to sensitive cells (Supplementary Fig. 5a , Supplementary Table 2 ). We further performed an in silico analysis of the interactome of β-catenin and Stat3 and found a set of 148 shared interactants including β-catenin and Stat3 itself ( Supplementary Fig. 5b ). We confirmed the increased interaction of β-catenin with Stat3 in resistant compared to sensitive 451Lu and Mel1617 cells by western blot analysis after immunoprecipitation of either of the interaction partners. The interaction could be detected in both directions by using antibodies specific for β-catenin or Stat3 for the precipitation and the reciprocal antibody for immune-detection with increased occurrence in resistant 451Lu and Mel1617 cells (Fig. 7 a). In A375-R cells we also found increased amounts of β-catenin in the anti-Stat3 precipitates confirming the increased interaction in BRAFi resistant cells. In SKMel19-R cells the interaction could be detected but was not increased when compared to SKMel19-S cells (Fig. 7 a). Stat1 could not be detected in immunoprecipitations of 451Lu and Mel1617 lysates using a β-catenin specific antibody (data not shown). In the BRAFi resistant melanoma cell lines 451Lu-R, Mel1617-R and A375-R that comprised the observed nuclear β-catenin stabilization and interaction with Stat3, we found increased phosphorylation of Stat3 at Ser727 and for 451Lu-R additionally at Tyr705 whereas total Stat3 protein levels were not consistently increased in the lysates (Fig. 7 b). Both phosphorylation sites are strong indicators of activated Stat3 signaling. Interestingly, the phospho-Ser727 correlated with β-catenin and phospho-Erk1/2 levels (Fig. 1 b & 7b). Indeed, Stat3 signaling activity was significantly increased in 451Lu-R and A375-R cells as measured by a Stat3 reporter assay (Fig. 7 c) and in Mel1617-R a non-significant trend to increased reporter activity was measured. Resistant cell lines had higher transcript levels of IL-6 which is a target of Stat3 signaling and at the same time activates Stat3 via binding to IL6Rα and gp130 activation ending up in a feedforward loop that further activates Stat3 in the tumor microenvironment ( Supplementary Fig. 1e ). Stabilization of β-catenin using LiCl increased the Stat3 reporter activity in sensitive 451Lu and Mel1617 cells to levels of the corresponding resistant cells (Fig. 7 c).
|
|
|
Fig. 7. Interaction of β-catenin with Stat3 occurs preferentially in BRAFi resistant melanoma cells. a) Co-immunoprecipitation experiments were done to confirm an interaction of β-catenin with Stat3. Lysates were prepared after crosslinking of the proteins with 0.4% buffered formaldehyde using the sensitive and resistant versions of 451Lu and Mel1617 cells. Precipitation was performed with a β-catenin or a Stat3 specific antibody and detected by immunoblot. Semi-quantification by densitometric analysis was done by calculating the ratios (bound fraction:input fraction) followed by normalization to the sensitive cells (see numbers below the blots and table). b) Immunoblot analysis for Stat3 expression and phosphorylation (Ser727 and Tyr705) in the sensitive and resistant 451Lu, Mel1617 and A375 cell lines. The same lysates as in Fig. 1 B were used. Beta-actin protein levels served as loading controls. c) Firefly reporter assay for Stat3 specific transciption was used to detect the nuclear Stat3 signaling activity in the sensitive and resistant melanoma cells of 451Lu, Mel1617 and A375. LiCl (15 mM for 24 h) treatment was used to induce β-catenin accumulation. Firefly signals were normalized to CMV-renilla control signals. Mean values +/− SD of sixtuplicates are shown. Multiple t-tests with Holm–Šídák correction were used to compare data of sensitive and resistant samples and p < 0.05 was considered as significant (asterisk). |
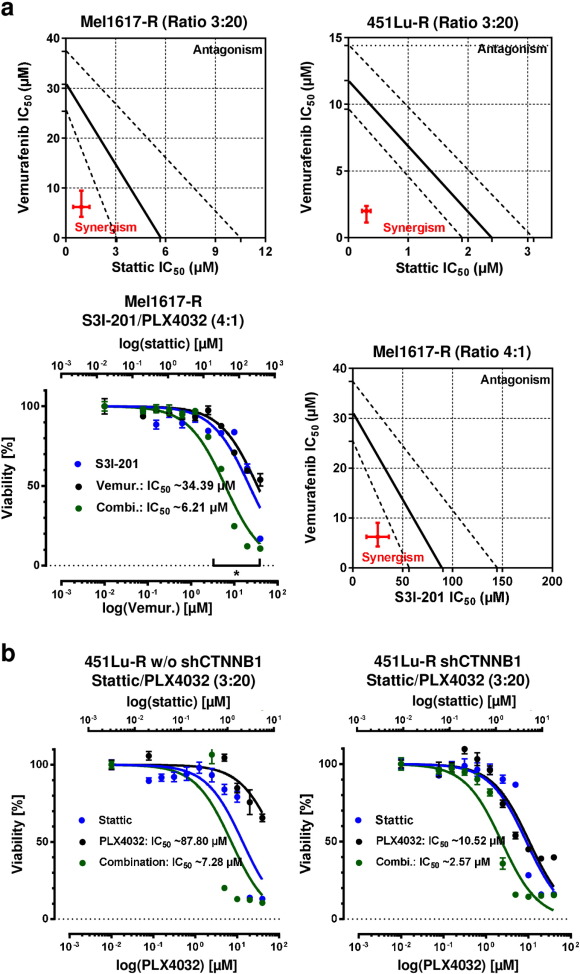
To reveal the functional role of Stat3 in the vemurafenib resistance mechanisms, we overexpressed Stat3 via lentiviral gene transfer in sensitive A375 and SKMel19 cell lines. As expected the increased expression of Stat3 resulted in a decreased sensitivity towards vemurafenib treatment ( Fig. 8 a). Furthermore, treatment with two different inhibitors of Stat3 activation, S3I-201 (Supplementary Fig. 6a ) and Stattic (Fig. 8 b), sensitized the resistance-acquired 451Lu-R, Mel1617-R and to a lesser extent A375-R and SKMel19-R cells towards vemurafenib treatment, yielding in a significantly reduced IC50 in the combination treatment of vemurafenib with Stattic compared to the mono-treatment with vemurafenib (Fig. 8 b). Isobologram analysis evidenced the synergistic character of combined inhibition of Stat3 and BRAFV600E in terms of reduced viability in the Mel1617-R and 451Lu-R BRAFi resistant cells (Supplementary Fig. 6a ). In order to test this combination in a more clinical model we used short term cultured melanoma cells (three models isolated from patient biopsies and one model from a patient derived xenograft mouse) which were derived from clinically BRAFi-resistant BRAFV600E melanomas. A similar beneficial effect of the combination could be measured after three days of treatment in the viability assay (Fig. 8 b).
|
|
|
Fig. 8. Stat3 and β-catenin levels cooperatively mediate resistance to BRAFi in melanoma cells. a) Stat3 was overexpressed in sensitive A375 and SKMel19 cells using a lentivirus (LV-STAT3). After selection the stable overexpression was tested by western blot and the cells were used for cell viability testings (MUH) after 72 h of treatment with vemurafenib. Signals were normalized to the control cells without vemurafenib treatment. Mean +/− SD values of six replicates are shown. Multiple t-tests with Holm–Šídák correction were used to compare data points of the two curves and p < 0.05 was considered as significant (asterisk). F-test revealed a significant different IC50 of the fitted curves. b) Cell viability (MUH assay) 72 h after combinatorial inhibition of Stat3 by Stattic and BRAFV600E by vemurafenib in resistant Mel1617-R, 451Lu-R, A375-R and SKMel19-R cells (top row). Stattic and vemurafenib were mixed in a fixed ratio (3:20) with the maximum concentrations 3 μM for Stattic and 20 μM for vemurafenib. Cells were treated with the inhibitors in ascending concentrations of the combined drugs. 72 h after treatment cell viablitiy was assessed in quintuplicates. The top x-axes represent the concentrations of Stattic and the bottom x-axes represent the concentrations of vemurafenib. Mean +/− SD values are shown. Multiple t-tests with Holm–Šídák correction were used to compare data points of the curves and p < 0.05 was considered as significant (asterisk). F-test revealed a significant different IC50 of the fitted curves. Four short-term melanoma cultures from clinically BRAFi resistant tumors were used for viability testing at 72 h after beginning with either mono or combinatorial treatment using Stattic and vemurafenib (bottom row). c) Cell viability after knockdown of β-catenin. siRNA (50 nM) was used to specifically downregulate either β-catenin or Stat3 in the resistant 451Lu-R, Mel1617-R and A375-R cells. 24 h post transfection cells were treated with increasing concentrations of vemurafenib for 72 h before the measurement of cell viability via MUH assay. Each assay was performed in quintuplicates. Multiple t-tests with Holm–Šídák correction were used to compare data points of the curves and p < 0.05 was considered as significant (asterisk). d) The 451Lu-TetOn-shCTNNB1 resistant cells were used for double (left and middle diagram) and triple targeting (right diagram) of β-catenin (using doxycycline), Stat3 (using Stattic) and BRAFV600E (using vemurafenib). For the knockdown of β-catenin cells were pretreated with 1 μg/ml doxycycline before the addition of the inhibitors. After 72 h of treatment cell viability was assessed. Multiple t-tests with Holm–Šídák correction were used to compare data points of the curves and p < 0.05 was considered as significant (asterisk). Knockdown of β-catenin reduced the IC50 of vemurafenib 3.7fold and the IC50 of Stattic 1.3fold. Combination of the knockdown with both inhibitors resulted in an additive effect as shown in the isobologram. Intersections with the axes denote the IC50 values of the corresponding inhibitors. The measured effective IC50 of the combinations are depicted by the dots. Data points of the assay with knockdown are shown in red, black data points represent the cells without knockdown of β-catenin. |
The sensitizing effect was confirmed by using siRNA against Stat3 combined with vemurafenib treatment in 451Lu-R, Mel1617-R and A375-R cells. The amount of re-sensitization was comparable to a siRNA specific for β-catenin (Fig. 8 c). These results were additionally substantiated with the results obtained using the Tet-inducible shCTNNB1 in combination with the Stat3 inhibitor Stattic or vemurafenib (Fig. 8 d, Supplementary Fig. 6b ). To analyze the role of Stat3 and β-catenin interaction in vemurafenib sensitivity, we treated the Tet-inducible shCTNNB1 451Lu-R cells with Stattic and vemurafenib in a fixed ratio (3:20) and additionally induced a β-catenin knockdown in these cells by a preceding doxyxcycline addition. The knockdown enabled a significant decline of cell viability at lower concentrations of Stattic and vemurafenib in comparison to the control cells without the induction of shCTNNB1 . As before, β-catenin down-regulation reduced vemurafenib resistance and lowered the IC50 for vemurafenib 3.7 fold and the IC50 of Stattic approx. 1.3 fold. In combination with the double treatment (using vemurafenib plus Stattic) the induced shRNA acted in an additive to superadditive manner in the resistance-acquired 451Lu-Tet cells (Fig. 8 d). These data strongly support an important role of β-catenin for the Stat3 mediated resistance towards vemurafenib.
4. Discussion
Acquired resistance to the second generation BRAF inhibitor vemurafenib is limiting the benefits of long term targeted therapy for patients with malignant melanomas that harbor the V600E BRAF mutation (Chapman, 2013 and Wagle et al., 2011 ). Recent studies revealed an enormous heterogeneity of underlying resistant mechanisms within the individual patients but also within single tumors (Hoogstraat et al., 2014 , Rizos et al., 2014 and Shi et al., 2014 ). The selective pressure of BRAFi can force clonal evolution of a variety of different phenotypes. This heterogeneity is at least partly reflected in our cell line models of resistance where we have diverse expression levels of RTKs and different effects on the level of BRAF that cause the reactivation of the MAPK signaling pathway under vemurafenib treatment. This demonstrates the need to identify common signaling nodes which merge several resistance mechanisms and could be used as additional target structures like the recent discovery of the translational regulator complex EIF4F (Boussemart et al., 2014 ) or Pax3/Mitf during the adaptation phase to BRAFi (Smith et al., 2016 ).
Our present results provide additional insight into the complexity of the vemurafenib resistance phenotype of melanomas and advance the basis upon which such resistance may be overcome. The first major finding in this work is the increase of β-catenin protein levels in about half of the melanoma metastases with acquired resistance towards vemurafenib. This suggests that β-catenin plays a role in a subset of melanoma cells with acquired resistance towards BRAFi. In line with the described heterogeneity of resistant melanomas this subset would be ideally suitable for β-catenin targeting. We clearly show that reduced β-catenin levels can strongly enhance the growth inhibitory and apoptotic effects of vemurafenib in a synergistic manner in these cells. Moreover, knocking down of β-catenin levels partially re-sensitized our resistant melanoma cell lines for the BRAF inhibitor vemurafenib. This demonstrates that an increased β-catenin protein level mediates survival of melanoma cells during a therapy resistant state which is in line with a recent finding of Chien et al. (Chien et al., 2014 ), showing a worse survival of BRAFi treated patients with melanomas containing the BRAFV600E mutation and high β-catenin levels compared to patients with tumors having low β-catenin levels. This supports our previous studies demonstrating that β-catenin can positively affect melanoma cell survival (Sinnberg et al., 2010 and Sinnberg et al., 2011 ). Moreover, we show that knocking down of β-catenin delays the development of resistance to vemurafenib under chronic BRAF inhibition, which further strengthens the important role of β-catenin in the formation of acquired resistance to BRAF inhibitors in melanoma. In the light of the role of Mitf during the adaptation of melanoma cells to BRAFi (Smith et al., 2016 ) this makes perfect sense since it is known that β-catenin regulates the expression of Mitf (Widlund et al., 2002 ). However enhanced β-catenin signaling seems not to be generally protective in BRAF mutated melanoma cell lines. Chien and Moon showed enhanced apoptosis induction in some cell lines showing an activated Wnt-/β-catenin signaling after BRAF inhibition (Biechele et al., 2012 ) as well as after MEK inhibition (Conrad et al., 2012 ). However, their results suggest that in melanoma cells exposed to long-term BRAFi, the transcriptional effects of Wnt/β-catenin signaling (measured by AXIN2 transcript levels) are uncoupled from the enhancement of apoptosis by Wnt/β-catenin signaling. A very early activation of the canonical Wnt signaling pathway after 3–6 h of BRAFi treatment could be detected in our cell lines, too (data not shown) which declined below starting levels after 24 h of BRAF inhibition.
In our resistant melanoma cells the elevated levels of β-catenin did not cause an increase in activity of the canonical Wnt signaling pathway although the increase in β-catenin was also detectable on a nuclear level. Target gene expression like MITF (Dorsky et al., 2000 and Widlund et al., 2002 ) and Tyrosinase (Wang et al., 2015 ) was even reduced in the resistant cells. This is in accordance with a previously identified resistant subpopulation of chronically treated melanomas with low MITF expression (Müller et al., 2014 ). Similar results were seen by Chien et al. after chronic treatment of cells with PLX4027 that did not cause increased canonical Wnt signaling activity (Chien et al., 2014 ). Beyond that, activation of Wnt signaling by Wnt3a stimulation did not mediate resistance to our melanoma cells tested and also did not further sensitize the cells to vemurafenib treatment. In contrast, stabilization of β-catenin by lithium chloride as well as the liberation of β-catenin from complexes with TCF4/LEF1 by treatment with PKF115–584 reduced melanoma susceptibility to the BRAF inhibitor vemurafenib. This hints at a Wnt signaling independent function of β-catenin in the course of resistance to BRAF inhibition. Beta-catenin can be also stabilized by non-canonical Wnt ligands like Wnt5a, dependent on the receptor context. While binding of Wnt5a to Ror2 usually inhibits the canonical β-catenin dependent signaling pathway (van Amerongen et al., 2012 ), its binding to frizzled 4/Lrp6 is known to stabilize β-catenin (Grossmann et al., 2013 and Mikels and Nusse, 2006 ). The non-canonical signaling of Wnt5a including the downstream activation of PKC is known to be activated in highly invasive melanomas (Dissanayake et al., 2007 , Hoek et al., 2008 and Weeraratna et al., 2002 ). Moreover Wnt5a was previously found to mediate intrinsic and acquired therapy resistance of melanoma cells to BRAFi by signaling through Ror2 (O'Connell et al., 2013 ). In another recent work Wnt5a expression was shown to be increased in a subset of melanoma cells with activation of Akt mediating acquired resistance to BRAFi (Anastas et al., 2014 ). Interestingly, the receptors frizzled-7 and Ryk were suspected of mediating this effect. In xenopus cells frizzled-7 is also capable to activate β-catenin after binding of Wnt5a (Umbhauer et al., 2000 ), which could be a hint on Wnt5a dependent β-catenin stabilization in therapy resistant melanoma cells. However, although the resistant form of Mel1617 showed increased expression of Wnt5a no tendency towards increased Wnt5a levels was detected in the other resistant cells tested. In addition, Wnt5a stimulation of sensitive melanoma cell lines did not significantly attenuate the efficacy of vemurafenib. Thereby, we exclude a Wnt3a or Wnt5a dependent stabilization of β-catenin in our cell lines with acquired resistance and we further reason that β-catenin has a Wnt independent function in the acquired resistance mechanisms to BRAFi. From previous studies it is known that the interaction partners of β-catenin seem to dictate its functional activities. In melanoma cells interaction of β-catenin with TCF4 drives invasive melanoma growth, whereas interaction with LEF1 predominated in a proliferative phenotype (Eichhoff et al., 2011 ).
Indeed the second major finding of our work is a so far non-described interaction of β-catenin with the signal transducer and activator of transcription 3 (Stat3) in resistant melanoma cells. Our data prove for the first time a physical interaction of the transcription factors β-catenin and Stat3. The levels of β-catenin in melanoma cells seem to change the signaling activity of Stat3 as determined by β-catenin stabilization. This hints on a regulatory function of β-catenin protein levels in Stat3 signaling. In cutaneous melanoma, activation of Stat3 signaling is known to be a negative prognostic factor (Lee et al., 2012 and Wang et al., 2007 ). Our observation that Stat3 overexpression increases vemurafenib resistance and Stat3 knockdown or inhibition increases sensitivity towards vemurafenib is in coincidence with a recent study showing an important role of Stat3 signaling in the mediation of resistance to BRAFi by the induction of the transcription factor Pax3 (Liu et al., 2013 ). In addition, we found an increase of Stat3 phosphorylation at Ser727 in the same cell lines that showed increased nuclear levels of β-catenin. Phosphorylation at Tyr705 was more variable in the resistant cells confirming the finding that phosphorylation of Ser727 can occur independently of Tyr705 in melanocytic cells (Sakaguchi et al., 2012 ). Furthermore Stattic reduced melanoma cell viability together with vemurafenib in a synergistic manner in BRAFi resistant melanoma cell lines and patient derived short-term cultures. Additional knockdown of β-catenin further enhanced this growth inhibitory effect. Interestingly, beyond the direct interaction, Stat3 and β-catenin share interaction partners that are known to be involved in the re-activation of MAPK signaling pathways or the hyperactivation of PI3K/Akt signaling during the acquisition of resistance to BRAFi. Among those are the receptor tyrosine kinases Erbb2/3 and c-Met, that mediate resistance via ligand dependent activation of down-stream signaling adaptor molecules like Src which is also among the common interactants. Nrg-1, a ligand of Erbb2/Erbb3 Egf-receptor complexes was shown to mediate resistance in melanoma cells to BRAFi ( Abel et al., 2013 , Tiwary et al., 2014 and Wilson et al., 2012 ) as well as Hgf, the ligand of c-Met ( Straussman et al., 2012 and Wilson et al., 2012 ). In malignant melanoma these growth factors can be expressed by stromal cells like cancer associated fibroblasts (Straussman et al., 2012 ). The corresponding receptors are capable to activate β-catenin upon ligand binding. A putative mechanism could be the activation of members of the Src kinase family. Interestingly, BRAF and MEK inhibition can directly cause an intermediate phosphorylation of Stat3 via RTK-driven Src activation in a subset of melanomas ( Vultur et al., 2014 ). A second mechanism of Stat3 activation in the course of resistance acquisition towards BRAFi might be the finding of increased expression of IL-6 which activates the Stat3 signaling pathway via its receptor gp130 homodimerization and janus kinase (JAK) activation. Again, JAK1 has gained attention in terms of mediation of primary resistance ( Sos et al., 2014 ) and of secondary resistance to BRAFi (Kim et al., 2015 ).
Our data indicate that β-catenin and Stat3 are both downstream targets of important mediators of resistance acquisition against chronic treatment with BRAFi and converge in a signaling node. During resistance development β-catenin obviously changes its nuclear interaction partners to Stat3. The convergence of resistance mediating pathways at the level of β-catenin and Stat3 by the formation of a novel protein complex turns this interaction into an important nodal point. According to our finding that inhibition of Stat3 and knockdown of β-catenin both synergized with vemurafenib in the resistant melanoma cells as well as in their combination, targeting of this signaling complex seems to be a promising strategy to improve targeted therapies with BRAFi. Further investigation is needed to decipher the importance of the β-catenin Stat3 interaction for the canonical Wnt pathway, the Stat3 signaling and the resistance acquisition to MAPK pathway inhibitors like BRAFi and MEKi. A further in-depth characterization of the activating mechanisms of Stat3 might lead to novel therapeutic target structures that are capable to overcome acquired resistance to BRAF and maybe even MEK inhibition.
The following are the supplementary data related to this article.
Supplementary material
|
|
|
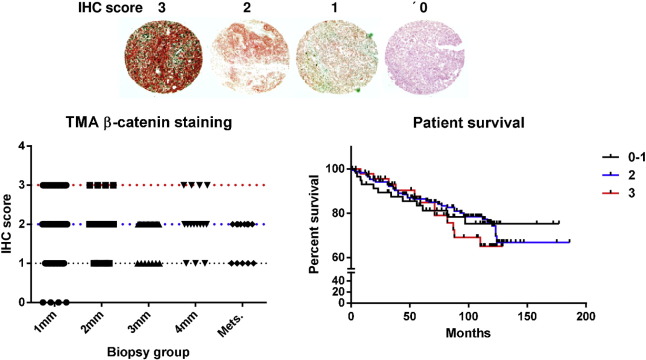
Supplementary Fig. 1. Beta-catenin IHC of a tissue microarray containing 270 melanoma biopsies. The slide was stained using a β-catenin specific antibody (Cell Signaling #9562 1:100). Staining intensities (IHC score) were judged by two experimenters from 0 (= absent) to 3(= strong) and grouped according tumor thickness and metastases. Kaplan Meier analysis was done using primary tumor data. |
|
|
|
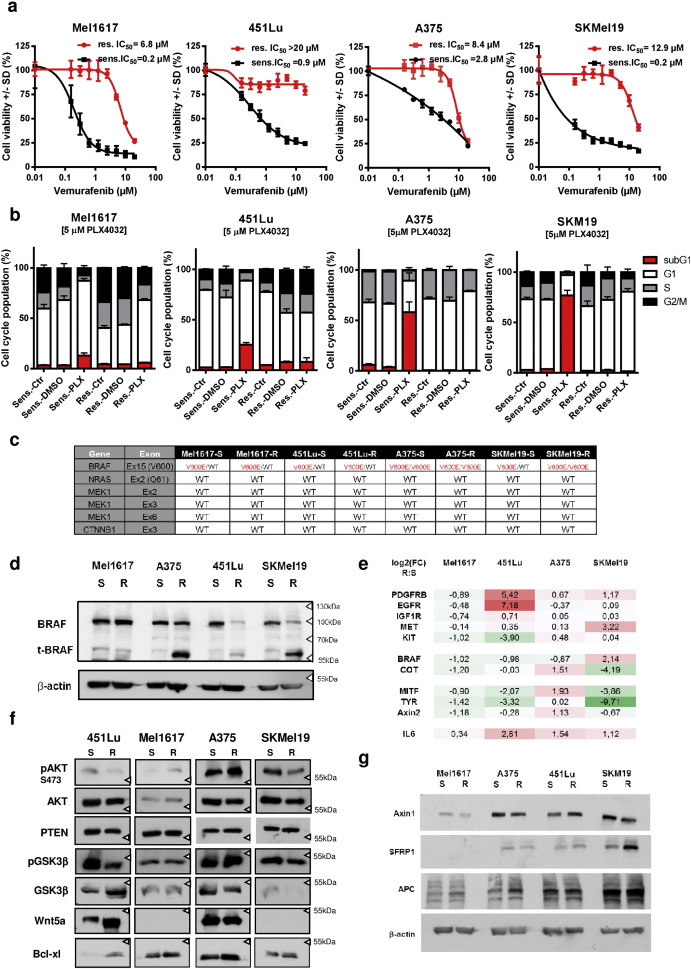
Supplementary Fig. 2. Increased resistance of melanoma cell lines after chronic treatment with 2 μM PLX4032 (vemurafenib). a) Cell viability assays (MUH) were performed after 72 h of treatment with increasing concentrations of vemurafenib. Black data points show the parental sensitive cells, red data points show the corresponding resistant cells. Assays were performed in sixtuplicates and data were normalized to the untreated control, mean values +/− SD are shown with the calculated IC50 values vemurafenib. b) Cell cycle analysis of the sensitive and resistant melanoma cell lines using propidium iodide staining. c) Mutation analysis of hotspot mutations in the four cell line pairs Mel167, 451Lu, A375 and SKMel19 for BRAF, NRAS, CTNNB1 and MEK1. d) Western blot to detect the expression levels of BRAF in the four cell lines together with truncated splice variants of BRAF. e) Transcript expression (real-time qPCR) of genes known to be involved in resistance mechanisms and target genes of Wnt-/β-catenin signaling (MITF and TYR) and Stat3 (IL6). f) Immunoblots corresponding to Fig. 1 with additional signaling proteins. g) Immunoblots showing the expression of regulators of β-catenin expression (APC, Sfrp1, Axin-1) in the four cell line pairs. |
|
|
|
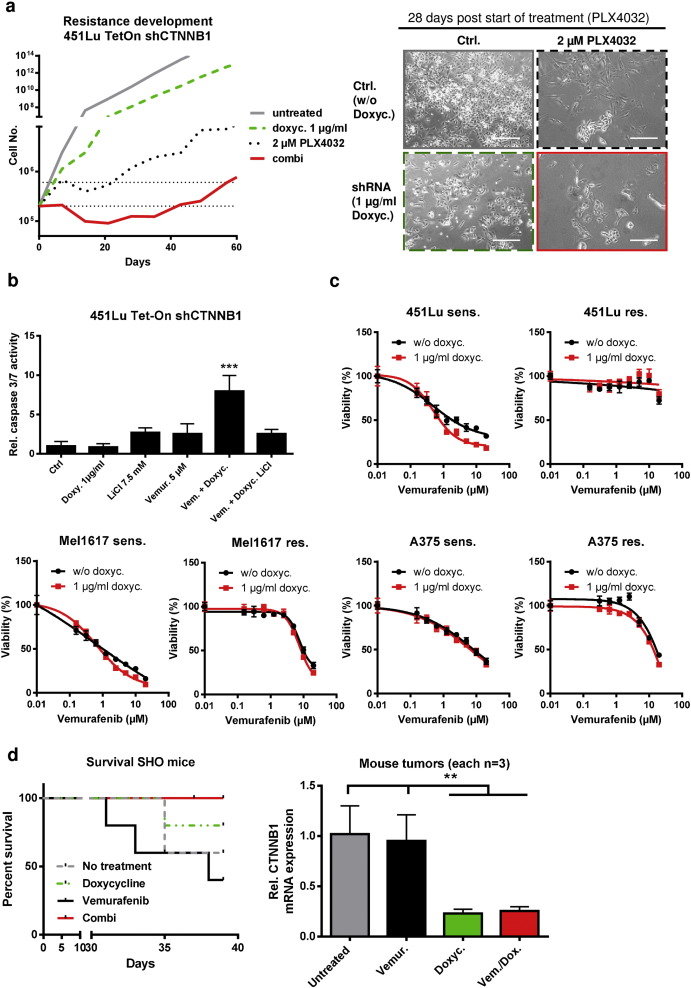
Supplementary Fig. 3. Beta-catenin is critically involved in the development of resistance to BRAFi. a) 451Lu-TetOn-shCTNNB1 cells were used for chronic treatment with 2 μM vemurafenib (1 μM during the first week) either in the presence (green and red curves) or absence (grey and black curves) of doxycyline (1 μg/ml doxycycline) and cell numbers were weekly counted by automated cell counting and viability assessment (CASY). Knockdown of β-catenin increased the time by approx. 2.5 fold until cell numbers exceeded the cell numbers of the maximum cell number in the first week. Morphology of the treated cells is shown after four weeks of cultivation with the indicated four different treatments (scale bar is 200 μm). b) The significantly (p < 0.001) increased caspase3/7 activation (DEVD-AMC as caspase 3/7 substrate) after 48 h post knockdown of β-catenin with combined BRAF inhibition can be blocked by β-catenin stablilization using a 6 hour pre-incubation with 7.5 mM LiCl. c) Control experiment to show that doxycyline treatment (1 μg/ml) without shCTNNB1 induction has no impact at the susceptibility of the indicated sensitive and resistant melanoma cell lines to increasing concentrations of vemurafenib using the MUH cell viability assay. Cells were treated for 72 h with vemurafenib after a pre-treatment for 24 h with the antibiotic. d) Kaplan–Meier-survival curves (left panel) showing the survival data of the SHO mice from Fig. 2 . Black curve: vemurafenib 25 mg/kg i.p. treated mice (n = 5), grey curve: untreated mice (n = 5), green curve: doxycycline 1 mg/ml p.o. (n = 5), red curve: doxycycline 1 mg/ml p.o. plus vemurafenib 25 mg/kg i.p. (n = 5). Bar diagram (right panel) shows the significant extent of CTNNB1 down-regulation after in vivo shRNA induction with 1 μg/ml doxycycline in the drinking water of SHO mice compared to untreated or vemurafenib (mono-therapy) treated mice (p < 0.01; each n = 3). mRNA expression was measured using SYBR-green qPCR. |
Supplementary Fig. 4.
Wnt signaling is not majorly involved in resistance mechanisms to BRAFi.
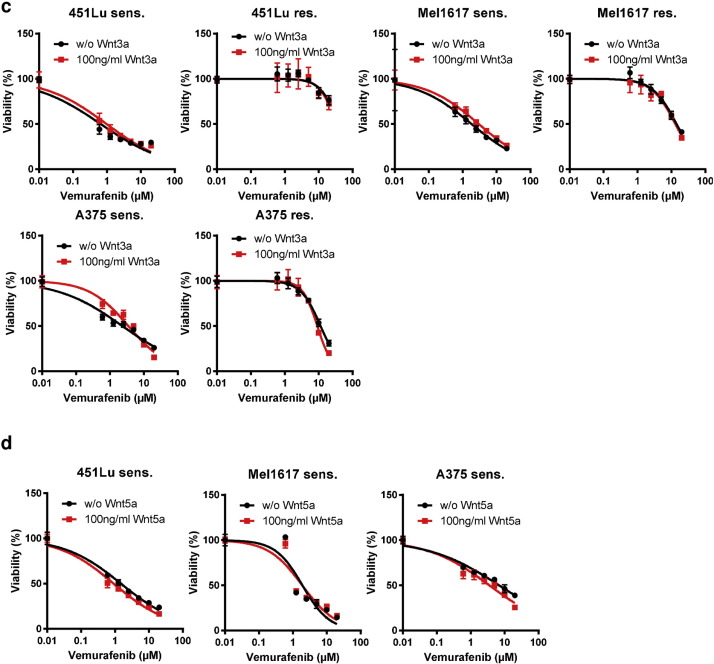
a) Bead-based GST-pull-down assay of “free” β-catenin using GST-tagged TCF4. Samples were prepared in triplicates after cultivation under the indicated conditions for 24 h. Signals were normalized to the total amount of β-catenin measured by a bead-based sandwich immunoassay and compared to the untreated controls. b) Heatmap of mRNA expression analysis using the Illumina beadchip-based microarray dataset GSE50509 dealing with 21 pairs of samples from melanoma patients treated with BRAFi33 . Fold changes in gene expression of BRAFi treated and matched pre-treatment tumors were calculated for each patient using log2 transformed and RMA normalized signal intensities. Exemplarily the gene sets M223 (canonical Wnt signaling) and M31 (non-canonical Wnt signaling) from the Gene set enrichment analysis (GSEA) tool from the Broad Institute is shown which shows nuclear β-catenin signaling and target gene transcription.
c) Cell viability assay of cells pre-treated with Wnt3a (100 ng/ml) before treatment with vemurafenib for 72 h reveals no effect of Wnt3a stimulation. After the pre-treatment, cells were re-stimulated with fresh Wnt3a when the BRAFi treatment was started. Black data represent unstimulated cells, red data show Wnt3a treated cells. d) Sensitive 451Lu-S, Mel1617-S and A375-S cells were pre-treated with Wnt5a (100ng/ml) and re-stimulated after 24 h. Vemurafenib treatment was applied for 72 h before measurement of the cell viability (MUH). No differences in susceptibility to BRAFi were found after treatment with Wnt3a or Wnt5a. e) Three different Gluc-reporter systems (AP1-Gluc, CRE-Gluc, NFAT-Gluc) were used to assay the non-canonical Wnt signaling activity in the sensitive and resistant versions of 451Lu, Mel1617 and A375 melanoma cells. After transfection with the reporter plasmids cells were treated for 24 h with 15 mM LiCl for stabilizing β-catenin. Secreted gaussia luciferase activity was measured and normalized to cytoplasmic firefly luciferase activity from a co-transfected SV40-fluc control plasmid. Assays were performed in sixtuplicates. Mean values and standard deviations of sixtuplicates are presented. Multiple t-tests with Holm–Šídák correction were used to compare data of sensitive and resistant samples and p < 0.05 was considered as significant (asterisk).
|
|
|
Supplementary Fig. 5. Novel Interaction partners of β-catenin in BRAFi resistant melanoma cells. a) Mel1617-S and -R cells were used for preparing nuclear enriched lysates in RIPA buffer before performing an immunoprecipitation with 500 μg of lysate and β-catenin specific antibody. LC-MS/MS analysis was performed using the co-precipitated and trypsin digested proteins. In Mel1617-R cells highly enriched and nuclear localized candidates for interaction with β-catenin were selected using DAVID and the raw intensity values plotted to compare the interaction in sensitive (black) versus resistant (red) Mel1617 melanoma cells. b) An in silico interactome analysis of β-catenin and Stat3 interaction partners was carried out using prePPI58 . 148 putative shared interaction partners of β-catenin and Stat3 (Venn diagram) were found with a (prePPI-) p-value > 0.5 and sorted according their p-value products. A direct interaction of β-catenin with Stat3 was calculated with the probability p = 0.83. |
|
|
|
Supplementary Fig. 6. Isobolograms and dose response curves of BRAFi resistant melanoma cell lines with combined Stat3 and BRAF inhibition. a) Isobologram analysis for Mel1617-R and 451Lu-R cells using the data of Fig. 7 b. Calculated IC50 values including their 95% confidence interval (red symbols) were used for synergism analysis and isobologram plotting. Dose response curves of Mel1617-R cells treated with the Stat3 inhibitor S3I-201 (blue curve), vemurafenib (black curves) and the combination (green curves) are shown in the diagram at the lower left. Cells were treated with the inhibitors in ascending concentrations using the indicated STAT3i to BRAFi ratio (4:1) of the combined drugs. 72 h after treatment cell viablitiy was assessed in quintuplicates. The top x-axes represent the concentrations of S3I-201 and the bottom x-axes represent the concentrations of vemurafenib. Mean +/− SD values are shown. Multiple t-tests with Holm–Šídák correction were used to compare data points of the curves and p < 0.05 was considered as significant (asterisk). F-test revealed a significant different IC50 of the fitted curves. b) Diagrams show dose response curves of 451Lu Tet-R cells doubly treated with Stattic (upper x-axes and blue data points) and vemurafenib (lower x-axes and black data points). Left diagram shows response on viability (MUH) after three days of treatment without knockdown of β-catenin and right diagram shows the response after additional induction of shCTNNB1. The diagrams correspond to the data presented in Fig. 8 d. |
Excel Table 1.
Identified proteins from the LC-MS/MS analysis using co-immunoprecipitates of β-catenin from nuclear enriched Mel1617-S and –R protein fractions.
Excel Table 2.
Identified proteins from Table 2 with known nuclear localization (analyzed using DAVID) and increased signal (> 5fold) in the Mel1617-R compared to the Mel1617-S sample.
Excel Table 3.
Oligonucleotides used as primers for qPCR and Sanger sequencing.
Contributions
T.S. and E.M. designed, performed and analyzed most of the experiments. T.S. wrote the manuscript. M.K. performed PET scans and together with T.S. and E.M. performed mice experiments. U.R., O.P. and N.G. assisted with protein interaction studies. A.V. and B.M. performed mass spectrometry. S.C. analyzed transcriptome data. H.N. and F.M. assisted at specimen collection and data collection. K.I. provided the tissue microarray. T.S., C.G. and B.S. conceived and oversaw the project. All authors discussed the results and commented on the manuscript.
Competing financial interest
The authors declare no competing financial interests.
Acknowledgments
We thank Marc Van de Wetering for giving us the Tet inducible shCTNNB1 plasmid, Leda Raptis with James Turkson for the Stat3 reporter, Larissa Ring with Alexander Faussner for the non-canonical Wnt signaling reporters and Jessie Villanueva with Meenhard Herlyn for the initial assistance with the generation of resistant melanoma cells. This work was supported by the German Cancer Aid (110-210 and the German Melanoma Research Network), the intramural fortuene program (2198-0-0 & 2291-0-0 ), the Deutsche Forschungsgemeinschaft (SFB773 ) and the Open Access Publishing Fund of the University of Tübingen .
References
- Abel et al., 2013 E.V. Abel, K.J. Basile, C.H. Kugel, A.K. Witkiewicz, K. Le, R.K. Amaravadi, G.C. Karakousis, X. Xu, W. Xu, L.M. Schuchter, J.B. Lee, A. Ertel, P. Fortina, A.E. Aplin; Melanoma adapts to RAF/MEK inhibitors through FOXD3-mediated upregulation of ERBB3; J. Clin. Invest., 123 (2013), pp. 2155–2168 http://dx.doi.org/10.1172/JCI65780
- Albert and Silvia, 2012 B. Albert, V. Silvia; The nuclear factor κB inhibitor (E)-2-fluoro-4′-methoxystilbene inhibits firefly luciferase; (2012)
- Anastas et al., 2014 J.N. Anastas, R.M. Kulikauskas, T. Tamir, H. Rizos, G.V. Long, E.M. von Euw, P.-T. Yang, H.-W. Chen, L. Haydu, R.A. Toroni, O.M. Lucero, A.J. Chien, R.T. Moon; WNT5A enhances resistance of melanoma cells to targeted BRAF inhibitors; J. Clin. Invest., 124 (2014), pp. 2877–2890 http://dx.doi.org/10.1172/JCI70156
- Benham-Pyle et al., 2015 B.W. Benham-Pyle, B.L. Pruitt, W.J. Nelson; Mechanical strain induces E-cadherin-dependent Yap1 and -catenin activation to drive cell cycle entry; Science, 348 (80-.) (2015), pp. 1024–1027 http://dx.doi.org/10.1126/science.aaa4559
- Biechele et al., 2012 T.L. Biechele, R.M. Kulikauskas, R.A. Toroni, O.M. Lucero, R.D. Swift, R.G. James, N.C. Robin, D.W. Dawson, R.T. Moon, A.J. Chien; Wnt/β-catenin signaling and AXIN1 regulate apoptosis triggered by inhibition of the mutant kinase BRAFV600E in human melanoma; Sci. Signal., 5 (2012), p. ra3 http://dx.doi.org/10.1126/scisignal.2002274
- Boussemart et al., 2014 L. Boussemart, H. Malka-Mahieu, I. Girault, D. Allard, O. Hemmingsson, G. Tomasic, M. Thomas, C. Basmadjian, N. Ribeiro, F. Thuaud, C. Mateus, E. Routier, N. Kamsu-Kom, S. Agoussi, A.M. Eggermont, L. Désaubry, C. Robert, S. Vagner; eIF4F is a nexus of resistance to anti-BRAF and anti-MEK cancer therapies; Nature, 513 (2014), pp. 105–109 http://dx.doi.org/10.1038/nature13572
- Carlino et al., 2013 M.S. Carlino, K. Gowrishankar, C.A.B. Saunders, G.M. Pupo, S. Snoyman, X.D. Zhang, R. Saw, T.M. Becker, R.F. Kefford, G.V. Long, H. Rizos; Antiproliferative effects of continued mitogen-activated protein kinase pathway inhibition following acquired resistance to BRAF and/or MEK inhibition in melanoma; Mol. Cancer Ther., 12 (2013), pp. 1332–1342 http://dx.doi.org/10.1158/1535-7163.MCT-13-0011
- Chapman, 2013 P.B. Chapman; Mechanisms of resistance to RAF inhibition in melanomas harboring a BRAF mutation; Am. Soc. Clin. Oncol. Educ. Book (2013) http://dx.doi.org/10.1200/EdBook_AM.2013.33.e80
- Chapman et al., 2011 P.B. Chapman, A. Hauschild, C. Robert, J.B. Haanen, P. Ascierto, J. Larkin, R. Dummer, C. Garbe, A. Testori, M. Maio, D. Hogg, P. Lorigan, C. Lebbe, T. Jouary, D. Schadendorf, A. Ribas, S.J. O'Day, J.A. Sosman, J.M. Kirkwood, A.M.M. Eggermont, B. Dreno, K. Nolop, J. Li, B. Nelson, J. Hou, R.J. Lee, K.T. Flaherty, G.A. McArthur; Improved survival with vemurafenib in melanoma with BRAF V600E mutation; N. Engl. J. Med., 364 (2011), pp. 2507–2516 http://dx.doi.org/10.1056/NEJMoa1103782
- Chapman et al., 2012 P.B. Chapman, A. Hauschild, C. Robert, J.M.G. Larkin, J.B.A.G. Haanen, A. Ribas, D. Hogg, O. Hamid, P.A. Ascierto, A. Testori, P. Lorigan, R. Dummer, J.A. Sosman, C. Garbe, M. Maio, K.B. Nolop, B.J. Nelson, A.K. Joe, K.T. Flaherty, G.A. McArthur; Updated overall survival (OS) results for BRIM-3, a phase III randomized, open-label, multicenter trial comparing BRAF inhibitor vemurafenib (vem) with dacarbazine (DTIC) in previously untreated patients with BRAFV600E-mutated melanoma; ASCO Meet. Abstr., 30 (2012) (8502–)
- Chien et al., 2014 A.J. Chien, L.E. Haydu, T.L. Biechele, R.M. Kulikauskas, H. Rizos, R.F. Kefford, R.A. Scolyer, R.T. Moon, G.V. Long; Targeted BRAF inhibition impacts survival in melanoma patients with high levels of Wnt/β-catenin signaling; PLoS One, 9 (2014), Article e94748 http://dx.doi.org/10.1371/journal.pone.0094748
- Conrad et al., 2012 W.H. Conrad, R.D. Swift, T.L. Biechele, R.M. Kulikauskas, R.T. Moon, A.J. Chien; Regulating the response to targeted MEK inhibition in melanoma: enhancing apoptosis in NRAS- and BRAF-mutant melanoma cells with Wnt/β-catenin activation; Cell Cycle, 11 (2012), pp. 3724–3730 http://dx.doi.org/10.4161/cc.21645
- Davies et al., 2002 H. Davies, G.R. Bignell, C. Cox, P. Stephens, S. Edkins, S. Clegg, J. Teague, H. Woffendin, M.J. Garnett, W. Bottomley, N. Davis, E. Dicks, R. Ewing, Y. Floyd, K. Gray, S. Hall, R. Hawes, J. Hughes, V. Kosmidou, A. Menzies, C. Mould, A. Parker, C. Stevens, S. Watt, S. Hooper, R. Wilson, H. Jayatilake, B.A. Gusterson, C. Cooper, J. Shipley, D. Hargrave, K. Pritchard-Jones, N. Maitland, G. Chenevix-Trench, G.J. Riggins, D.D. Bigner, G. Palmieri, A. Cossu, A. Flanagan, A. Nicholson, J.W.C. Ho, S.Y. Leung, S.T. Yuen, B.L. Weber, H.F. Seigler, T.L. Darrow, H. Paterson, R. Marais, C.J. Marshall, R. Wooster, M.R. Stratton, P.A. Futreal; Mutations of the BRAF gene in human cancer; Nature, 417 (2002), pp. 949–954 http://dx.doi.org/10.1038/nature00766
- Desbois-Mouthon et al., 2001 C. Desbois-Mouthon, A. Cadoret, M.J. Blivet-Van Eggelpoël, F. Bertrand, G. Cherqui, C. Perret, J. Capeau; Insulin and IGF-1 stimulate the beta-catenin pathway through two signalling cascades involving GSK-3beta inhibition and Ras activation; Oncogene, 20 (2001), pp. 252–259 http://dx.doi.org/10.1038/sj.onc.1204064
- Dissanayake et al., 2007 S.K. Dissanayake, M. Wade, C.E. Johnson, M.P. O'Connell, P.D. Leotlela, A.D. French, K.V. Shah, K.J. Hewitt, D.T. Rosenthal, F.E. Indig, Y. Jiang, B.J. Nickoloff, D.D. Taub, J.M. Trent, R.T. Moon, M. Bittner, A.T. Weeraratna; The Wnt5A/protein kinase C pathway mediates motility in melanoma cells via the inhibition of metastasis suppressors and initiation of an epithelial to mesenchymal transition; J. Biol. Chem., 282 (2007), pp. 17259–17271 http://dx.doi.org/10.1074/jbc.M700075200
- Dorsky et al., 2000 R.I. Dorsky, D.W. Raible, R.T. Moon; Direct regulation of nacre, a zebrafish MITF homolog required for pigment cell formation, by the Wnt pathway; Genes Dev., 14 (2000), pp. 158–162
- Eichhoff et al., 2011 O.M. Eichhoff, A. Weeraratna, M.C. Zipser, L. Denat, D.S. Widmer, M. Xu, L. Kriegl, T. Kirchner, L. Larue, R. Dummer, K.S. Hoek; Differential LEF1 and TCF4 expression is involved in melanoma cell phenotype switching; Pigment Cell Melanoma Res. (2011) http://dx.doi.org/10.1111/j.1755-148X.2011.00871.x
- Flaherty et al., 2010 K.T. Flaherty, I. Puzanov, K.B. Kim, A. Ribas, G.A. McArthur, J.A. Sosman, P.J. O'Dwyer, R.J. Lee, J.F. Grippo, K. Nolop, P.B. Chapman; Inhibition of mutated, activated BRAF in metastatic melanoma; N. Engl. J. Med., 363 (2010), pp. 809–819 http://dx.doi.org/10.1056/NEJMoa1002011
- Girotti et al., 2013 M.R. Girotti, M. Pedersen, B. Sanchez-Laorden, A. Viros, S. Turajlic, D. Niculescu-Duvaz, A. Zambon, J. Sinclair, A. Hayes, M. Gore, P. Lorigan, C. Springer, J. Larkin, C. Jorgensen, R. Marais; Inhibiting EGF receptor or SRC family kinase signaling overcomes BRAF inhibitor resistance in melanoma; Cancer Discov., 3 (2013), pp. 158–167 http://dx.doi.org/10.1158/2159-8290.CD-12-0386
- Grossmann et al., 2013 A.H. Grossmann, J.H. Yoo, J. Clancy, L.K. Sorensen, A. Sedgwick, Z. Tong, K. Ostanin, A. Rogers, K.F. Grossmann, S.R. Tripp, K.R. Thomas, C. D'Souza-Schorey, S.J. Odelberg, D.Y. Li; The small GTPase ARF6 stimulates β-catenin transcriptional activity during WNT5A-mediated melanoma invasion and metastasis; Sci. Signal., 6 (2013), p. ra14 http://dx.doi.org/10.1126/scisignal.2003398
- Hauschild et al., 2012 A. Hauschild, J.J. Grob, L.V. Demidov, T. Jouary, R. Gutzmer, M. Millward, P. Rutkowski, C.U. Blank, W.H. Miller, E. Kaempgen, S. Martín-Algarra, B. Karaszewska, C. Mauch, V. Chiarion-Sileni, A.M. Martin, S. Swann, P. Haney, B. Mirakhur, M.E. Guckert, V. Goodman, P.B. Chapman; Dabrafenib in BRAF-mutated metastatic melanoma: a multicentre, open-label, phase 3 randomised controlled trial; Lancet, 380 (2012), pp. 358–365 http://dx.doi.org/10.1016/S0140-6736(12)60868-X
- Hoek et al., 2008 K.S. Hoek, O.M. Eichhoff, N.C. Schlegel, U. Döbbeling, N. Kobert, L. Schaerer, S. Hemmi, R. Dummer; In vivo switching of human melanoma cells between proliferative and invasive states; Cancer Res., 68 (2008), pp. 650–656 http://dx.doi.org/10.1158/0008-5472.CAN-07-2491
- Holderfield et al., 2014 M. Holderfield, M.M. Deuker, F. McCormick, M. McMahon; Targeting RAF kinases for cancer therapy: BRAF-mutated melanoma and beyond; Nat. Rev. Cancer, 14 (2014), pp. 455–467 http://dx.doi.org/10.1038/nrc3760
- Hoogstraat et al., 2014 M. Hoogstraat, C.G. Gadellaa-van Hooijdonk, I. Ubink, N.J.M. Besselink, M. Pieterse, W. Veldhuis, M. van Stralen, E.F.J. Meijer, S.M. Willems, M.A. Hadders, T. Kuilman, O. Krijgsman, D.S. Peeper, M.J. Koudijs, E. Cuppen, E.E. Voest, M.P. Lolkema; Detailed imaging and genetic analysis reveal a secondary BRAF(L) (505H) resistance mutation and extensive intrapatient heterogeneity in metastatic BRAF mutant melanoma patients treated with vemurafenib; Pigment Cell Melanoma Res. (2014) http://dx.doi.org/10.1111/pcmr.12347
- Ji et al., 2009 H. Ji, J. Wang, H. Nika, D. Hawke, S. Keezer, Q. Ge, B. Fang, X. Fang, D. Fang, D.W. Litchfield, K. Aldape, Z. Lu; EGF-induced ERK activation promotes CK2-mediated disassociation of alpha-Catenin from beta-Catenin and transactivation of beta-Catenin; Mol. Cell, 36 (2009), pp. 547–559 http://dx.doi.org/10.1016/j.molcel.2009.09.034
- Johnson et al., 2014 D.B. Johnson, K.T. Flaherty, J.S. Weber, J.R. Infante, K.B. Kim, R.F. Kefford, O. Hamid, L. Schuchter, J. Cebon, W.H. Sharfman, R.R. McWilliams, M. Sznol, D.P. Lawrence, G.T. Gibney, H.A. Burris, G.S. Falchook, A. Algazi, K. Lewis, G.V. Long, K. Patel, N. Ibrahim, P. Sun, S. Little, E. Cunningham, J.A. Sosman, A. Daud, R. Gonzalez; Combined BRAF (Dabrafenib) and MEK inhibition (Trametinib) in patients with BRAFV600-mutant melanoma experiencing progression with single-agent BRAF inhibitor; J. Clin. Oncol., 32 (2014), pp. 3697–3704 http://dx.doi.org/10.1200/JCO.2014.57.3535
- Kendziorra et al., 2011 E. Kendziorra, K. Ahlborn, M. Spitzner, M. Rave-Fränk, G. Emons, J. Gaedcke, F. Kramer, H.A. Wolff, H. Becker, T. Beissbarth, R. Ebner, B.M. Ghadimi, T. Pukrop, T. Ried, M. Grade; Silencing of the Wnt transcription factor TCF4 sensitizes colorectal cancer cells to (chemo-) radiotherapy; Carcinogenesis, 32 (2011), pp. 1824–1831 http://dx.doi.org/10.1093/carcin/bgr222
- Kim et al., 2015 H. Kim, D.T. Frederick, M.P. Levesque, Z.A. Cooper, Y. Feng, C. Krepler, L. Brill, Y. Samuels, N.K. Hayward, A. Perlina, A. Piris, T. Zhang, R. Halaban, M.M. Herlyn, K.M. Brown, J.A. Wargo, R. Dummer, K.T. Flaherty, Z.A. Ronai; Downregulation of the ubiquitin ligase RNF125 underlies resistance of melanoma cells to BRAF inhibitors via JAK1 deregulation; Cell Rep., 11 (2015), pp. 1458–1473 http://dx.doi.org/10.1016/j.celrep.2015.04.049
- Kumar et al., 2003 R. Kumar, S. Angelini, K. Czene, I. Sauroja, M. Hahka-Kemppinen, S. Pyrhönen, K. Hemminki; BRAF mutations in metastatic melanoma: a possible association with clinical outcome; Clin. Cancer Res., 9 (2003), pp. 3362–3368
- Lee et al., 2012 I. Lee, P.S. Fox, S.D. Ferguson, R. Bassett, L.-Y. Kong, C.W. Schacherer, J.E. Gershenwald, E.A. Grimm, G.N. Fuller, A.B. Heimberger; The expression of p-STAT3 in stage IV melanoma: risk of CNS metastasis and survival; Oncotarget, 3 (2012), pp. 336–344
- Lim et al., 2009 J.C. Lim, Z. Mickute, M. Zaman, S. Hopkins, H. Wijesuriya, T. Steckler, D. Moechars, F. Van Leuven, Z. Sarnyai, S.B. Hladky, M.A. Barrand; Decreased expression of multidrug efflux transporters in the brains of GSK-3beta transgenic mice; Brain Res., 1276 (2009), pp. 1–10 http://dx.doi.org/10.1016/j.brainres.2009.04.031
- Liu et al., 2013 F. Liu, J. Cao, J. Wu, K. Sullivan, J. Shen, B. Ryu, Z. Xu, W. Wei, R. Cui; Stat3-targeted therapies overcome the acquired resistance to vemurafenib in melanomas; J. Investig. Dermatol., 133 (2013), pp. 2041–2049 http://dx.doi.org/10.1038/jid.2013.32
- Long et al., 2011 G.V. Long, A.M. Menzies, A.M. Nagrial, L.E. Haydu, A.L. Hamilton, G.J. Mann, T.M. Hughes, J.F. Thompson, R.A. Scolyer, R.F. Kefford; Prognostic and clinicopathologic associations of oncogenic BRAF in metastatic melanoma; J. Clin. Oncol., 29 (2011), pp. 1239–1246 http://dx.doi.org/10.1200/JCO.2010.32.4327
- Long et al., 2014 G.V. Long, D. Stroyakovskiy, H. Gogas, E. Levchenko, F. de Braud, J. Larkin, C. Garbe, T. Jouary, A. Hauschild, J.J. Grob, V.C. Sileni, C. Lebbe, M. Mandalà, M. Millward, A. Arance, I. Bondarenko, J.B.A.G. Haanen, J. Hansson, J. Utikal, V. Ferraresi, N. Kovalenko, P. Mohr, V. Probachai, D. Schadendorf, P. Nathan, C. Robert, A. Ribas, D.J. DeMarini, J.G. Irani, M. Casey, D. Ouellet, A.-M. Martin, N. Le, K. Patel, K. Flaherty; Combined BRAF and MEK inhibition versus BRAF inhibition alone in melanoma; N. Engl. J. Med., 371 (2014) http://dx.doi.org/10.1056/NEJMoa1406037 (140929070023009)
- Luckert et al., 2012 K. Luckert, T.S. Gujral, M. Chan, M. Sevecka, T.O. Joos, P.K. Sorger, G. MacBeath, O. Potz; A dual array-based approach to assess the abundance and posttranslational modification state of signaling proteins; Sci. Signal., 5 (2012), p. pl1-pl1 http://dx.doi.org/10.1126/scisignal.2002372
- Menzies and Long, 2014 A.M. Menzies, G.V. Long; Systemic treatment for BRAF-mutant melanoma: where do we go next?; Lancet Oncol., 15 (2014), pp. e371–e381 http://dx.doi.org/10.1016/S1470-2045(14)70072-5
- Mikels and Nusse, 2006 A.J. Mikels, R. Nusse; Purified Wnt5a protein activates or inhibits beta-catenin-TCF signaling depending on receptor context; PLoS Biol., 4 (2006), Article e115 http://dx.doi.org/10.1371/journal.pbio.0040115
- Müller et al., 2014 J. Müller, O. Krijgsman, J. Tsoi, L. Robert, W. Hugo, C. Song, X. Kong, P.A. Possik, P.D.M. Cornelissen-Steijger, M.H.G. Foppen, K. Kemper, C.R. Goding, U. McDermott, C. Blank, J. Haanen, T.G. Graeber, A. Ribas, R.S. Lo, D.S. Peeper; Low MITF/AXL ratio predicts early resistance to multiple targeted drugs in melanoma; Nat. Commun., 5 (2014), p. 5712 http://dx.doi.org/10.1038/ncomms6712
- Nazarian et al., 2010 R. Nazarian, H. Shi, Q. Wang, X. Kong, R.C. Koya, H. Lee, Z. Chen, M.-K. Lee, N. Attar, H. Sazegar, T. Chodon, S.F. Nelson, G. McArthur, J.A. Sosman, A. Ribas, R.S. Lo; Melanomas acquire resistance to B-RAF(V600E) inhibition by RTK or N-RAS upregulation; Nature, 468 (2010), pp. 973–977 http://dx.doi.org/10.1038/nature09626
- O'Connell et al., 2013 M.P. O'Connell, K. Marchbank, M.R. Webster, A.A. Valiga, A. Kaur, A. Vultur, L. Li, M. Herlyn, J. Villanueva, Q. Liu, X. Yin, S. Widura, J. Nelson, N. Ruiz, T.C. Camilli, F.E. Indig, K.T. Flaherty, J.A. Wargo, D.T. Frederick, Z.A. Cooper, S. Nair, R.K. Amaravadi, L.M. Schuchter, G.C. Karakousis, W. Xu, X. Xu, A.T. Weeraratna; Hypoxia induces phenotypic plasticity and therapy resistance in melanoma via the tyrosine kinase receptors ROR1 and ROR2; Cancer Discov., 3 (2013), pp. 1378–1393 http://dx.doi.org/10.1158/2159-8290.CD-13-0005
- Purcell et al., 2011 R. Purcell, M. Childs, R. Maibach, C. Miles, C. Turner, A. Zimmermann, M. Sullivan; HGF/c-Met related activation of β-catenin in hepatoblastoma; J. Exp. Clin. Cancer Res., 30 (2011), p. 96 http://dx.doi.org/10.1186/1756-9966-30-96
- Ring et al., 2014 L. Ring, P. Neth, C. Weber, S. Steffens, A. Faussner; β-Catenin-dependent pathway activation by both promiscuous “canonical” WNT3a-, and specific “noncanonical” WNT4- and WNT5a-FZD receptor combinations with strong differences in LRP5 and LRP6 dependency; Cell. Signal., 26 (2014), pp. 260–267 http://dx.doi.org/10.1016/j.cellsig.2013.11.021
- Rizos et al., 2014 H. Rizos, A.M. Menzies, G.M. Pupo, M.S. Carlino, C. Fung, J. Hyman, L.E. Haydu, B. Mijatov, T.M. Becker, S.C. Boyd, J. Howle, R. Saw, J.F. Thompson, R.F. Kefford, R.A. Scolyer, G.V. Long; BRAF inhibitor resistance mechanisms in metastatic melanoma: spectrum and clinical impact; Clin. Cancer Res., 20 (2014), pp. 1965–1977 http://dx.doi.org/10.1158/1078-0432.CCR-13-3122
- Robert et al., 2014 C. Robert, B. Karaszewska, J. Schachter, P. Rutkowski, A. Mackiewicz, D. Stroiakovski, M. Lichinitser, R. Dummer, F. Grange, L. Mortier, V. Chiarion-Sileni, K. Drucis, I. Krajsova, A. Hauschild, P. Lorigan, P. Wolter, G.V. Long, K. Flaherty, P. Nathan, A. Ribas, A.-M. Martin, P. Sun, W. Crist, J. Legos, S.D. Rubin, S.M. Little, D. Schadendorf; Improved overall survival in melanoma with combined dabrafenib and trametinib; N. Engl. J. Med., 372 (2014) http://dx.doi.org/10.1056/NEJMoa1412690 (141116004513004)
- Saifo et al., 2010 M.S. Saifo, D.R. Rempinski, Y.M. Rustum, R.G. Azrak; Targeting the oncogenic protein beta-catenin to enhance chemotherapy outcome against solid human cancers; Mol. Cancer, 9 (2010), p. 310 http://dx.doi.org/10.1186/1476-4598-9-310
- Sakaguchi et al., 2012 M. Sakaguchi, M. Oka, T. Iwasaki, Y. Fukami, C. Nishigori; Role and regulation of STAT3 phosphorylation at Ser727 in melanocytes and melanoma cells; J. Investig. Dermatol., 132 (2012), pp. 1877–1885 http://dx.doi.org/10.1038/jid.2012.45
- Seiler et al., 2013 C.Y. Seiler, J.G. Park, A. Sharma, P. Hunter, P. Surapaneni, C. Sedillo, J. Field, R. Algar, A. Price, J. Steel, A. Throop, M. Fiacco, J. LaBaer; DNASU plasmid and PSI: biology-materials repositories: resources to accelerate biological research; Nucleic Acids Res., 42 (2013), pp. D1253–D1260 http://dx.doi.org/10.1093/nar/gkt1060
- Shi et al., 2014 H. Shi, W. Hugo, X. Kong, A. Hong, R.C. Koya, G. Moriceau, T. Chodon, R. Guo, D.B. Johnson, K.B. Dahlman, M.C. Kelley, R.F. Kefford, B. Chmielowski, J.A. Glaspy, J.A. Sosman, N. van Baren, G.V. Long, A. Ribas, R.S. Lo; Acquired resistance and clonal evolution in melanoma during BRAF inhibitor therapy; Cancer Discov., 4 (2014), pp. 80–93 http://dx.doi.org/10.1158/2159-8290.CD-13-0642
- Sinnberg et al., 2010 T. Sinnberg, M. Menzel, S. Kaesler, T. Biedermann, B. Sauer, S. Nahnsen, M. Schwarz, C. Garbe, B. Schittek; Suppression of casein kinase 1alpha in melanoma cells induces a switch in beta-catenin signaling to promote metastasis; Cancer Res., 70 (2010), pp. 6999–7009 http://dx.doi.org/10.1158/0008-5472.CAN-10-0645
- Sinnberg et al., 2011 T. Sinnberg, M. Menzel, D. Ewerth, B. Sauer, M. Schwarz, M. Schaller, C. Garbe, B. Schittek; β-Catenin signaling increases during melanoma progression and promotes tumor cell survival and chemoresistance; PLoS One, 6 (2011), p. e23429 http://dx.doi.org/10.1371/journal.pone.0023429
- Smith et al., 2016 M.P. Smith, H. Brunton, E.J. Rowling, J. Ferguson, I. Arozarena, Z. Miskolczi, J.L. Lee, M.R. Girotti, R. Marais, M.P. Levesque, R. Dummer, D.T. Frederick, K.T. Flaherty, Z.A. Cooper, J.A. Wargo, C. Wellbrock; Inhibiting drivers of non-mutational drug tolerance is a salvage strategy for targeted melanoma therapy; Cancer Cell, 29 (2016), pp. 270–284 http://dx.doi.org/10.1016/j.ccell.2016.02.003
- Sos et al., 2014 M.L. Sos, R.S. Levin, J.D. Gordan, J.A. Oses-Prieto, J.T. Webber, M. Salt, B. Hann, A.L. Burlingame, F. McCormick, S. Bandyopadhyay, K.M. Shokat; Oncogene mimicry as a mechanism of primary resistance to BRAF inhibitors; Cell Rep., 8 (2014), pp. 1037–1048 http://dx.doi.org/10.1016/j.celrep.2014.07.010
- Straussman et al., 2012 R. Straussman, T. Morikawa, K. Shee, M. Barzily-Rokni, Z.R. Qian, J. Du, A. Davis, M.M. Mongare, J. Gould, D.T. Frederick, Z.A. Cooper, P.B. Chapman, D.B. Solit, A. Ribas, R.S. Lo, K.T. Flaherty, S. Ogino, J.A. Wargo, T.R. Golub; Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion; Nature, 487 (2012), pp. 500–504 http://dx.doi.org/10.1038/nature11183
- Sun et al., 2014 C. Sun, L. Wang, S. Huang, G.J.J.E. Heynen, A. Prahallad, C. Robert, J. Haanen, C. Blank, J. Wesseling, S.M. Willems, D. Zecchin, S. Hobor, P.K. Bajpe, C. Lieftink, C. Mateus, S. Vagner, W. Grernrum, I. Hofland, A. Schlicker, L.F.A. Wessels, R.L. Beijersbergen, A. Bardelli, F. Di Nicolantonio, A.M.M. Eggermont, R. Bernards; Reversible and adaptive resistance to BRAF(V600E) inhibition in melanoma; Nature, 508 (2014), pp. 118–122 http://dx.doi.org/10.1038/nature13121
- Tiwary et al., 2014 S. Tiwary, M. Preziosi, P.G. Rothberg, N. Zeitouni, N. Corson, L. Xu; ERBB3 is required for metastasis formation of melanoma cells; Oncogenesis, 3 (2014), Article e110 http://dx.doi.org/10.1038/oncsis.2014.23
- Traenkle et al., 2015 B. Traenkle, F. Emele, R. Anton, O. Poetz, R.S. Haeussler, J. Maier, P.D. Kaiser, A.M. Scholz, S. Nueske, A. Buchfellner, T. Romer, U. Rothbauer; Monitoring interactions and dynamics of endogenous beta-catenin with intracellular nanobodies in living cells; Mol. Cell. Proteomics, 14 (2015), pp. 707–723 http://dx.doi.org/10.1074/mcp.M114.044016
- Umbhauer et al., 2000 M. Umbhauer, A. Djiane, C. Goisset, A. Penzo-Méndez, J.F. Riou, J.C. Boucaut, D.L. Shi; The C-terminal cytoplasmic Lys-thr-X-X-X-Trp motif in frizzled receptors mediates Wnt/beta-catenin signalling; EMBO J., 19 (2000), pp. 4944–4954 http://dx.doi.org/10.1093/emboj/19.18.4944
- van Amerongen et al., 2012 R. van Amerongen, C. Fuerer, M. Mizutani, R. Nusse; Wnt5a can both activate and repress Wnt/β-catenin signaling during mouse embryonic development; Dev. Biol., 369 (2012), pp. 101–114 http://dx.doi.org/10.1016/j.ydbio.2012.06.020
- van de Wetering et al., 2003 M. van de Wetering, I. Oving, V. Muncan, M.T. Pon Fong, H. Brantjes, D. van Leenen, F.C.P. Holstege, T.R. Brummelkamp, R. Agami, H. Clevers; Specific inhibition of gene expression using a stably integrated, inducible small-interfering-RNA vector; EMBO Rep., 4 (2003), pp. 609–615 http://dx.doi.org/10.1038/sj.embor.embor865
- Villanueva et al., 2010 J. Villanueva, A. Vultur, J.T. Lee, R. Somasundaram, M. Fukunaga-Kalabis, A.K. Cipolla, B. Wubbenhorst, X. Xu, P.A. Gimotty, D. Kee, A.E. Santiago-Walker, R. Letrero, K. D'Andrea, A. Pushparajan, J.E. Hayden, K.D. Brown, S. Laquerre, G.A. McArthur, J.A. Sosman, K.L. Nathanson, M. Herlyn; Acquired resistance to BRAF inhibitors mediated by a RAF kinase switch in melanoma can be overcome by cotargeting MEK and IGF-1R/PI3K; Cancer Cell, 18 (2010), pp. 683–695 http://dx.doi.org/10.1016/j.ccr.2010.11.023
- Vultur et al., 2014 A. Vultur, J. Villanueva, C. Krepler, G. Rajan, Q. Chen, M. Xiao, L. Li, P.A. Gimotty, M. Wilson, J. Hayden, F. Keeney, K.L. Nathanson, M. Herlyn; MEK inhibition affects STAT3 signaling and invasion in human melanoma cell lines; Oncogene, 33 (2014), pp. 1850–1861 http://dx.doi.org/10.1038/onc.2013.131
- Wagle et al., 2011 N. Wagle, C. Emery, M.F. Berger, M.J. Davis, A. Sawyer, P. Pochanard, S.M. Kehoe, C.M. Johannessen, L.E. Macconaill, W.C. Hahn, M. Meyerson, L.A. Garraway; Dissecting therapeutic resistance to RAF inhibition in melanoma by tumor genomic profiling; J. Clin. Oncol., 29 (2011), pp. 3085–3096 http://dx.doi.org/10.1200/JCO.2010.33.2312
- Wang et al., 2007 W. Wang, H.D. Edington, U.N.M. Rao, D.M. Jukic, S.R. Land, S. Ferrone, J.M. Kirkwood; Modulation of signal transducers and activators of transcription 1 and 3 signaling in melanoma by high-dose IFNalpha2b; Clin. Cancer Res., 13 (2007), pp. 1523–1531 http://dx.doi.org/10.1158/1078-0432.CCR-06-1387
- Wang et al., 2015 X. Wang, Y. Liu, H. Chen, L. Mei, C. He, L. Jiang, Z. Niu, J. Sun, H. Luo, J. Li, Y. Feng; LEF-1 regulates tyrosinase gene transcription in vitro; PLoS One, 10 (2015), Article e0143142 http://dx.doi.org/10.1371/journal.pone.0143142
- Watanabe and Dai, 2011 K. Watanabe, X. Dai; A WNTer revisit: new faces of β-catenin and TCFs in pluripotency; Sci. Signal., 4 (2011), p. pe41 http://dx.doi.org/10.1126/scisignal.2002436
- Weeraratna et al., 2002 A.T. Weeraratna, Y. Jiang, G. Hostetter, K. Rosenblatt, P. Duray, M. Bittner, J.M. Trent; Wnt5a signaling directly affects cell motility and invasion of metastatic melanoma; Cancer Cell, 1 (2002), pp. 279–288
- Widlund et al., 2002 H.R. Widlund, M.A. Horstmann, E.R. Price, J. Cui, S.L. Lessnick, M. Wu, X. He, D.E. Fisher; Beta-catenin-induced melanoma growth requires the downstream target Microphthalmia-associated transcription factor; J. Cell Biol., 158 (2002), pp. 1079–1087 http://dx.doi.org/10.1083/jcb.200202049
- Wilson et al., 2012 T.R. Wilson, J. Fridlyand, Y. Yan, E. Penuel, L. Burton, E. Chan, J. Peng, E. Lin, Y. Wang, J. Sosman, A. Ribas, J. Li, J. Moffat, D.P. Sutherlin, H. Koeppen, M. Merchant, R. Neve, J. Settleman; Widespread potential for growth-factor-driven resistance to anticancer kinase inhibitors; Nature, 487 (2012), pp. 505–509 http://dx.doi.org/10.1038/nature11249
- Zhang et al., 2013 J. Zhang, J.R. Shemezis, E.R. McQuinn, J. Wang, M. Sverdlov, A. Chenn; AKT activation by N-cadherin regulates beta-catenin signaling and neuronal differentiation during cortical development; Neural Dev., 8 (2013), p. 7 http://dx.doi.org/10.1186/1749-8104-8-7
- Zhou, 2011 S. Zhou; TGF-β regulates β-catenin signaling and osteoblast differentiation in human mesenchymal stem cells; J. Cell. Biochem., 112 (2011), pp. 1651–1660 http://dx.doi.org/10.1002/jcb.23079
Document information
Published on 06/04/17
Licence: Other
Share this document
Keywords
claim authorship
Are you one of the authors of this document?