Abstract
Porcine cytomegalovirus (PCMV) is a Betaherpesvirus that causes lifelong latent infections in swine; occasionally, it may be associated with inclusion body rhinitis in piglets and reproductive disorders in pregnant sows. Post-weaning multisystemic wasting syndrome (PMWS) a condition where porcine circovirus type 2 (PCV2) infection is necessary – though not sufficient – to trigger disease, has become one of the major health problems to the porcine productive chain. Despite the high expected prevalence of both PCMV and PCV2 in swine-raising farms, no links between PCMV and PMWS have been investigated so far. In view of that, the present study was conducted to search for relations between PCMV infections and the occurrence of PMWS. Spleen and sera of PMWS-affected and non-PWMS-affected animals were examined. In PMWS-affected animals, PCMV DNA was detected in 88.4% of the spleen samples and 7.6% of the sera, whereas in non-PMWS-affected pigs, PCMV DNA was detected in 72.7% of the spleens and 10% of sera. Such differences were not statistically significant. These findings showed despite the high prevalence of PCMV infections in the swine population examined, no positive or negative association could be inferred from the presence of PCMV DNA and the occurrence of PMWS.
Introduction
Post-weaning multisystemic wasting syndrome (PMWS) is a porcine disease which has become a major burden to the swine productive chain (Gillespie et al. 2009). PMWS is one of the more often reported conditions presently known as ‘porcine circovirus-associated diseases’ (PCVAD). The virus involved, porcine circovirus type 2 (PCV2), is a member of the genus Circovirus, family Circoviridae (International Committee on Taxonomy of Viruses – ICTV, ). PMWS is characterized clinically by pallor of the skin, respiratory distress, occasional diarrhea, jaundice and, most prominently, wasting in post-weaning pigs of about 2–4 months of age (Ellis et al. 1998; Allan & Ellis 2000). PCV2 is necessary, though not sufficient, to induce PMWS (Krakowka et al. 2000; Harms et al. 2001). Variables affecting the virus, the host – including co-infections – and herd management conditions have been shown to influence in development of disease (Opriessnig et al. 2007). Moreover, one or more agents may be involved. Several studies have been conducted in search for links between PCVAD and other swine pathogens (Ellis et al. 1999, 2000; Allan & Ellis 2000; Allan et al. 2000; Krakowka et al. 2000; Pogranichniy et al. 2002; Rose et al. 2003; McMahon et al. 2006; Segales et al. 2009; Blomstrom et al. 2010).
Porcine cytomegalovirus (PCMV, also known as swine herpesvirus 2 - SuHV-2) is a Betaherpesvirus that causes lifelong latent infections in swine; occasionally, it may be associated with inclusion body rhinitis in piglets and reproductive disorders in pregnant sows (Edington et al. 1976). The virus is ubiquitous in swine, where it induces latent infections in the host, a common feature between members of the Herpesviridae family (Tucker et al. 1999; Garkavenko et al. 2004; Davison et al. 2009). Infection can be reactivated and viral replication enhanced by immunosuppressive conditions (Fishman & Rubin 1998; Mueller et al. 2002, 2003; Gollackner et al. 2003). Despite the high prevalence PCMV in swine worldwide (Rondhuis et al. 1980; Assaf et al. 1982; Tajima et al. 1993; Hamel et al. 1999), associations between PCMV and PMWS have not been investigated so far. As PCV2 is an immunosuppressing virus, it became of interest to examine whether PCMV would play any role in the development of PMWS.
Evidence pointing out the immunosuppressive effect of PVC2 is abundant. PMWS-affected pigs show increased incidence of diseases associated with opportunistic pathogens or unexpected pathogens (Carrasco et al. 2000; Nunez et al. 2003; Cavallini Sanches et al. 2006; Zlotowski et al. 2006; Szeredi & Szentirmai 2008). However, the mechanisms by which immunosuppression occurs are not fully understood. Immune function studies in PMWS-affected pigs revealed extensive depletion of lymphocytes in lymphoid tissues (Chianini et al. 2003) and reduction in circulating B and T cells (Nielsen et al. 2003). Moreover, increased interleukin 10 (IL-10) expression was observed in PCV2 infection suggesting that IL-10-mediated immunosuppression may play an important role in the pathogenesis of PMWS (Crisci et al. 2010; Doster et al. 2010). These findings support the involvement of the immune system in PMWS pathogenesis. The aim of this study was to search for some associations between the detection of PCMV DNA in pigs and the occurrence of PMWS.
Materials and methods
Spleen and serum samples
Spleen and serum samples were received from pig farms in Rio Grande do Sul State, Brazil. The case group (PMWS-affected pigs) consisted of 77 spleens from 1 to 4 months old piglets and 92 serum samples from 1 to 4 months old piglets. In the case group, the pigs displayed dyspnoea, enlargement of superficial inguinal lymph nodes, pallor, jaundice and diarrhoea. The diagnosis of PMWS was confirmed by typical macroscopic lesions at necropsy, histopathology and demonstration of PCV2 DNA in tissues by PCR.
The control group (non-PMWS-affected pigs) consisted of 11 samples of spleen tissue from healthy pigs at slaughtering age and serum samples of 119 pigs >6 months old. Additionally, 24 serum samples from healthy from 1 to 4 months old piglets were included as controls.
DNA extraction
DNA extraction from spleens was performed with sodium iodide as described previously (Teixeira et al. 2013). DNA of sera was extracted from 500 μL volumes using a phenol–chloroform method. The extracted DNA was resuspended with 50 μL of TE buffer. The quantity and quality of the extracts were analysed with the aid of a spectrophotometer (Nanodrop® 1000 DE, USA).
PCR for PCMV
Primers for PCMV detection were designed by Hamel and collaborators (Hamel et al. 1999). The primer sequences correspond to nucleotides 37 to 64 and 449 to 420 on the PCMV DNA polymerase gene. The expected size of the amplification product is 413 bp. The PCR was carried out in 25 μL volumes containing 2 μL of DNA (100 ng for spleen samples and 2 μL of DNA extracted from serum samples), 5 pmol of each primer, 0.8 mmol/L dNTPs, 1.5 mmol/L MgCl2 and 1 U Taq DNA polymerase (Invitrogen Carlsbad, USA). The PCR program consisted of an initial reaction at 94°C for 3 min, followed by 35 cycles at 94°C (30 s), 61°C (30 s) and 72°C (30 s), with a final extension of 5 min at 72°C. PCR products were subjected to electrophoresis on a 1% agarose gel, stained with ethidium bromide and visualized on UV light.
To avoid contamination, separate rooms were used to prepare reaction buffers and to perform the PCR reactions, to extract DNA, and to examine PCR products. Disposable filter tips were used throughout. Additional controls with ultra-pure water instead of sample DNA were included in every 10 PCR tubes. Possible inhibitory effects of serum DNA on PCR reaction were evaluated by amplification of an unrelated amplicon in a separate PCR performed with each serum sample (data not shown).
Sensitivity assay
In order to determine the PCMV PCR sensitivity, an amplicon from a spleen containing PCMV DNA (as later confirmed by sequencing) was cloned in plasmid pCR 2.1 following the manufacturers protocol (pCR 2.1 TOPO TA Cloning Kit; Invitrogen Carlsbad, USA). The sensitivity of the PCR was determined by amplification of 10-fold dilutions of known amounts of plasmid DNA. These experiments were repeated three times. The plasmid containing PCMV genome fragment was used as positive control in the PCR throughout the study.
Statistical analysis
Statistical analysis was performed applying the chi-square test available in DagStat (Mackinnon 2000) to compare the proportion of PCMV detected in the case and control groups. The level of significance was set to P ≤ 0.05.
Results
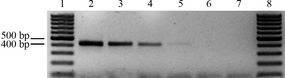
The PCR employed to amplify PCMV genome fragments in spleen and sera of PMWS-affected and non-PMWS-affected pigs was shown to amplify a minimum number of genome copies equivalent to 100 molecules/reaction of PCMV DNA, as extracted from the PCMV DNA-containing plasmid (Fig. 1).
|
|
|
Figure 1. Determination of PCR sensitivity for detection of PCMV. Known copy numbers of the plasmid containing the PCR- targeted region of the PCMV DPOL gene (lanes 2–6: 105–101 molecules per reaction, respectively) were submitted to PCR amplification (for details refer to methods). The detection limit of the PCR was 100 molecules/reaction. Lanes 1 and 8, molecular size marker; lane 7: negative control. DPOL, DNA polymerase; PCMV, porcine cytomegalovirus. |
In spleen tissues, PCMV DNA was detected in 88.3% (68/77) of samples from PMWS-affected pigs and in 72.7% (8/11) from non-PMWS-affected pigs (Table 1). Such differences were not statistically significant (P ≥ 0.05).
| Groups | No. of samples tested | Samples with PCMV DNA | |
|---|---|---|---|
| Spleen | PMWS-affected pigs (1–4 months old) | 77 | 68 (88.4%) |
| Non-PMWS-affected pigs (>6 months old) | 11 | 8 (72.7%) | |
| Total | 88 | 76 (86.4%) | |
| Serum | PMWS-affected pigs (1–4 months old) | 92 | 7 (7.6%) |
| Non-PMWS-affected pigs (>6 months old) | 119 | 14 (11.8%) | |
| Non-PMWS-affected pigs (1–4 months old) | 24 | 2 (8.3%) | |
| Total | 235 | 24 (10.2%) | |
| PCMV, porcine cytomegalovirus; PMWS, post-weaning multisystemic wasting syndrome. | |||
In sera, PCMV DNA was identified in 7.6% (7/92) of PMWS-affected pigs (Table 1), whereas in non-PMWS-affected pigs PCMV DNA was detected in 8.3% (2/24) in 1–4 months old pigs 11.8% (14/119) of >6 months old pigs (Table 1). Statistical analysis of the frequencies of PCMV DNAemia in PMWS-affected and non-PMWS-affected groups revealed that such frequencies were not significantly different (P ≥ 0.05).
Discussion and conclusions
Associations of PCV2 with other swine pathogens have been shown to be play some role in the development of PCVAD. Co-infections with PCV2 and other pig pathogens may lead to the development of PMWS (Allan & Ellis 2000). Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) can act synergistically with PCV2 and lead to PMWS development (Pogranichniy et al. 2002; Rose et al. 2003). Porcine parvovirus (PPV) has also been found to play a role in PMWS: co-infections with PCV2 and PPV were detected in approximately 15% of PMWS cases (Ellis et al. 2000). In addition, experimental co-infections with PCV2 and PPV resulted in PMWS reproduction in gnotobiotic, colostrum-deprived pigs (Ellis et al. 1999). Other pathogens which have been reported to be capable of participating in the establishment of PMWS include Mycoplasma hyopneumoniae (Pallares et al. 2002), Cryptosporidium parvum (Nunez et al. 2003), Aujeszkys disease virus (Quintana et al. 2001; Maldonado et al. 2005), influenza virus and bacterial pneumonias (Kim et al. 2002; Pogranichniy et al. 2002; Dorr et al. 2007; Wei et al. 2010). Against such a background, a search for a role for PCMV in PMWS, particular in view of the immunosuppressive potential of PCV2, would be expected to reveal some sort of interaction. PCV2 is known to induce immunosuppressive effects on the host, predisposing to viral, bacterial and mycotic infections (Segales et al. 2004). Opportunistic infections have often been observed in herds with PMWS (Carrasco et al. 2000; Kim et al. 2002; Pallares et al. 2002; Nunez et al. 2003; Cavallini Sanches et al. 2006; Zlotowski et al. 2006). Whether the immunosuppressing effect of PCV2 facilitates co-infection or whether a co-infecting pathogen would induce immunosuppression that could trigger PMWS still remains a matter of controversy.
In the present study, a search was made for PCMV in spleens and sera of PMWS-affected and non-PMWS-affected pigs. The presence of PCMV DNA in spleen is indicative of previous infection. This is important in view of the biology of herpesviruses, where latent infections are produced and active virus replication may not be directly linked to any particular disease.
In humans, human cytomegalovirus-HCMV-DNA in serum was considered a sign of active viral replication, and was associated to clinical disease (Spector et al. 1992; Wolf & Spector 1993). On the other hand, HCMV DNA in leukocytes was considered indicative of latent infections (Ishigaki et al. 1991). Here, PCMV DNA was detected in serum in proportions that did not differ significantly between groups of PWMS-affected and non-PMWS-affected groups of pigs. A slightly larger proportion of PCMV DNA-containing samples was found in older, non-PMWS-affected pigs, (Table 1). However, this finding was probably more related to exposure related to aging than to other conditions, since these pigs had no signs of disease. The proportions of PCMV DNA-bearing animals were not significantly different between the two groups, revealing that the presence of PCMV DNA in serum is not associated to the occurrence of PMWS in the sampled population. More likely, PCMV DNA in sera of both PMWS-affected and non-affected pigs at equivalent proportions suggests that serum PCMV DNA is indicative of latent, rather than active PCMV infections.
The results presented here show that PCMV DNA was detected in spleens of a high proportion of the sampled animals – either with or without PWMS. The frequency of detection of PCMV DNA in the sampled population is similar to those reported in previous studies (Rondhuis et al. 1980; Assaf et al. 1982; Tajima et al. 1993; Hamel et al. 1999; Goltz et al. 2000). Despite the high frequency of PCMV and PCV2 infections in both groups of animals, no association between the presence of PCMV DNA and the occurrence of PMWS could be inferred. Therefore, despite the high prevalence of PCMV in the examined population of pigs, PCMV does not seem to play any significant role in the occurrence of PMWS.
In summary, results reported here show that the prevalence of PCMV infections in the population examined is high in both, PMWS-affected and non-PMWS-affected animals. Therefore, PCMV does not seem to play any significant role in the development of PMWS.
Acknowledgements
We are grateful to Doctor David Driemeier and David Barcellos for providing sera samples. SPC was in receipt of a Ph.D. grant from Conselho Nacional de Desenvolvimento Científico – CNPq.
Source of funding
This work was financially supported by FINEP (Grant 01.10.0783.04), CNPq (Grant 302427/2010-4) and FAPERGS (Proj. 19712.341.15150.16012015).
Conflicts of interest
The authors have declared that no competing interests exist.
Contributions
Conceived and designed the experiments: SPC & PMR. Performed the experiments: SPC, GP, TFT, CLH & APMV. Analyzed the data: SPC, DD, ACF. Contributed reagents/materials/analysis tools: PMR ACF. Wrote the paper: SPC & PMR.
References
- Allan G.M. & Ellis J.A. (2000) Porcine circoviruses: a review. Journal of Veterinary Diagnostic Investigation12, 3–14.
- Allan G.M., McNeilly F., Ellis J., Krakowka S., Meehan B., McNair I.et al. (2000) Experimental infection of colostrum deprived piglets with porcine circovirus 2 (PCV2) and porcine reproductive and respiratory syndrome virus (PRRSV) potentiates PCV2 replication. Archives of Virology145, 2421–2429.
- Assaf R., Bouillant A.M. & Di Franco E. (1982) Enzyme linked immunosorbent assay (ELISA) for the detection of antibodies to porcine cytomegalovirus. Canadian Journal of Comparative Medicine46, 183–185.
- Blomstrom A.L., Belak S., Fossum C., Fuxler L., Wallgren P. & Berg M. (2010) Studies of porcine circovirus type 2, porcine boca-like virus and torque teno virus indicate the presence of multiple viral infections in postweaning multisystemic wasting syndrome pigs. Virus Research152, 59–64.
- Carrasco L., Segales J., Bautista M.J., Gomez-Villamandos J.C., Rosell C., Ruiz-Villamor E.et al. (2000) Intestinal chlamydial infection concurrent with postweaning multisystemic wasting syndrome in pigs. The Veterinary Record146, 21–23.
- Cavallini Sanches E.M., Borba M.R., Spanamberg A., Pescador C., Corbellini L.G., Ravazzolo A.P.et al. (2006) Co-infection of Pneumocystis carinii f. sp. suis and porcine circovirus-2 (PCV2) in pig lungs obtained from slaughterhouses in southern and midwestern regions of Brazil. The Journal of Eukaryotic Microbiology53 (Suppl. 1), S92–S94.
- Chianini F., Majo N., Segales J., Dominguez J. & Domingo M. (2003) Immunohistochemical characterisation of PCV2 associate lesions in lymphoid and non-lymphoid tissues of pigs with natural postweaning multisystemic wasting syndrome (PMWS). Veterinary Immunology and Immunopathology94, 63–75.
- Crisci E., Ballester M., Dominguez J., Segales J. & Montoya M. (2010) Increased numbers of myeloid and lymphoid IL-10 producing cells in spleen of pigs with naturally occurring postweaning multisystemic wasting syndrome. Veterinary Immunology and Immunopathology136, 305–310.
- Davison A.J., Eberle R., Ehlers B., Hayward G.S., McGeoch D.J., Minson A.C.et al. (2009) The order herpesvirales. Archives of Virology154, 171–177.
- Dorr P.M., Baker R.B., Almond G.W., Wayne S.R. & Gebreyes W.A. (2007) Epidemiologic assessment of porcine circovirus type 2 coinfection with other pathogens in swine. Journal of the American Veterinary Medical Association230, 244–250.
- Doster A.R., Subramaniam S., Yhee J.Y., Kwon B.J., Yu C.H., Kwon S.Y.et al. (2010) Distribution and characterization of IL-10-secreting cells in lymphoid tissues of PCV2-infected pigs. Journal of Veterinary Science11, 177–183.
- Edington N., Plowright W. & Watt R.G. (1976) Generalized porcine cytomegalic inclusion disease: distribution of cytomegalic cells and virus. Journal of Comparative Pathology86, 191–202.
- Ellis J., Hassard L., Clark E., Harding J., Allan G., Willson P.et al. (1998) Isolation of circovirus from lesions of pigs with postweaning multisystemic wasting syndrome. The Canadian Veterinary Journal39, 44–51.
- Ellis J., Krakowka S., Lairmore M., Haines D., Bratanich A., Clark E.et al. (1999) Reproduction of lesions of postweaning multisystemic wasting syndrome in gnotobiotic piglets. Journal of Veterinary Diagnostic Investigation11, 3–14.
- Ellis J.A., Bratanich A., Clark E.G., Allan G., Meehan B., Haines D.M.et al. (2000) Coinfection by porcine circoviruses and porcine parvovirus in pigs with naturally acquired postweaning multisystemic wasting syndrome. Journal of Veterinary Diagnostic Investigation12, 21–27.
- Fishman J.A. & Rubin R.H. (1998) Infection in organ-transplant recipients. The New England Journal of Medicine338, 1741–1751.
- Garkavenko O., Muzina M., Muzina Z., Powels K., Elliott R.B. & Croxson M.C. (2004) Monitoring for potentially xenozoonotic viruses in New Zealand pigs. Journal of Medical Virology72, 338–344.
- Gillespie J., Opriessnig T., Meng X.J., Pelzer K. & Buechner-Maxwell V. (2009) Porcine circovirus type 2 and porcine circovirus-associated disease. Journal of Veterinary Internal Medicine23, 1151–1163.
- Gollackner B., Mueller N.J., Houser S., Qawi I., Soizic D., Knosalla C.et al. (2003) Porcine cytomegalovirus and coagulopathy in pig-to-primate xenotransplantation. Transplantation75, 1841–1847.
- Goltz M., Widen F., Banks M., Belak S. & Ehlers B. (2000) Characterization of the DNA polymerase loci of porcine cytomegaloviruses from diverse geographic origins. Virus Genes21, 249–255.
- Hamel A.L., Lin L., Sachvie C., Grudeski E. & Nayar G.P. (1999) PCR assay for detecting porcine cytomegalovirus. Journal of Clinical Microbiology37, 3767–3768.
- Harms P.A., Sorden S.D., Halbur P.G., Bolin S.R., Lager K.M., Morozov I.et al. (2001) Experimental reproduction of severe disease in CD/CD pigs concurrently infected with type 2 porcine circovirus and porcine reproductive and respiratory syndrome virus. Veterinary Pathology38, 528–539.
- ICTV- International Committee on Taxonomy of Viruses. In: http://www.ictvonline.org/virusTaxonomy.asp. Accessed on 12/15/2014.
- Ishigaki S., Takeda M., Kura T., Ban N., Saitoh T., Sakamaki S.et al. (1991) Cytomegalovirus DNA in the sera of patients with cytomegalovirus pneumonia. British Journal of Haematology79, 198–204.
- Kim J., Chung H.K., Jung T., Cho W.S., Choi C. & Chae C. (2002) Postweaning multisystemic wasting syndrome of pigs in Korea: prevalence, microscopic lesions and coexisting microorganisms. The Journal of Veterinary Medical Science64, 57–62.
- Krakowka S., Ellis J.A., Meehan B., Kennedy S., McNeilly F. & Allan G. (2000) Viral wasting syndrome of swine: experimental reproduction of postweaning multisystemic wasting syndrome in gnotobiotic swine by coinfection with porcine circovirus 2 and porcine parvovirus. Veterinary Pathology37, 254–263.
- Mackinnon A. (2000) A spreadsheet for the calculation of comprehensive statistics for the assessment of diagnostic tests and inter-rater agreement. Computers in Biology and Medicine30, 127–134.
- Maldonado J., Segales J., Martinez-Puig D., Calsamiglia M., Riera P., Domingo M.et al. (2005) Identification of viral pathogens in aborted fetuses and stillborn piglets from cases of swine reproductive failure in Spain. Veterinary Journal169, 454–456.
- McMahon K.J., Minihan D., Campion E.M., Loughran S.T., Allan G., McNeilly F.et al. (2006) Infection of pigs in Ireland with lymphotropic gamma-herpesviruses and relationship to postweaning multisystemic wasting syndrome. Veterinary Microbiology116, 60–68.
- Mueller N.J., Barth R.N., Yamamoto S., Kitamura H., Patience C., Yamada K.et al. (2002) Activation of cytomegalovirus in pig-to-primate organ xenotransplantation. Journal of Virology76, 4734–4740.
- Mueller N.J., Sulling K., Gollackner B., Yamamoto S., Knosalla C., Wilkinson R.A.et al. (2003) Reduced efficacy of ganciclovir against porcine and baboon cytomegalovirus in pig-to-baboon xenotransplantation. American Journal of Transplantation3, 1057–1064.
- Nielsen J., Vincent I.E., Botner A., Ladekaer-Mikkelsen A.S., Allan G., Summerfield A.et al. (2003) Association of lymphopenia with porcine circovirus type 2 induced postweaning multisystemic wasting syndrome (PMWS). Veterinary Immunology and Immunopathology92, 97–111.
- Nunez A., McNeilly F., Perea A., Sanchez-Cordon P.J., Huerta B., Allan G.et al. (2003) Coinfection by Cryptosporidium parvum and porcine circovirus type 2 in weaned pigs. Journal of Veterinary Medicine B, Infectious Diseases and Veterinary Public Health50, 255–258.
- Opriessnig T., Meng X.J. & Halbur P.G. (2007) Porcine circovirus type 2 associated disease: update on current terminology, clinical manifestations, pathogenesis, diagnosis, and intervention strategies. Journal of Veterinary Diagnostic Investigation19, 591–615.
- Pallares F.J., Halbur P.G., Opriessnig T., Sorden S.D., Villar D., Janke B.H.et al. (2002) Porcine circovirus type 2 (PCV-2) coinfections in US field cases of postweaning multisystemic wasting syndrome (PMWS). Journal of Veterinary Diagnostic Investigation14, 515–519.
- Pogranichniy R.M., Yoon K.J., Harms P.A., Sorden S.D. & Daniels M. (2002) Case-control study on the association of porcine circovirus type 2 and other swine viral pathogens with postweaning multisystemic wasting syndrome. Journal of Veterinary Diagnostic Investigation14, 449–456.
- Quintana J., Segales J., Rosell C., Calsamiglia M., Rodriguez-Arrioja G.M., Chianini F.et al. (2001) Clinical and pathological observations on pigs with postweaning multisystemic wasting syndrome. The Veterinary Record149, 357–361.
- Rondhuis P.R., de Jong M.F. & Schep J. (1980) Indirect fluorescence antibody studies of porcine cytomegalo virus infections in the Netherlands. Tijdschrift voor Diergeneeskunde105 (Suppl. 2), 56–68.
- Rose N., Larour G., Le Diguerher G., Eveno E., Jolly J.P., Blanchard P.et al. (2003) Risk factors for porcine post-weaning multisystemic wasting syndrome (PMWS) in 149 French farrow-to-finish herds. Preventive Veterinary Medicine61, 209–225.
- Segales J., Domingo M., Chianini F., Majo N., Dominguez J., Darwich L.et al. (2004) Immunosuppression in postweaning multisystemic wasting syndrome affected pigs. Veterinary Microbiology98, 151–158.
- Segales J., Martinez-Guino L., Cortey M., Navarro N., Huerta E., Sibila M.et al. (2009) Retrospective study on swine Torque teno virus genogroups 1 and 2 infection from 1985 to 2005 in Spain. Veterinary Microbiology134, 199–207.
- Spector S.A., Merrill R., Wolf D. & Dankner W.M. (1992) Detection of human cytomegalovirus in plasma of AIDS patients during acute visceral disease by DNA amplification. Journal of Clinical Microbiology30, 2359–2365.
- Szeredi L. & Szentirmai C. (2008) Gastric zygomycosis in a pig affected with postweaning multisystemic wasting syndrome–case report. Acta Veterinaria Hungarica56, 207–213.
- Tajima T., Hironao T., Kajikawa T. & Kawamura H. (1993) Application of enzyme-linked immunosorbent assay for the seroepizootiological survey of antibodies against porcine cytomegalovirus. The Journal of Veterinary Medical Science55, 421–424.
- Teixeira T.F., Dezen D., Cibulski S.P., Varela A.P., Sheffer C.M., Holz C.L.et al. (2013) Torque teno sus virus (TTSuV) in tissues of pigs and its relation with the occurrence of postweaning multisystemic wasting syndrome. Virus Genes47, 276–281.
- Tucker A.W., Galbraith D., McEwan P. & Onions D. (1999) Evaluation of porcine cytomegalovirus as a potential zoonotic agent in xenotransplantation. Transplantation Proceedings31, 915.
- Wei H., Lenz S.D., Van Alstine W.G., Stevenson G.W., Langohr I.M. & Pogranichniy R.M. (2010) Infection of cesarean-derived colostrum-deprived pigs with porcine circovirus type 2 and Swine influenza virus. Comparative Medicine60, 45–50.
- Wolf D.G. & Spector S.A. (1993) Early diagnosis of human cytomegalovirus disease in transplant recipients by DNA amplification in plasma. Transplantation56, 330–334.
- Zlotowski P., Rozza D.B., Pescador C.A., Barcellos D.E., Ferreiro L., Sanches E.M.et al. (2006) Muco-cutaneous candidiasis in two pigs with postweaning multisystemic wasting syndrome. Veterinary Journal171, 566–569.
Document information
Published on 09/06/17
Submitted on 09/06/17
Licence: Other
Share this document
Keywords
claim authorship
Are you one of the authors of this document?