Abstract
Dermatophilus congolensis, the aetiological agent of dermatophilosis, is a pleomorphic, Gram-positive actinomycete, which infects animals and humans. Often, there is a wrong diagnosis of the infection in animals because of the close resemblance of the organism with other members of the family Actinomycetaceae. In this study, molecular tools were applied to suspected isolates of D. congolensis obtained from naturally infected cattle in Nigeria for confirmation of dermatophilosis. DNA extraction from 54 suspected pure colonies of D. congolensis was carried out using the QIAamp® DNA Mini extraction kit. PCR targeted at the 16S rRNA gene was employed for the confirmation of D. congolensis using 5′-ACATGCAAGTCGAACGATGA-3′ and 5′-ACGCTCGCACCCTACGTATT-3′ as forward and reverse primers, respectively. Positive amplicons were then sequenced directly using Big Dye Terminator Cycle Sequencing Kit with the forward primers and AmpliTaq-FS DNA Polymerase. Nucleotide sequences were aligned using bioedit (Ibis Biosciences Carlsbad, CA USA) and the phylogenetic analysis was carried out using mega 5.2 (Center for Evolutionary Medicine and Informatics, The Biodesign Institute, Tempe, Arizona, USA) software programme. The aligned nucleotide sequences of 10 positive D. congolensis isolates had between 94% to 99% homology with the sequences of D. congolensis satellite DNA in GenBank. This result also revealed that the sequenced D. congolensis are of different strains. Phylogenetic analysis revealed that D. congolensis, though closely related to Nocardia brasiliensis (NR 074743.01) and Streptomyces sp. (JN 400114.1), belongs to different genus. In conclusion, molecular tools employed in the study were able to confirm the identity of the test organisms as D. congolensis. It can also be concluded that two strains of D. congolensis obtained from the study can still be accommodated within the previously listed strains available in GenBank while the remaining eight may be different strains of D. congolensis not yet listed in GenBank.
Introduction
Dermatophilus congolensis, the aetiological agent of dermatophilosis (lumpy wool, mycotic dermatitis, streptotrichosis, streptothricosis, kirchi), is a pleomorphic, Gram-positive actinomycete which primarily infects the epidermis of many species of animals, and humans (Pal & Matsusaka 1993; Hirsh et al. 2004). It presents as a chronic or acute exudative dermatitis with scab formation on the skin. The disease had been reported to be more severe in ruminants and is of particular importance in the tropics and sub-tropics where it can cause substantial economic losses (Zaria 1993; Aning & Koney 1996; Ambrose et al. 1999; Makinde et al. 2001). The diagnosis of dermatophilosis is routinely made using conventional methods like clinical signs, direct smear and isolation of the organism, other methods such as serology and animal inoculation have been employed, though they are not used routinely (Makinde et al. 1991). Generally, difficulty abounds in confirming cases of dermatophilosis due to close resemblance of the organism with other members of the family Actinomycetaceae (Makinde et al. 2001). Isolates of D. congolensis obtained from different animal species have also been found to react differently to biochemical test reagents thereby giving inconsistent/variable reactions that are not specific for characterization and identification of the organism. For example, varying results have been reported by different researchers with maltose, sucrose, galactose and fructose and other reagents (Van Sacegham 1934; Macadam & Haalstra 1971; Gordon 1976). The tendency of a mixed infection and or difficulty in isolating the organism from field cases using the conventional techniques is the impetus for a search for a more efficient diagnostic approach. In this study, we employed molecular techniques to authenticate field isolates of D. congolensis obtained from natural disease outbreaks and made attempts at elucidating the evolutionary relationships between the strains of D. congolensis and other key members of the family Actinomycetaceae that can mask the diagnosis of the infection.
Materials and methods
Source of isolates
Fifty-four suspected isolates of D. congolensis obtained from infected cattle in Abeokuta, Ogun State and Ilorin, Kwara State Nigeria were examined in this study. The isolates were from skin scrapings collected from clinically infected cattle at a major abattoir in Abeokuta and from nomadic cattle herds in Ilorin.
Dermatophilus congolensis detection by PCR
Dermatophilus congolensis was diagnosed using PCR according to the method described by Shaibu et al. (2010).
DNA extraction
Genomic DNA extraction was carried out using the QIAamp® DNA Mini extraction kit (Qiagen Inc. Valencia, CA USA) according to the manufacturers protocol. Suspected bacterial colonies were inoculated in buffered peptone broth and incubated for 24 h at 37°C. Bacteria were pelleted by centrifugation for 10 min at 5000 × g (7500 rpm) and the pellets were suspended in 180 μL of enzyme solution (20 mg mL−1 lysozyme; 20 mM Tris·HCl, pH 8.0; 2 mM EDTA; 1.2% Triton). This was incubated for 30 min at 37°C.
To the solution, 20 μL of proteinase K and 200 μL of Buffer AL were added. This was completely mixed by vortexing. The mixture was incubated at 56°C for 30 min and then for a further 15 min at 95°C. To this solution, 200 μL of ethanol (100%) was added and mixed again by pulse-vortexing for 15 s.
The mixture (including the precipitate) was applied to the QIAamp Mini spin column (in a 2 mL collection tube) and centrifuged at 6000 × g (8000 rpm) for 1 min. The spin column was then placed in a clean 2 mL collection tube, and the tube containing the filtrate was discarded. Five hundred microlitres Buffer AW1 was added to the spin column and the mixture centrifuged at 6000 × g (8000 rpm) for 1 min. The spin column was placed in a clean 2 mL collection tube, and the collection tube containing the filtrate was discarded.
To the solution in the spin column, 500 μL Buffer AW2 was added and centrifuged at full speed (20 000 × g; 14 000 rpm) for 3 min. The spin column was placed in a new 2 mL collection tube and the old collection tube with the filtrate discarded. This was centrifuged at full speed for 1 min. The spin column was placed in a clean 1.5 mL microcentrifuge tube, and the collection tube containing the filtrate discarded. Two hundred microlitres of Buffer AE was added and the solution was incubated at room temperature for 1 min before being centrifuged at 6000 × g (8000 rpm) for 1 min. The eluted DNA was collected in a clean 1.5 mL microcentrifuge tube and stored in a refrigerator until further use.
The PCR assay
The primers used for the PCR assay were designed from the 16S rRNA gene of an isolate of D. congolensis in the GenBank according to the method described by Han et al. (2007). The sequences of specific primers used in this study were 5′-ACATGCAAGTCGAACGATGA-3′ and 5′-ACGCTCGCACCCTACGTATT-3′ (synthesized by Bioneer Incorporation Alameda, CA, USA) as forward and reverse primers, respectively.
DNA extracted from the suspected D. congolensis was used for the amplification, which was targeted at 500 bp fragment of the 16S rRNA gene of the organism. This was done in a total reaction mixture of 25 μL, consisting of buffer (2.5 μL), Mg2+ (1.0 μL), primer (0.3 μL each), dNTPs (0.8 μL), Taq (0.4 μL), sample DNA (1.0 μL) and nuclease free water (18.7 μL).
With the use of a thermo cycler, the DNA was initially denatured at 95°C for 1 min, followed by denaturation at 94°C for 30 s, annealing at 60°C for 30 s and extension at 72°C for 1 min. This was done for a total of 39 cycles, and a final extension at 72°C for 10 min. Nuclease free water was used as a negative control while Escherichia coli (ATCC 25922) was included as non-specific DNA template.
Ten microlitres of the PCR products was electrophoresed in 1.5% agarose gel containing 3 μL of 10 mg mL−1 GelRed at 80 V for 45 min. One hundred base pair marker (Phenix Research products,Candler, NC, USA) was used as a molecular size ladder. The amplified DNA bands were examined on a transilluminator and the sizes of the amplified PCR products were taken as positives at 500 bp of the molecular size ladder.
DNA sequencing and sequence analysis
To study the gene products further and conduct phylogenetic analysis, 10 positive samples of D. congolensis were selected for sequencing. The PCR products of D. congolensis were sequenced directly using Big Dye Terminator Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA) with the forward amplification PCR primers and AmpliTaq-FS DNA Polymerase.
Bioinformatics analysis
The sequences of 10 D. congolensis were blasted according to Altschul et al. (1990) against existing sequences in the GenBank anchored by National Center for Biotechnology Information (NCBI). The nucleotide sequences were aligned using bioedit and the phylogenetic analysis was carried out using mega 5.2 software programme. Bootstrap confidence values were obtained with 100 resamplings.
Results
PCR amplification
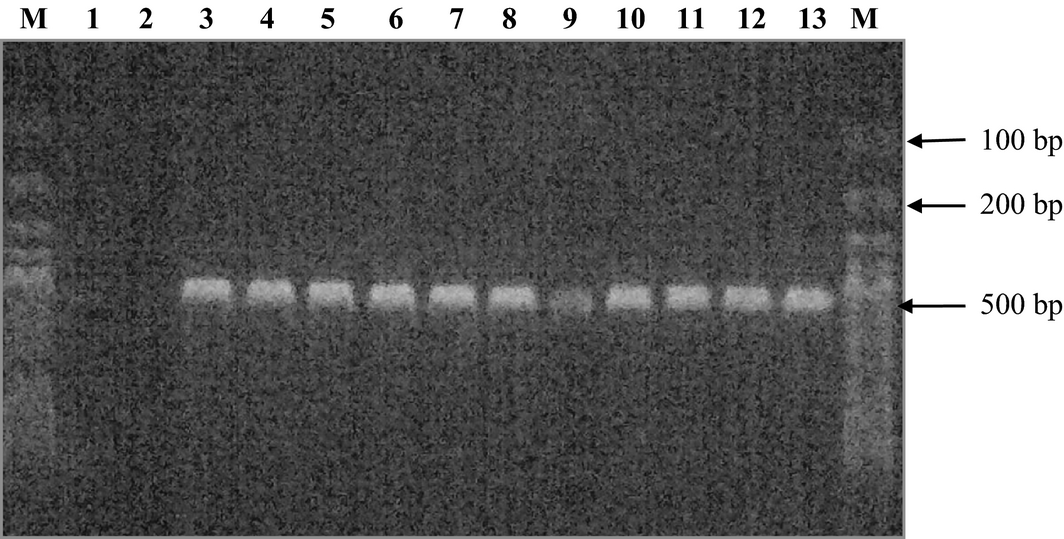
The results of the amplification of the D. congolensis isolates from infected cattle are presented in Fig. 1. It shows the amplification of the partial 16S rRNA gene of a band at approximately 500 base pairs (bp).
|
|
|
Figure 1. PCR amplification of a fragment of 16S rRNA gene from Dermatophilus congolensis isolates. Lane M; 100 bp ladder, lanes 1–2; Nuclease free water and E.coli (accession number: ATCC 25922), respectively, lanes 3–13; cattle isolates of D. congolensis. |
Sequence analysis of the amplified PCR products
Using sequences retrieved from GenBank, the aligned D. congolensis sequences from 10 positive samples analysed had between 94% to 99% homology with the sequence of D. congolensis satellite DNA in the GenBank (Table 1).
| Strain | Homology (%) | Strain | Accession number |
|---|---|---|---|
| D356 | 94 | FJ708621.1 | S3 |
| D357 | 99 | AB550800.1 | NBRC105199 |
| D360 | 94 | AB550800.1 | NBRC105199 |
| D362 | 99 | NR041990.1 | DSM44180 |
| D363 | 99 | L40615.1 | D.C.16SrRNA |
| D364 | 96 | FJ708620.1 | S2 |
| D367 | 98 | AB550800.1 | NBRC105199 |
| D368 | 99 | FJ708617.1 | C2 |
| D373 | 97 | AB550800.1 | NBRC105199 |
| D374 | 96 | NR041990.1 | DSM44180 |
Alignment of nucleotide sequences of 10 strains of D. congolensis
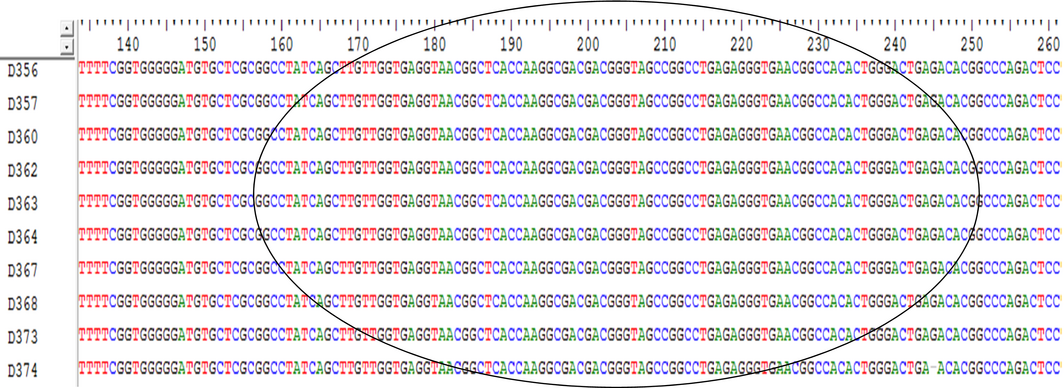
The alignment of the nucleotide sequences of 10 positive strains of D. congolensis organism isolated from infected cattle are shown below in Fig. 2 and Table 2. Variations in the aligned nucleotide sequences were observed as points of alterations, deletions and insertions.
|
|
|
Figure 2. Showing conserved regions in the multiple sequence alignments of 10 positive PCR products of Dermatophilus congolensis. |
| Strain | Point of deletion | Point of insertion | Point of alteration |
|---|---|---|---|
| D356 | – | – | 21, 22 and 32 |
| D357 | – | – | 37, 41, 43 and 77 |
| D360 | – | – | 32, 37 and 44 |
| D362 | – | – | 22, 32, 43 and 77 |
| D363 | – | – | 21, 22, 32, 41, 43 and 77 |
| D364 | – | – | 32, 37 and 44 |
| D367 | – | 65 | 21, 22 and 77 |
| D368 | – | – | 37, 41, 43 and 77 |
| D373 | – | – | 21, 22, 32, 37 and 44 |
| D374 | 80 and 81 | – | 44 |
Phylogenetic analysis of D. congolensis from infected cattle
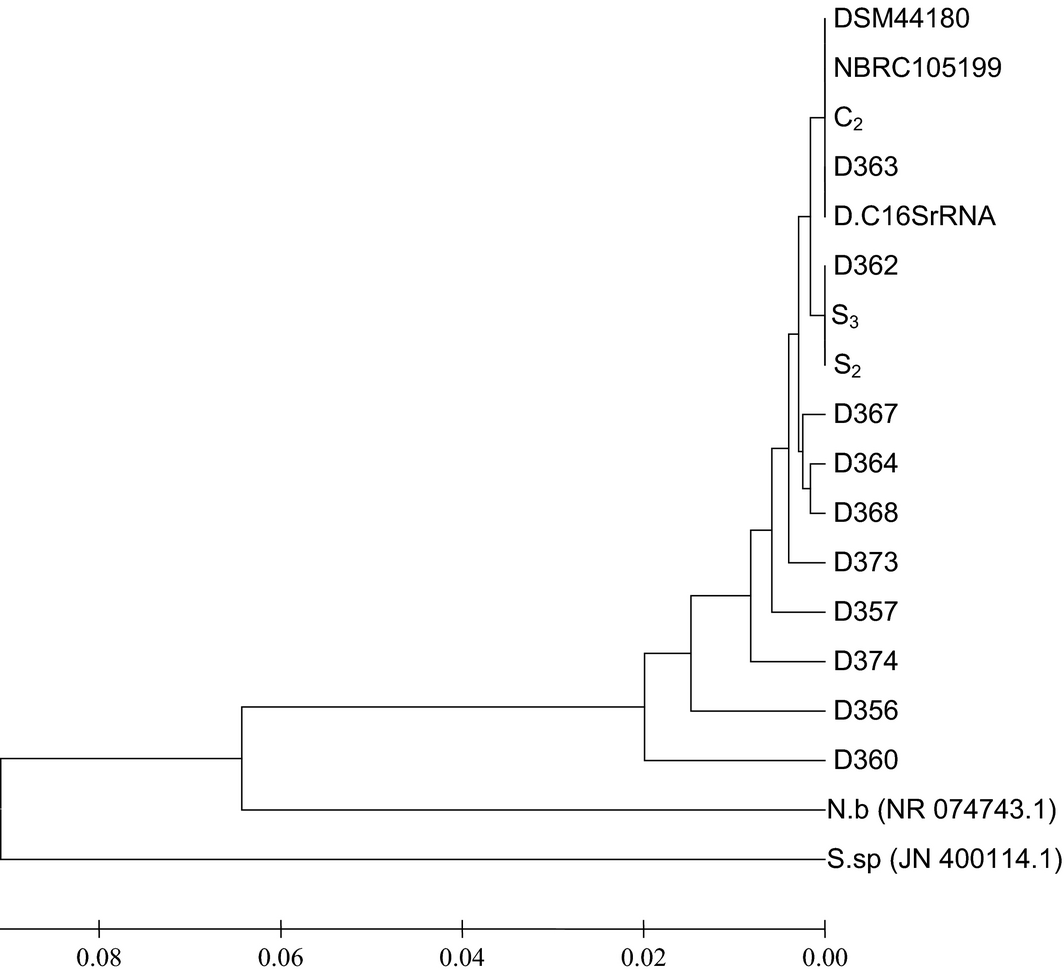
Figure 3 shows the analysis of the phylogenetic relatedness of strains of D. congolensis organisms from infected cattle as examined in the study using 100 bootstrap values. Nocardia brasiliensis; NR 074743.01 and Streptomyces sp.; JN 400114.1 were included in the phylogenetic tree as non-members of the genus Dermatophilus. The result indicates three different clades of bacterial pathogens and reveals a striking genetic diversity between the isolated D. congolensis organisms and the other bacteria pathogens which are non-members of the genus Dermatophilus (P ˂ 0.05).
|
|
|
Figure 3. Phylogenetic tree showing the genetic diversity between the sequenced Dermatophilus congolensis strains and the control bacteria pathogens; Nocardia brasiliensis (accession number: NR 074743.01) and Streptomyces sp. (accession number: JN 400114.1). |
Discussion
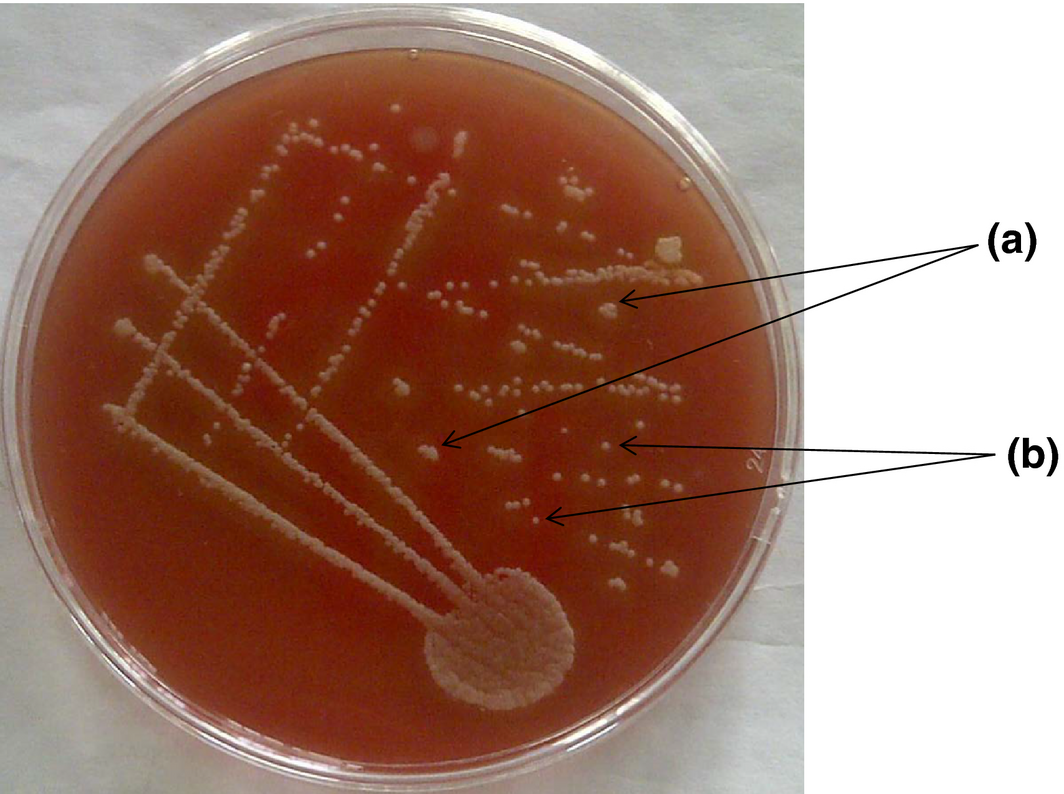
The results obtained from the amplification of a 500 bp fragment of the 16S ribosomal RNA gene of D. congolensis agrees with the works of previous researchers; Han et al. (2007); Shaibu et al. (2010). Using the PCR technique, we were able to confirm 51 of the 54 isolates as D. congolensis (Fig. 4).
|
|
|
Figure 4. Suspected colonies of Dermatophilus congolensis from Abeokuta on 5% sheep blood agar showing the rough (a) and the smooth colonies (b) surrounded by zones of beta haemolysis after 48 h of incubation. |
The study revealed great similarities in the nucleotide sequences of all the 10 positive isolates of D. congolensis sequenced. The observation of nucleotide similarities within isolates of D. congolensis had also been previously reported by Shaibu et al. (2011). These conserved regions may be responsible for a shared functionality of the encoded genes across the strains sequenced. However, some areas of nucleotide variations were observed in the study, which could be due to evolutionary changes of the organism to changing host immunity or mutations caused by drug effects over a period of time.
The result from the phylogenetic analysis revealed that D. congolensis, though closely related to N. brasiliensis (NR 074743.01) and Streptomyces sp. (JN 400114.1), belongs to different clusters in the phylogenetic tree. This result confirmed the isolates from the study as D. congolensis and further revealed that N. brasiliensis is more closely related to D. congolensis than Streptomyces sp. It had been previously reported that colonies of D. congolensis have great semblance with those of Nocardia spp. and Streptomyces spp. (OIE, 2008) which all belong to the same order, i.e. Actinomycetales.
Two strains of D. congolensis from the study clustered with previously reported strains of D. congolensis elsewhere outside the study areas both within Nigeria and outside the shores of Nigeria while the remaining eight clustered out separately.
The results from this study also indicate that there is more genetic resemblance between strain D 363 from the study and strains DSM44180, NBRC105199, C2 and D.C16SrRNA obtained from GenBank. In the same vein, strain D 362 from the study is closely related to strains S3 and S2 obtained from the GenBank.
It is worth mentioning that strain D 362 from the study was obtained in the same geopolitical zone (North Central Nigeria) as strains FJ 708621.1 and FJ 708620.1. It is possible that the movement of nomadic cattle in this zone increased the propensity of spreading the disease, which may help explain the genetic linkage between the strains. The remaining eight strains (i.e. strains D 360, D 356, D 374, D 357, D 373, D 368, D 364 and D 367) clustered out differently and do not show as close genetic relatedness to the D. congolensis strains obtained from the GenBank as the other two strains (i.e. D 363 and D 362) from the study.
Conclusion
This study further indicates that the PCR technique is a preferable technique to the conventional culture technique for the confirmation of D. congolensis organisms during epidemiological surveys. Similarities of nucleotide sequences observed in all the 10 positive PCR products imply structural and functional conservation. Phylogenetic analysis revealed that minimal genetic diversity occurred between the strains of D. congolensis from the study. It can also be deduced that of the 10 strains of D. congolensis sequenced, two can still be accommodated in the previously listed strains available in the GenBank while the remaining eight may be different strains of D. congolensis.
Acknowledgement
The authors appreciate the generous technical assistance of Mr Akinduti P. A, Miss Shomake A. O and Mrs Ayinde O. O. We also value, in no small measure, contributions from the other members of the Veterinary Microbiology laboratory, Federal University of Agriculture, Abeokuta. This includes Dr. Durosimi O. E, Dr. Opeyi O. O and Dr. Omoshaba E. O. Efforts from Dr. Famuyide I. M and Dr. Anifowose I. O during the latter part of this study are highly commendable.
Source of funding
This research was personally funded. No fund was obtained from an external source.
Conflicts of interest
The authors declare no conflicts of interest in the conduct of this research.
Contributions
Fatai S. Oladunni contributed in sample collection, sample processing, data analysis and manuscript write up and revision. Mufutau A. Oyekunle, Adewale O. Talabi, Olufemi E. Ojo, Michael I. Takeet, Mohammed Adam, and Ibrahim A. Raufu all contributed in data analysis, manuscript write up and revision.
References
- Altschul S.F., Gish W., Miller W., Myers E.W. & Lipman D.J. (1990) Basic local alignment search tool. Journal of Molecular Biology215, 403–410.
- Ambrose N., Lloyd D.H. & Maillard J.C. (1999) Immune responses to Dermatophilus congolensis infections. Parasitology Today15, 295–300.
- Aning K.G. & Koney E.B.M. (1996) Chemotherapy of dermatophilosis-A preliminary study. Tropical Animal Health and Production28, 39–43.
- Gordon M.A. (1976) Characterisation of Dermatophilus congolensis its activities with the Actinomycetales and differentiation from GeoDermatophilus. In: Dermatophilus Infection in Animals and Man (eds by D.H. Lloyd , K.C. Sellers), pp 49, Academic Press, London.
- Han W., Chen Y., Wang J., Wang Y., Yan G. (2007) Establishment of Polymerase Chain Reaction Assay for Detection of Dermatophilosis in Sheep. Available at: http://www.paper.edu.Cn.
- Hirsh D.C., Maclachlan N.J. & Walker R.L. (2004) Veterinary Microbiology. 2nd edn, pp 220–221. Black Well Publishing Company: Malden, MA, USA.
- Macadam I. & Haalstra R.T. (1971) Bacteriology of Nigerian strains of Dermatophilus congolensis. Tropical Animal Health and Production3, 225–231.
- Makinde A.A., Majiyagbe K.A., Orhue P. & Garba A. (1991) Serodiagnosis of dermatophilosis IV. Antigenic selection for optimal antibody response determination by counter immune electrophoresis. Nigerian Journal of Immunology5, 56–60.
- Makinde A.A., Egwuekwe C., Ezeh A.O.I., Calla C.A., Okafor I. & Garba A. (2001) Survival of Dermatophilus congolensis under laboratory conditions in Nigeria. Veterinary Record149, 154–156.
- OIE (2008) Manual of Diagnostic Tests and Vaccine for Terrestrial Animals: Dermatophilosis, 5th edn, pp 725–727. Available at: http://www.oie.int/Dermatophilosis.
- Pal M. & Matsusaka N. (1993) Cutaneous streptothricosis in a fallow deer (Dama dama). Verhandlungsbericht des Erkrankungen des Zootiere35, 167–170.
- Shaibu S.J., Kazeem H.M., Abdullahi U.S. & Fatihu M.Y. (2010) The use of polymerase chain reaction in the diagnosis of dermatophilosis from cattle, sheep and goats in Nigeria. Journal of Animal and Veterinary Advances6, 1034–1036.
- Shaibu S.J., Kazeem H.M., Abdullahi U.S. & Fatihu M.Y. (2011) Phenotypic and genotypic characterization of isolates of Dermatophilus congolensis from cattle, sheep and goats in Jos, Nigeria. African Journal of Microbiology Research5, 467–474.
- Van Sacegham R. (1934) La Dermatose ditto contagieuse des bovines. Bulletin of Agriculture25, 590–598.
- Zaria L.T. (1993) Dermatophilus congolensis infection (Dermatophilosis) in animals and man: an update. Comparative Immununology, Microbiology and Infectious Diseases163, 179–222.
Document information
Published on 01/01/2017
Licence: Other
Share this document
claim authorship
Are you one of the authors of this document?