Abstract
Acrylamide is known to produce follicular cell tumors of the thyroid in rats. RccHan Wistar rats were exposed in utero to a carcinogenic dose of acrylamide (3 mg/Kg bw/day) from gestation day 6 to delivery and then through their drinking water to postnatal day 35. In order to identify potential mechanisms of carcinogenesis in the thyroid glands, we used a transcriptomics approach. Thyroid glands were collected from male pups at 10 PM and female pups at 10 AM or 10 PM in order to establish whether active exposure to acrylamide influenced gene expression patterns or pathways that could be related to carcinogenesis. While all animals exposed to acrylamide showed changes in expected target pathways related to carcinogenesis such as DNA repair, DNA replication, chromosome segregation, among others; animals that were sacrificed while actively drinking acrylamide-laced water during their active period at night showed increased changes in pathways related to oxidative stress, detoxification pathways, metabolism, and activation of checkpoint pathways, among others. In addition, thyroid hormones, triiodothyronine (T3) and thyroxine (T4), were increased in acrylamide-treated rats sampled at night, but not in quiescent animals when compared to controls. The data clearly indicate that time of day for sample collection is critical to identifying molecular pathways that are altered by the exposures. These results suggest that carcinogenesis in the thyroids of acrylamide treated rats may ensue from several different mechanisms such as hormonal changes and oxidative stress and not only from direct genotoxicity, as has been assumed to date.
Abbreviations
ADA , adenosine Deaminase ; ADRB2 , adrenergic ; ASF1B , anti-Silencing Function 1B Histone Chaperone ; BRIP1 , BRCA1 Interacting Protein C-Terminal Helicase 1 ; BUB1B , BUB1 Mitotic Checkpoint Serine/Threonine Kinase B ; C1QTNF3 , C1q and Tumor Necrosis Factor Related Protein 3 ; C5 , complement Component 5 ; CALCR , calcitonin receptor ; CARD9 , caspase recruitment domain family ; CCNA2 , cyclin A2 ; CCNG1 , cyclin G1 ; CD45 , protein tyrosine phosphatase ; CD46 , CD46 molecule ; CDC45 , cell division cycle 45 ; CDCA2 , cell division cycle associated 2 ; CDCA5 , cell division cycle associated 5 ; CENPT , centromere protein T ; CFB , complement factor B ; CGA , glycoprotein hormones ; CTLA4 , cytotoxic T-lymphocyte-associated protein 4 ; DAD1 , defender against cell death 1 ; DCTPP1 , DCTP pyrophosphatase 1 ; DNMT3A , DNA (cytosine-5-)-methyltransferase 3 alpha ; DUOX2 , dual oxidase 2 ; GCG , glucagon ; GCLC , glutamate-cysteine ligase ; GOLGA3 , golgin A3 ; GSTM1 , glutathione S-transferase Mu 1 ; GSTP1 , glutathione S-transferase Pi 1 ; HPSE , heparanase ; HSPA5 , heat shock 70 kDa protein 5 ; HSPB1 , heat shock 27 KDa protein ; HSPB2 , heat shock 27 kDa protein 2 ; HSPH1 , heat shock 105 kDa/110 kDa protein 1 ; HTATIP2 , HIV-1 tat interactive protein 2 ; ID1 , inhibitor of DNA binding 1 ; IGF2 , Insulin-like growth factor 2 (somatomedin A) ; IL1B , interleukin 1 ; INHBA , inhibin ; IYD , iodotyrosine deiodinase ; KIF20B , kinesin family member 20B ; KIF22 , kinesin family Member 22 ; KLK1 , kallikrein 1 ; LAMA2 , laminin, alpha 2 ; MCM8 , minichromosome maintenance complex component 8 ; MIF , macrophage migration inhibitory factor ; MIS18A , MIS18 kinetochore protein A ; NDC80 , NDC80 kinetochore complex component ; NPPC , natriuretic peptide precursor C ; NPY , neuropeptide ; NUBP1 , nucleotide binding protein 1 ; ORC1 , origin recognition complex ; PDE3A , phosphodiesterase 3A ; PINK1 , PTEN induced putative kinase 1 ; PLCD1 , phospholipase C ; PLK1 , polo-like kinase 1 ; POMC , proopiomelanocortin ; PRL , prolactin ; PTGIS , prostaglandin I2 (prostacyclin) synthase ; PTGS1 , prostaglandin-endoperoxide synthase 1 ; PRKAA2 , protein kinase ; PRODH , proline dehydrogenase ; RAB5A , RAB5A ; RAN , ras-related nuclear protein ; RRM2 , ribonucleotide reductase M2 ; SCL5A5 , solute carrier family 5 (sodium iodide symporter) ; SELP , selectin P (granule membrane protein 140 kDa ; SPAG8 , sperm associated antigen 8 ; TACC3 , transforming ; TBCB , tubulin folding cofactor B ; TFRC , transferrin receptor ; TOP2A , topoisomerase (DNA) II alpha ; TPO , thyroid peroxidase ; TSHR , thyroid stimulating hormone receptor ; TSN , translin ; VWF , Von Willebrand Factor
Keywords
Acrylamide ; RccHan Wistar ; Transcriptomics ; Thyroid
1. Introduction
Acrylamide (AA) is a monomer used in the manufacture of polymers for mining, oil and natural gas processing, paper manufacture, waste processing, hospital laboratories, among other uses. Adverse health effects from worker exposure to AA have been extensively studied [44] . No adverse effects have been reported with daily human exposure up to 2.1 mg/kg/day [19] . Exposure to AA in foodstuffs has become a worldwide concern because of its generation in a variety of carbohydrate rich foods when these are cooked at temperatures exceeding 120 °C. At these temperatures, AA is made from the Maillard reaction of sugars with asparagine residues [24] and [55] .
The World Health Organization (WHO) and Food and Agriculture Organization [22] , the Environmental Protection Agency [18] , and the European Food Safety Authority [16] have classified AA as a “probable human carcinogen” by virtue of its conversion to glycidamide, as the ultimate carcinogen in rodents. In vivo , AA is metabolized by CYP 2E1 to glycidamide (an epoxide), which also is very reactive toward nucleophiles and has been implicated in adduct formation at active site cysteines of proteins and the amino terminus of hemoglobin and also with nucleic acids [24] and [47] . Only a portion of the AA gets converted to glycidamide. The classification as a probable human carcinogen is based primarily upon reproducible carcinogenicity studies in Fischer rats [2] , [26] and [34] . Tumors were observed in the mammary gland (fibroadenomas), thyroid (follicular tumors) and tunica vaginalis testes in rats [2] , [26] and [33] . These tumor sites have well documented rat-specific modes of action, which are not relevant to humans [3] , [31] , [56] and [59] .
Thyroid follicular tumors can arise in rats specifically from chemical disturbance of thyroid-pituitary homeostasis, or from a combination of genotoxicity and hormonal alterations. External factors that change thyroid hormone metabolism result in a marked change in thyroid behavior in rats [36] . AA treatment for 7, 14 or 28 days results in increased mitotic figures and Dfigures and DNA synthesis in the thyroids of Fischer rats [43] . The Fischer rat has a unique hormonal milieu that may be responsible for these tumors. However, specific changes in thyroid transcriptomics have yet to be documented. It has been suggested that alteration of the dopamine receptor is responsible for the change in thyroid metabolism [65] . AA also causes oxidative stress in rats [72] and the thyroid has a very high oxidation level [72] and [73] .
AA is mutagenic in rats and mice [12] . This mutagenic profile does not fit the pattern of tumors observed in Fischer rats, as the specificity cannot be explained [27] , [52] , [65] and [74] . AA is a very weak carcinogen and mutagen [74] . Since mutagenicity is the regulatory default assumption as a mechanism for carcinogenicity, use of other mechanistic data for regulatory purposes has been limited. In contrast to the Fischer rat, the only significant tumors observed in male Wistar rats were in the thyroid gland [51] . While thyroid tumors in rats are not considered to be relevant to humans, mechanisms for genotoxicity in the thyroid still represent a possible significant risk. This study was designed to build upon the large volume of data published to date about AA exposure of rats to determine whether tumors formed in the thyroid after chronic exposure to AA were produced only through a genotoxic component.
The research conducted here determines the effect of AA on gene expression in weanling Wistar rat thyroids following in utero exposure to a carcinogenic dose of AA. This research is intended to differentiate between DNA reactivity, oxidative stress and direct action on hormone receptors. A transcriptomics approach (microarrays and bioinformatics) was used to investigate changes in gene expression and the association with physiological responses (changes in plasma hormone levels) in rats treated with AA from gestational day 6 to post-natal day 35. We hypothesized that 3 mg AA/Kg bw/day exposure would produce changes in plasma hormone levels associated with key genes and biochemical pathways involved with molecular actions of AA.
2. Materials and methods
2.1. Test material
AA (C3H3NO, CAS no 79-06-1, 1,2-propenamide; >99.9% pure; Sigma Aldrich) was dissolved in tap water and evaluated for stability at room temperature at 6, 13, 20, and 27 test days after preparation. Recovery ranged from 96.9% to 102.6%.
2.2. Animal exposures
The methods used to conduct this study have been previously published [51] . The in vivo phase of this study was conducted under GLP guidelines and was externally audited. It was approved by the responsible local government office according to the German animal welfare law “Tierschutzgesetz” (TierSchG). AA solutions were prepared weekly and concentrations were adjusted for body weight. Water bottles were changed weekly. AA concentration in the drinking water was determined at test week 4 and 10.
Sperm positive female Wistar Han™/RccHan™:WIST rats were obtained from Harlan Laboratories GmbH, Serumweg 48, 27324 Eystrup, Germany in multiple deliveries. At gestation day 6, dams were provided AA in their drinking water. Exposures continued in F1 offspring through postnatal day (pnd) 35 ± 3. Rats were housed 1 per cage in MACROLON cages with granulated wood bedding (Brandenburg, 49424 Goldenstedt/Arkeburg). Animal rooms were alternately lit (about 150 lx at approximately 1.50 m room height) and darkened in a 12-hour lighting cycle. Cage side observations were conducted twice per day during the week and once per day on weekends.
On day 4 after birth, the weights of the pups were determined. The size of each litter was adjusted by eliminating extra pups to yield, as nearly as possible, five males and five females per litter and remaining animals remained with the dams until day 21 of lactation (weaning). On lactation day 21, the F1 animals were randomized using a computer randomization program to assign the animals to the subsets within each group.
A separate cohort of the animals was allowed to continue on the same regimen for two years. At the end of the two years, mammary gland fibroadenomas were identified in females and thyroid follicular cell tumors were identified in both sexes [51] .
2.3. Plasma TSH, T3 and T4 analysis
On pnd 35 ± 3, at approximately 10 AM and 10 PM, respectively, as much blood as possible was withdrawn from 5 male and 5 female F1 rats/dose. The thyroids were removed and frozen. Blood samples were divided into 5 aliquots. Four aliquots of at least 75 μL were frozen and stored at −20 °C. Two aliquots were analyzed using the rat pituitary panel from Millipore for adrenocorticotropic hormone (ACTH), brain-derived neurotrophic factor (BDNF), growth hormone (GH), follicle-stimulating hormone (FSH), luteinizing hormone (LH), and prolactin. Then two aliquots were analyzed using the rat thyroid milliplex kit from Millipore for T3, T4 and TSH.
2.4. Extraction of total RNA and RNA quality control
Total RNA was extracted from the thyroid gland of rats (n = 4; 4 controls and 4 AA-treated per tissue) using RNA STAT-60 reagent (Tel-Test Inc., Friendswood, TX, USA) according to the manufacturer’s instructions. A total of 500 ng of RNA was DNase-treated with Turbo DNA-free (Ambion Austin, TX) following the manufacturer’s protocol. RNA quantity for microarray analysis was measured using the NanoDrop ND-1000 (Nanodrop Technologies, Wilmington, DE) and RNA quality was evaluated using the Agilent 2100 BioAnalyzer with the RNA 6000 Nanochip. RNA integrity values (RIN) were >8.0 for all samples used in the analysis. Total RNA was used for microarray analysis and real-time quantitative PCR (qPCR).
2.5. Microarray analysis
Gene expression patterns of thyroid samples were identified using the Rat V1 8 × 60 K oligonucleotide microarray manufactured by Agilent (Palo Alto, CA, USA). 100 ng of total RNA per sample (n = 4 biological replicates for each condition) were used for the analysis following the manufacturer’s kits and protocols (Agilent Low input Quick-amp labeling kit one color; Agilent, Santa Clara, CA, USA). All samples used for microarrays contained a specific activity >9.0 pmol Cy3/ml and amounts were adjusted to a final mass of 600 ng. Microarrays were kept in the dark until scanning using an Agilent G2505B microarray scanner. Data extraction was performed using Agilent Feature Extraction software (v 9.5). All microarrays were submitted to NCBI’s Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/ ) with accession number (GSE 62026). We had technical issues with samples from male rats collected in the morning and for this reason, these results are not being presented here.
2.6. Bioinformatics
The microarray data were analyzed independently by two different groups using JMP Genomics v5 (SAS, Cary, NC, USA) and by open source R-based software packages. The microarrays were performed in two different batches, one year apart, and it was impossible to analyze both batches in one ANOVA because of significant batch effects. Thus, each of the batches was analyzed separately and comparisons at the pathway level were possible between the two batches.
In JMP Genomics, raw expression data were log2-transformed and all control and non-uniform spots were removed prior to the final analysis. All probe responses per gene were averaged before using median normalization. One-way ANOVA was used to find genes that were significantly altered (p < 0.05; fold change greater than ±1.2). The list of genes from the ANOVA was subjected to hierarchical clustering for each batch of arrays independently. The arrays were also analyzed by other non-parametric algorithms including Gene Set Enrichment Analysis (GSEA) [69] and Fischer Exact Test with the Sub-Network Enrichment Analysis (SNEA) [40] available through PathwayStudio™ (Elsevier Inc., Philadelphia, PA, USA), described in more detail below.
The second analysis used R-based software packages in a step-wise process consisting of a quality assessment of the array data, processing of the arrays for background correction and group normalization, unsupervised clustering, and class comparison for differential expression. The software components required to execute the source code include the following: (1) R environment for statistical programming (R package Limma) [67] ; (2) Several package libraries from BioConductor [35] ; (3) R markdown text formatting system; and (4) Knitr engine for dynamic report generation with R. Statistical criteria for identification of differentially expressed genes were false discovery rate (FDR) adjusted P* < 0.05 and fold change greater than 1.5. Based upon these filter criteria no differential gene expression was observed between animals exposed to AA and controls. Further analysis with FDR adjusted P* < 0.10 resulted in 1 gene being differentially expressed between AA treated animals and controls (Slc35b1, log FC = 1.6, P* = 0.05, solute carrier family 35, member B1).
2.7. PathwayStudio™ analysis
Pathway Studio™ V9 (operating with the ResNet 9.0 database) was used to identify molecular pathways associated with chronic toxicity of AA. Specifically two datasets were generated using (1) Fisher’s Exact Test with (p -value < 0.05) and SNEA and (2) GSEA, using the Kolmogorov-Smirnov test (p < 0.05) [40] and [69] . GSEA is an analytical method used to evaluate microarray data at the level of gene sets, which are defined based on prior biological knowledge of gene ontology and curated pathways [40] and [69] .
2.8. Quantitative real-time PCR (qPCR)
Quantitative reverse transcriptase polymerase chain reaction (qPCR) was performed in order to confirm some of the results of differential gene expression identified by microarray analysis. We selected five different genes for the morning animals and five for the night animals and used TaqMan probes designed and validated by Applied Biosystems (Foster City, CA, USA). Gene symbols and ID of the TaqMan assays used are identified in Table 1 .
| Night group genes | |||
|---|---|---|---|
| Gen bank Accession | Gene Name | Gene Symbol | IDa |
| NM_022303 | Caspase recruitment domain family, member 9 | Card9 | Rn00673582_m1 |
| NM_012547 | Dopamine receptor D2 | Drd2 | Rn00561126_m1 |
| NM_080906 | DNA-damage-inducible transcript 4 | Ddit4 | Rn01433735_g1 |
| NM_019353 | Thyroid peroxidase | Tpo | Rn00571159_m1 |
| NM_001009623 | Tumor necrosis factor (ligand) superfamily, member 13 | Tnfsf13 | Rn01467490_g1 |
| Morning group gene | |||
| NM_053749 | Aurora kinase B | Aurkb | Rn01460656_m1 |
| NM_024141.1 | Dual oxidase 2 | Duox2 | Rn01514628_m1 |
| NM_001106335.1 | Inner centromere protein | Incenp | Rn01478880_g1 |
| NM_139326.2 | Proopiomelanocortin | Pomc | Rn00595020_m1 |
| NM_022183 | Topoisomerase (DNA) II alpha | Top2a | Rn00573347_m1 |
| Normalizing Gene | |||
| NR_046239.1 | 45S pre-ribosomal RNA | 18S | Rn03928990_g1 |
a. TaqMan assays ID (Applied Biosystems).
We used the same total RNA as used for microarrays. First-strand cDNA was synthesized from 250 ng DNAse treated total RNA using 250 ng random primers and SuperScript™ II Reverse Transcriptase (Invitrogen/Life Technologies, Grand Island, NY, USA) in a 20 μL reaction, as per the manufacturer’s protocol. qPCR analysis was performed in an ABI 7500 Fast System instrument (Applied Biosystems, Foster City, CA, USA). In brief, a mixture was made with 1 μL cDNA template (diluted to 100 ng/L), 0.5 μL of each probe from the specific TaqMan assay (20X), 1 μL of a 2X TaqMan Fast Advanced master mix and 3 μL Nuclease-free water in 10 μL total volume per reaction, as indicated in the manufacturer’s protocol. TaqMan PCR cycling conditions were set at 95 °C for 20 s for the first cycle and 3 s at 95 °C followed by 30 s at 60 °C for the remaining 40 cycles, as indicated in the manufacturer’s protocol. All experimental samples were run in duplicate, along with two negative controls; a “no reverse transcriptase (-RT)” control, in which DNase-treated RNA samples were pooled and water was used in place of reverse transcriptase during the reverse transcription reaction, and a “no template control (NTC), ” in which water was used in place of template cDNA during the real-time PCR reaction. 18S ribosomal RNA was used as the reference gene to normalize expression data. The results were analyzed using the ΔΔCt method of relative quantification [45] .
2.9. Statistical analyses
Significant differences in plasma levels of hormones in AA treated animals were determined using one-way analysis of variance (ANOVA) followed by a Tukey’s Test. For qPCR a t-test was performed on normalized gene expression to evaluate whether the expressions were statistically different compared to controls (p < 0.05).
For both plasma hormone levels and qPCR when data were not normally distributed, the Kruskal–Wallis non-parametric test was followed by the Mann-Whitney U-test. A p -value ≤0.05 was considered statistically significant. Statistical analyses were performed using Sigma Plot (SYSTAT, Chicago, IL, USA) and figures were plotted using Prism 5.0 (GraphPad Software Inc., San Diego CA).
3. Results
3.1. Phenotypic anchoring: measurement of plasma hormones in treated and control rats
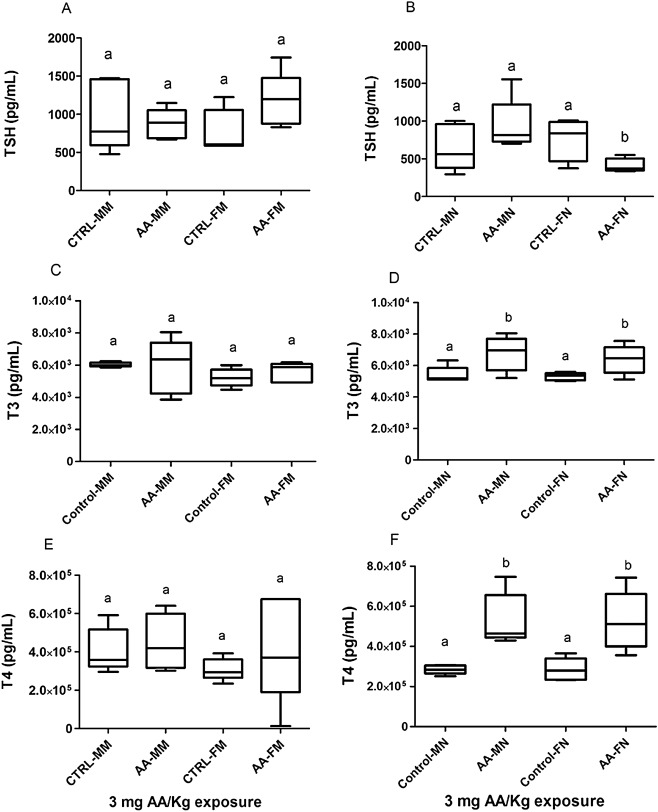
Changes in plasma hormone concentrations in pnd 35 rats occurred mainly in nocturnal animals of both sexes. Male rat plasma collected at night showed significant increases in T4, T3 and pituitary ACTH by 87%, 24.6% and 223%, respectively compared to controls. In females collected at night, plasma TSH was reduced by 44.7% and T4 and T3 were increased by 84.6% and 20.2%, respectively, compared to controls. The changes in T3 and T4 levels in female and male rats sampled at night closely matched each other (Fig. 1 ). Prolactin was only decreased in male rats sampled in the morning with levels decreased by 76% (data not shown). Other hormones tested were not significantly altered in any of the animals.
|
|
|
Fig. 1. Plasma levels of thyroid hormones and TSH in animals collected in the morning and at night (n = 5 per group). (A, C, E) Animals sampled in the morning; (B, D, F) animals sampled at night with (A, B) TSH; (C, D) T3; and (E, F) T4. Box plots indicate the upper and lower percentile (10–90th) and the median values (horizontal black line within box). Whiskers represent the highest and lowest points. Different letters indicate significant differences among groups. CTRL (control group); AA (Acrylamide-treated group); MM (male morning); FM (female morning); MN (male night); FN (female night). |
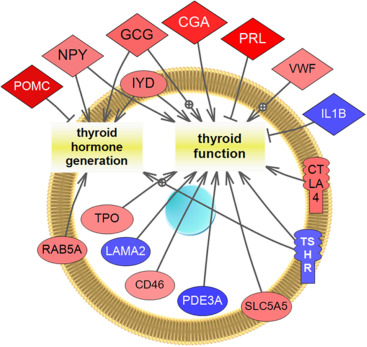
Interestingly, only night animals showed changes in genes related to thyroid hormone biosynthesis and function. Genes pertaining to this pathway that were changed in males sampled at night are shown in Fig. 2 . Fewer genes in this pathway were changed in females sampled at night.
|
|
|
Fig. 2. Changes in gene expression that were related to thyroid hormone generation and thyroid function in male rats sampled at night. Pathways were built by sub-networks enrichment analysis (p-value >0.05; fold change > ± 1.2; statistically analyzed by Fisher’s Exact Test) using PathwayStudio™. Blue indicates down-regulation of gene expression; red, up-regulation; and yellow indicates the processes likely affected. Color intensity correlates with the degree of response. The abbreviations of the genes are listed in the abbreviations list. |
3.2. Gene expression changes in male and female rats
Microarray analysis was performed separately on thyroids of animals collected in the morning or at night. ANOVA results indicated a large number of differentially regulated genes for rats exposed to AA. Male rat thyroids collected at night exhibited 1800 altered genes (1157 up-regulated and 642 down-regulated). With an FDR correction of 10%, only one gene (solute carrier family 35, member B1 (Slc35b1) was significantly altered in this group. There were procedural issues for the microarrays for males collected in the morning and for that reason the results are not presented here.
Female thyroids sampled at night presented 2221 genes (1155 up-regulated and 1066 down-regulated), whereas female thyroids collected in the morning presented 2793 genes (854 up-regulated and 1939 down-regulated). Application of a FDR at 5% showed no altered genes. In order to get pathway information, we broadened the criteria to include genes that were altered with a p value <0.05 and fold expression > ±1.2 and an FDR was not applied. Changes in gene expression are shown in Table SI-1.
The differentially regulated transcripts were ordered according to their fold-expression levels in males sampled at night (MN) followed by their expression levels in females sampled at night (FN), and then females sampled in the morning (FM) (Fig SI-1), keeping the same order of genes. This evaluation showed that many genes were significantly altered in the same direction in both male and female samples collected at night, but some of these same genes were altered in different directions in females collected in the morning. A partial list of these common genes can be found in Table 2 .
| Probe Name | Gen bank Accession | Gene Symbol | Gene Name | Fold Change (p ≤ 0.05 ) | |
|---|---|---|---|---|---|
| Male | Female | ||||
| Thyroid hormone | |||||
| A_64_P076450 | NM_139326 | Pomc | Proopiomelanocor-tin | 18.4 | 6.3 |
| A_64_P155193 | NM_021653 | Dio1 | Deiodinase, iodothyronine, type I | 1.4 | 1.6 |
| A_42_P625922 | NM_012547 | Drd2 | Dopamine receptor | 5.2 | 2.4 |
| A_44_P238257 | NM_053920 | Trip10 | Thyroid hormone receptor interactor 10 | −1.5 | −1.3 |
| Processing and protein transport | |||||
| A_43_P16457 | NM_001037208 | Creld2 | Cysteine-rich with EGF-like domains 2 | 2.2 | 2.1 |
| A_64_P056846 | NM_012815 | Gclc | Glutamate-cysteine ligase, catalytic subunit | 2.0 | 1.4 |
| A_64_P021433 | NM_001007755 | Scly | Selenocysteine lyase | 1.7 | 1.3 |
| Motor proteins | |||||
| A_42_P757258 | NM_001107609 | Kif20b | Kinesin family member 20B | 1.8 | 1.3 |
| A_64_P108200 | NM_001009645 | Kif22 | Kinesin family member 22 | 1.7 | 1.2 |
| A_64_P111749 | XM_001057533 | Kif2a | Kinesin family member 2A | 1.7 | 1.9 |
| A_44_P163018 | NM_001009619 | Nubp1 | Nucleotide binding Protein 1 | 1.3 | 1.2 |
| A_44_P463313 | NM_017100 | Plk1 | polo-like kinase 1 | 1.5 | 1.3 |
| A_44_P213149 | NM_001107369 | Mastl | Microtubule associated serine/threonine kinase-like | 2.3 | 1.6 |
| Detoxifying enzymes/Oxidative Stress | |||||
| A_64_P137037 | NM_017156 | Cyp2b12 | Cytochrome P450, family 2, subfamily b, polypeptide 12 | 1.4 | 2.0 |
| A_64_P004231 | NM_001159739 | Gsta5 | Glutathione S-transferase Yc2 subunit | 2.5 | 1.5 |
| A_42_P678430 | NM_022303 | Card9 | Caspase recruitment domain family, member 9 | 1.6 | 1.4 |
| A_64_P119916 | NM_024141 | Duox2 | Dual oxidase 2 | 1.7 | 1.2 |
| A_64_P095830 | NM_152242 | Gpr56 | G protein-coupled receptor 56 | 1.6 | 1.3 |
| A_42_P636627 | NM_017110 | Cartpt | CART prepropeptide | 1.6 | 2.4 |
| A_64_P069374 | NM_017247 | Scn10a | Sodium channel, voltage-gated, type X, alpha | 3.1 | 1.9 |
| A_64_P035564 | NM_001108682 | Tlr12 | Toll-like receptor 12 | 1.6 | 1.8 |
| Checkpoint pathways and cancer | |||||
| A_64_P012009 | NM_001106787 | Mov10L1 | Mov10L1, Moloney leukemia virus 10-like 1, homolog (mouse) | 3.2 | 2.4 |
| A_44_P478066 | NM_001106335 | Incenp | Inner centromere protein | 1.4 | 1.4 |
| A_64_P033214 | XM_346406 | Top2a | Topoisomerase (DNA) II alpha | 1.5 | 1.5 |
| A_43_P10723 | XM_001080736 | Bub1b | Budding uninhibited by benzimidazoles 1 homolog, beta | 1.7 | 1.4 |
| A_44_P1045354 | NM_199081 | Slc35b1 | Solute carrier family 35, member B1 | 1.9 | 1.3 |
These genes are involved in processes including, xenobiotic metabolism, motor proteins, oxidative stress, and cell cycle checkpoint pathways, among others. Genes that were commonly altered in females collected in the morning and at night are presented in Table 3 . Remarkably, females collected in the morning also showed a large group of altered genes that were not altered in animals collected at night. Some of these genes are in the same families as genes altered at night and their non-correspondence may be due to the use of different microarrays for these analyses. Thus, the best way to jointly analyze these data is at the pathway level rather than at the specific gene level.
| Fold Change (p < 0.05) | |||||
|---|---|---|---|---|---|
| Probe Name | Gen bank Accession | Gene Symbol | Gene Name | AM | PM |
| Up-regulated | |||||
| A_44_P533786 | NM_053749 | Aurkb | Aurora kinase B | 2.8 | 1.4 |
| A_44_P461544 | NM_001009470 | Ccnb2 | Cyclin B2 | 2.8 | 1.5 |
| A_44_P534089 | NM_171991 | Ccnb1 | Cyclin B1 | 2.7 | 1.4 |
| A_42_P535608 | NM_001107160 | Asf1b | ASF1 anti-silencing function 1 homolog B (S. cerevisiae) | 2.6 | 1.4 |
| A_44_P189375 | NM_001001719 | Fancd2 | Fanconi anemia, complementation group D2 | 2.6 | 1.5 |
| A_44_P381917 | NM_133386 | Sphk1 | Sphingosine kinase 1 | 2.5 | 1.7 |
| A_44_P223446 | NM_001107873 | Mcm2 | Minichromosome maintenance complex component | 2.4 | 1.4 |
| A_44_P478066 | NM_001106335 | Incenp | Inner centromere protein | 2.3 | 1.4 |
| A_44_P213149 | NM_001107369 | Mastl | Microtubule associated serine/threonine kinase-like | 2.3 | 1.6 |
| A_64_P075910 | NM_001101014 | Ajap1 | Adherens junction associated protein 1 | 2.2 | 1.6 |
| A_64_P129618 | NM_001017459 | Mdm1 | Mdm1 nuclear protein homolog (mouse) | 2.2 | 1.2 |
| A_42_P643574 | NM_001106795 | Aaas | Achalasia, adrenocortical insufficiency, alacrimia (Allgrove, triple-A) | 1.7 | 1.2 |
| A_42_P502590 | NM_001107424 | Slc7a6 | Solute carrier family 7 (cationic amino acid transporter, y+ system), member 6 | 1.6 | 1.2 |
| A_64_P097193 | XM_001072207 | Topbp1 | Topoisomerase (DNA) II binding protein 1 | 1.5 | 1.2 |
| Down-regulated | |||||
| A_64_P008477 | NM_001000247 | Olr325 | Olfactory receptor 325 | −1.4 | −1.2 |
| A_42_P463998 | NM_031013 | Abcc6 | ATP-binding cassette, sub-family C (CFTR/MRP), member 6 | −1.5 | −2.0 |
| A_64_P029596 | NM_001102417 | Svs3b | Seminal vesicle secretory protein 3B | −1.6 | −3.0 |
| A_44_P302179 | XM_001070842 | Naip5 | NLR family, apoptosis inhibitory protein 5 | −1.6 | −1.4 |
| A_44_P612186 | NM_001107344 | Mylip | Myosin regulatory light chain interacting protein | −1.6 | −1.2 |
| A_44_P279116 | XM_002726450 | Dyrk4 | Dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 4 | −1.6 | −1.3 |
| A_64_P083349 | NM_021843 | Kitlg | KIT ligand | −1.6 | −1.3 |
| A_64_P010648 | NM_001105822 | Ccl12 | Chemokine (C-C motif) ligand 12 | −1.7 | −1.4 |
| A_64_P008330 | NM_001000020 | Olr1454 | Olfactory receptor 1454 | −1.7 | −1.3 |
| A_44_P1000391 | NM_001106140 | Atp8b1 | ATPase, Class I, type 8B, member 1 | −1.8 | −1.2 |
| A_64_P050650 | XM_344594 | Sox4 | SRY (sex determining region Y)-box 4 | −1.9 | −1.3 |
| A_64_P127823 | NM_001108629 | Ggct | Gamma-glutamyl cyclotransferase | −1.9 | −1.2 |
| A_44_P684740 | NM_001107279 | Pcdh17 | Protocadherin 17 | −2.3 | −1.4 |
| A_64_P008502 | NM_001109574 | Tmem169 | Transmembrane protein 169 | −2.7 | −1.5 |
| A_43_P12257 | NM_022604 | Esm1 | Endothelial cell-specific molecule 1 | −2.9 | −1.6 |
3.3. Common alterations of the transcriptome for rats of both genders
The genes selected by ANOVA were used for SNEA. Several sub-networks were common to all of the animals tested (Table 4 ). Common pathways altered in both males and females at the lowest p values (<8.0E-07) included cell proliferation, cell differentiation, apoptosis, cell death, cell migration, cell growth, cell cycle, protein folding, and myogenesis, among others. The top 100 predicted pathways via SNEA for animals sampled at night males had a p value <0.002 for males and <0.009 for females (Table SI-2). However, it is clear from Table 5 that the top gene sets altered in females sampled at night were different from those altered in females in the morning.
| Sub-networks | p-value (<0.05) | ||
|---|---|---|---|
| Gene Set Seed | Male at night | Female at night | Female at morning |
| Cell proliferation | 8.3E-14 | 1.1E-10 | 9.4E-11 |
| Cell differentiation | 2.0E-11 | 1.9E-10 | 5.4E-10 |
| Apoptosis | 3.4E-10 | 9.2E-15 | 4.1E-13 |
| Cell death | 4.9E-10 | 7.6E-09 | 1.3E-07 |
| Cell migration | 6.3E-09 | 5.7E-06 | 1.2E-06 |
| Vascularization | 2.7E-08 | 1.7E-03 | 3.2E-07 |
| Cell growth | 5.0E-08 | 2.5E-08 | 6.1E-11 |
| Cell cycle | 9.2E-08 | 6.3E-08 | 9.1E-07 |
| Pregnancy | 7.3E-07 | 3.7E-03 | 2.3E-03 |
| Oxidative stress | 7.6E-07 | 8.6E-04 | 7.9E-06 |
| Cell survival | 1.7E-06 | 5.1E-05 | 1.7E-05 |
| Endocytosis | 2.3E-06 | 5.2E-04 | 4.5E-07 |
| Heart function | 4.6E-06 | 6.3E-03 | 6.9E-03 |
| Contraction | 1.4E-05 | 2.3E-03 | 4.9E-03 |
| G2/M transition | 1.7E-05 | 1.4E-05 | 2.6E-04 |
| Morphogenesis | 2.1E-05 | 1.9E-06 | 2.4E-03 |
| Ossification | 3.6E-05 | 1.2E-04 | 1.8E-02 |
| Inflammatory response | 6.2E-05 | 4.7E-03 | 4.9E-03 |
| Protein folding | 2.6E-04 | 1.0E-03 | 1.5E-02 |
| Myogenesis | 1.4E-04 | 8.6E-04 | 4.0E-04 |
| DNA replication | 5.0E-04 | 8.9E-04 | 2.8E-03 |
| Response to hypoxia | 9.8E-04 | 8.9E-03 | 5.0E-03 |
| Gene Set Seed | p-value (FN) | p-value (FM) |
|---|---|---|
| Mitosis | 8.05E-07 | NE |
| Embryonal development | 6.85E-06 | NE |
| M phase | 4.61E-05 | NE |
| Kinetochore assembly | 5.01E-05 | NE |
| Meiosis | 7.94E-05 | NE |
| Oncogenesis | 1.13E-04 | NE |
| DNA replication checkpoint | 1.30E-04 | NE |
| Macrophage differentiation | 1.67E-04 | NE |
| G2 phase | 1.67E-04 | NE |
| Genetic instability | 1.99E-04 | NE |
| G2/M checkpoint | 2.60E-04 | NE |
| Regeneration | 2.83E-04 | NE |
| Response to stress | 3.52E-04 | NE |
| Mitotic entry | 4.76E-04 | NE |
| Mitotic checkpoint | 4.87E-04 | NE |
| Muscle development | 4.98E-04 | NE |
| Fibroblast accumulation | 5.22E-04 | NE |
| Muscle regeneration | 5.41E-04 | NE |
| ER-associated protein catabolism | 6.76E-04 | NE |
| Microtubule cytoskeleton assembly | 7.21E-04 | NE |
| Erythrocyte differentiation | 7.79E-04 | NE |
| Nerve maturation | 8.50E-04 | NE |
| Xenobiotic metabolism | 8.61E-04 | NE |
| Mitotic spindle assembly | 9.06E-04 | NE |
| Chromosome condensation | 1.03E-03 | NE |
| Cell adhesion | NE | 1.77E-05 |
| Mitochondrial damage | NE | 2.81E-05 |
| Regulation of cell size | NE | 5.23E-05 |
| Muscle cell differentiation | NE | 9.85E-05 |
| Kidney function | NE | 1.19E-04 |
| Glial cell response | NE | 1.40E-04 |
| Cell-cell adhesion | NE | 1.57E-04 |
| Adipocyte differentiation | NE | 1.61E-04 |
| Wound healing | NE | 4.84E-04 |
| SMC proliferation | NE | 5.90E-04 |
| Neuron apoptosis | NE | 6.80E-04 |
| Luteinization | NE | 8.26E-04 |
| G1 phase | NE | 8.44E-04 |
| Senescence | NE | 8.79E-04 |
| Actin organization | NE | 9.21E-04 |
| Secretory pathway | NE | 1.14E-03 |
| Endothelial cell function | NE | 1.15E-03 |
| Endothelial cell proliferation | NE | 1.22E-03 |
| Myocyte proliferation | NE | 1.26E-03 |
| Colonization | NE | 1.28E-03 |
| Arteriogenesis | NE | 1.39E-03 |
| Embryo implantation | NE | 1.40E-03 |
| Biomineral formation | NE | 1.44E-03 |
| Autophagic cell death | NE | 1.65E-03 |
| Endomitotic cell cycle | NE | 1.74E-03 |
NE; No effect.
When both genders were analyzed together for this time point, the top 12 common pathways altered were cell death, apoptosis, protein folding, kinetochore assembly, decidualization, pregnancy, colorectal motility, muscle development, eating behavior, response to heat shock, oocyte development and heart function (p value <0.0008) (Table SI-3A). When females from the night and morning sampling times were analyzed together to find common pathways, the top pathways included DNA replication, apoptosis, cell growth and cell proliferation, among others (Table SI-3B).
The lists of genes from the microarray analysis were also subjected to a non-parametric statistical analysis, GSEA, which provided additional pathway information. An analysis for pathways for all three groups yielded 10 cellular processes that were commonly altered with DNA replication heading the list (Table 6 ). When considering only animals sampled at night, there were 20 common pathways, including INO80Chromatin Remodeling, TRRAP/Tip60Chromatin Remodeling, mRNA Transcription and Processing, Double Strand DNA Homologous Repair, cell cycle and DNA replication, among others (Table SI-4A and 4B). There was good concordance between the pathways selected via SNEA and the GSEA.
| Gene Set Category: Ariadne Cell Process Pathways | Male at night | Female at night | Female at morning | |||
|---|---|---|---|---|---|---|
| Median change | p-value | Median change | p-value | Median change | p-value | |
| DNA replication | 1.3 | 0.01 | 1.1 | 0 | 1.1 | 0 |
| Histone and DNA methylation | 1.2 | 0.02 | 1.1 | 0 | NE | 0.01 |
| Histone phosphorylation | 1.1 | 0 | 1.1 | 0 | 1.1 | 0 |
| Direct DNA repair | 1.1 | 0.02 | 1.1 | 0 | 1.1 | 0.01 |
| Single-strand mismatch DNA repair | 1.2 | 0 | 1.1 | 0 | 1.1 | 0 |
| Double strand DNA homologous repair | 1.2 | 0 | 1.1 | 0 | 1.1 | 0 |
| Single-strand base excision DNA repair | 1.2 | 0.01 | 1.1 | 0 | 1.1 | 0 |
| Cell cycle | 1.2 | 0 | 1.1 | 0 | 1.1 | 0 |
| Actin cytoskeleton assembly | 1.1 | 0 | NE | 0.02 | 1.2 | 0.01 |
| Tight junction assembly (Occludin) | 1.2 | 0 | 1.1 | 0.03 | NE | 0.01 |
NE; No effect.
Selected pathways graphically show similarities between males and females sampled at night (Fig. 3 , Fig. 4 and Fig. 5 ).
|
|
|
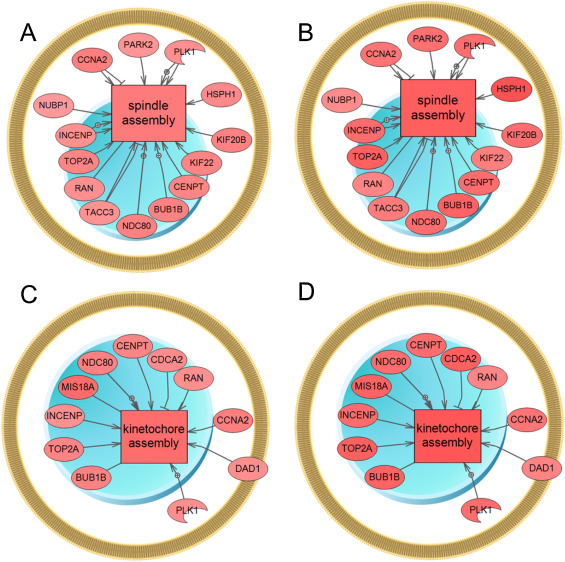
Fig. 3. Alterations in gene expression related to (A–B) spindle assembly and (C–D) kinetochore assembly. These pathways were built using sub-networks enrichment analysis (SNEA) (p-value > 0.05; fold change >± 1.2; by Fisher’s Exact Test) using PathwayStudio™. (A, C) Male rats sampled at night; and (B, D) female rats sampled at night. Blue, down-regulated genes; red, up-regulated genes. Abbreviations for gene names are in the abbreviation list. |
|
|
|
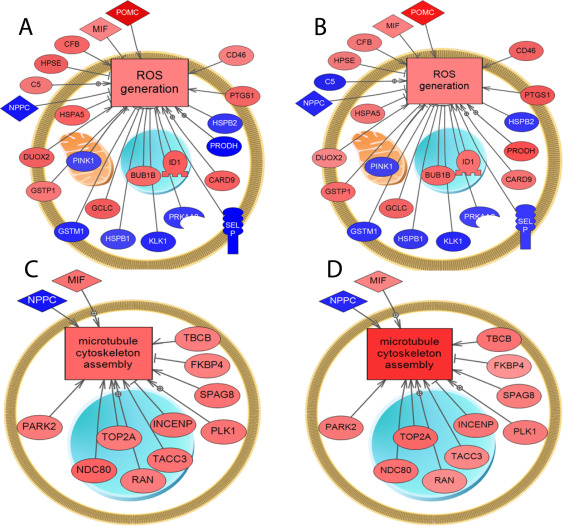
Fig. 4. Alterations in gene expression related to reactive oxygen species (A-B) (ROS) and (C-D) microtubule cytoskeleton assembly. These pathways were built using sub-networks enrichment analysis (SNEA) (p-value > 0.05; fold change >± 1.2; by Fisher’s Exact Test) using PathwayStudio™. (A, C) Male rats sampled at night; and (B, D) female rats sampled at night. Blue, down-regulated genes; red, up-regulated genes. Abbreviations for gene names are in the abbreviation list. |
|
|
|
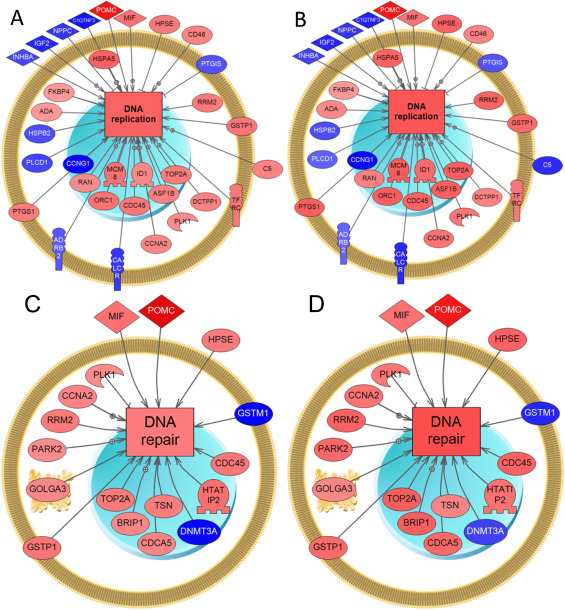
Fig. 5. Genes altered by AA exposure of rats that relate to (A-B) DNA replication and (C-D) DNA repair. These pathways were built using sub-networks enrichment analysis (p-value > 0.05; fold change >± 1.2; by Fisher’s Exact Test) using PathwayStudio™. (A, C) Male rats sampled at night; and (B, D) female rats sampled at night. Blue, down-regulated genes; and red, up-regulated genes. Abbreviations for gene names are in the abbreviation list. |
3.4. Diurnal changes in female rats sampled either at night or in the morning.
Diurnal changes in gene expression could only be observed in female rats, since we had issues with the microarrays for male rats. We observed many more changes in pathways in female rats sampled at night than those sampled in the morning (Table SI-4B and 4C). While both groups showed changes in similar repair mechanisms including DNA repair, protein degradation and recycling, chromatin remodeling, and the immune system; there were many more entities present in the affected pathways in animals sampled at night than those sampled in the morning. However, among the top pathways with the lowest P values (<0.01) by GSEA, 15 were only altered in females collected at night which involved methylation of histones and DNA, chromatin remodeling, mRNA and rRNA transcription and processing, tight junction assembly, and immune function, such as peptide antigen and complement pathways. One pathway was shared by both morning and night animals, which involved ubiquitin-dependent protein degradation, and 1 pathway was altered only in females collected in the morning and this involved actin cytoskeleton assembly (Table 7 ).
| Females | ||||
|---|---|---|---|---|
| Gene Set Category: | Night | Morning | ||
| Ariadne Cell Process Pathways | Median change | p-value | Median change | p-value |
| Histone and DNA methylation | 1.1 | 0 | NE | NE |
| SWI/SNF BRG1/BAF Chromatin remodeling | 1.1 | 0 | NE | NE |
| SWI/SNF BRG1/PBAF Chromatin remodeling | 1.1 | 0 | NE | NE |
| NURD Chromatin remodeling | 1.1 | 0 | NE | NE |
| NURF Chromatin remodeling | 1.1 | 0 | NE | NE |
| CHRAC Chromatin remodeling | 1.1 | 0 | NE | NE |
| SRCAP Chromatin remodeling | 1.1 | 0 | NE | NE |
| Presentation of endogenous peptide antigen | 1.1 | 0 | NE | NE |
| mRNA Transcription and processing | 1.1 | 0 | NE | NE |
| rRNA Transcription and processing | 1.1 | 0 | NE | NE |
| Histones sumoylation | 1.1 | 0 | NE | NE |
| Single-strand nucleotide excision DNA repair | 1.1 | 7.5E-03 | NE | NE |
| SWI/SNF BRM/BAF Chromatin remodeling | 1.1 | 2.0E-02 | NE | NE |
| Tight junction assembly (Occludin) | 1.1 | 2.8E-02 | NE | NE |
| Classical complement pathway | −1.2 | 4.0E-02 | NE | NE |
| Ubiquitin-dependent protein degradation | 1.1 | 4.2E-02 | 1.10 | 4.9E-03 |
| Actin cytoskeleton assembly | NE | NE | 1.15 | 1.0E-02 |
NE; No effect.
3.5. Validation of gene expression changes by qPCR
Genes implicated in cell cycle checkpoint pathways were investigated due to their presence in many of the sub-networks. There is good concordance between microarray and qRT-PCR (Table 8 ). Only changes in the mRNA for card 9 and TPO were significantly altered (p < 0.05) in the qPCR assay when compared to controls but the fold differences observed for microarray and qPCR for these genes were similar. qPCR is an exponential expansion of the data and small pipetting errors sometimes lead to non-significant differences being observed. We probably needed a higher “n” for statistical significance.
| (A) Morning group | Female | |
|---|---|---|
| Gene Symbols | Microarray | qPCR |
| Aurkb | 2.83a | 2.52 |
| Duox2 | −1.73a | −1.30 |
| Incenp | 2.33a | 1.85 |
| Pomc | −1.50a | −1.05 |
| Top2a | 3.27a | 3.44 |
| (B) Night group | Male | Female | ||
|---|---|---|---|---|
| Gene Symbols | Microarrays | qPCR | Microarrays | qPCR |
| Card9 | 1.62a | 1.64** | 1.42 | 1.20 |
| Ddit 4 | 2.63a | 1.42 | 1.21 | 1.02 |
| Drd2 | 5.17a | 5.18 | 2.42 | 3.58 |
| Tpo | 1.59a | 1.90** | 1.18 | 1.18 |
| Tnfs13 | 1.63a | 1.60 | 1.34 | 1.1 |
Data are presented as mean fold changes in relative mRNA expression from control (n = 5 per treatment).
a. significantly altered in microarrays.
- . significantly altered in qPCR (p < 0.05).
4. Discussion
AA is known to produce thyroid gland follicular cell tumors and mammary gland fibroadenomas in Wistar rats. AA has been classified as a genotoxic chemical because it can alkylate DNA and cause chromosome breaks and rearrangements (clastogenic mode of action), but only at high concentrations. In the current experiment, glycidamide adducts were measured from AA [23] . Cyp2E1 is moderately expressed in the thyroid [4] . But, the active concentrations of glycidamide would have been quite low since we used a relatively low concentration of AA in a chronic exposure scenario. We tested the hypothesis that AA would alter gene expression and physiological endpoints in Wistar rats at night when they are active more so than in the daytime, when rats are sleeping. AA and glycidamide have very short biological half-lives so we thought the evening measurement would be a more sensitive indicator of AA immediate effects in these neonatal rats because AA and glycidamide would still be present in the circulation. While the overall gene expression changes observed clearly involve DNA damage and repair, there were also changes involving oxidative stress, cellular motility, motor proteins such as the kinesin-related proteins, and kinases, all of which may be causally related to genotoxicity and oncogenicity.
4.1. Phenotypic anchoring
A separate cohort of the same animals was allowed to continue on the same regimen for 2 years [51] . At the end of the two-year period, histopathology indicated tumors of the mammary gland (fibroadenomas) in females and thyroid follicular cell tumors in both sexes. No other increase of tumors was found. Hemoglobin adducts of acrylamide and glycidamide were measured over several timepoints throughout the 2 years and these showed dose and time response [23] , confirming the exposures. We did not check for DNA adducts, but in other studies, these types of exposures have been shown to induce DNA adducts [74] . In addition, other non-neoplastic changes were observed including sciatic nerve nephropathy, spinal cord degeneration and hind limb myopathy [51] . Previous studies with rats have shown similar thyroid follicular cell neoplasms [2] and [26] .
The only phenotypic endpoints we measured in the pnd 35 animals were plasma hormones. Thyroid hormones were altered only in animals sacrificed at night with both males and females showing increased plasma T3 and T4, and only females showing depressed TSH. Normally one would expect that disruption of the HPT axis in rodents in a manner that induces thyroid carcinogenesis would be through increased TSH secretion that provides growth stimulus to the thyroid gland [8] . Our results suggest that the changes in T3 and T4 were primary as opposed to secondary due to an elevation in TSH. The decreased TSH in females might reflect negative feedback suppression due to the elevated thyroid hormones.
In a study with male adult Fischer 344 rats exposed to three concentrations of AA (2.5 mg/kg/d; 10 mg/kg/d and 50 mg/kg/d) in their drinking water for 14 days and sacrificed presumably during the day, only the highest dose of AA showed a significant decrease in T4 while T3 levels showed a non-significant increasing trend [5] . These authors concluded that AA did not affect hormone production in the animals. But, critical differences exist with our study. Animals collected in the day in our study also did not show any specific effects in the plasma, but animals collected at night, did, as discussed above.
Several genes associated with thyroid function were altered in rats in the current study that point to dysregulation of hormone production. The altered genes included neuropeptide Y (NPY), solute carrier family 5 (sodium iodide symporter), member 5 (SLC5A5), iodotyrosine deiodinase (IYD) and thyroid stimulating hormone receptor (TSHR) (Table SI-1). Sub-network analysis suggested that thyroid hormone generation and thyroid function were altered in males (Fig. 2 and Table SI-2).
4.2. Commonalities and differences in gene expression changes in males and females sampled at night and in the morning
Consistent with the known effects of AA, genes associated with spindle and kinetochore assemblies were altered (Fig. 3 ). Both male and female night animals showed alteration of many elements in these pathways, including three kinesins, Kif20b, Kif22, and Kif2a. Kinesins are microtubule-associated motor proteins, which are essential components for intracellular transport and cell division [57] . There are 45 members of this superclass of proteins in mammals and their main attribute is having a microtubule binding domain and an ATPase domain. These proteins can walk along microtubules to deliver cargo to the appropriate places. Kinesins have been associated with cancer. For example, over expression of Kif2a has been associated with poor outcome for colorectal cancer [21] .
Kinesins have been demonstrated to be direct targets for AA and glycidamide [27] , [53] and [66] . While different kinesins were studied than the three upregulated in the current study, the fact that the activity of KIFC5A and KRP2, representative of two different kinesin families, could be inhibited by relatively low concentrations of AA or glycidamide (range 100–5 mM for 60–80% inhibition), suggests that the entire family may be vulnerable [66] , leading to cell division defects and potential carcinogenicity.
Interestingly, pathways involved with oxidative stress were also prominent for night animals (Fig. 4 ). While ROS was not directly measured in this study, steady state levels of mRNAs encoding proteins involved in this process were up-regulated. The overall pathway for ROS was up-regulated in both males and females sampled at night but down-regulated in females sampled in the morning. ROS generation by acrylamide has been observed in other studies [32] and [46] .
A similar pattern is seen for microtubule cytoskeleton assembly showing up-regulation of this process in night animals but an overall down-regulation in females sampled in the morning. Interestingly, night animals showed down-regulation of genes involved in muscle development suggesting that energy was diverted toward repair and detoxification and away from general anabolic metabolism (data not shown).
DNA repair, DNA replication, cell proliferation, checkpoint pathways, and apoptosis were altered by the low chronic dose of AA employed in the current study. Fig. 5 illustrates some of the entities that were common for DNA replication and DNA repair in males and females sampled at night (Fig. 5 ). As prognostic markers of cancer, transcripts involved in key checkpoint pathways were altered, including inner centromere protein (Incenp), topoisomerase (DNA) II alpha (Top2A) and budding uninhibited by benzimidazoles 1 homolog, beta (Saccharomyces cerevisiae ) (Bub1B), among others ( Table 2 ). Transcripts involved in neurotoxicity also were affected, for example dopamine receptor D2 (Drd2), which was increased by 5.2 (p = 0.018) fold in males and by 2.4 (p = 0.05) fold in females collected at night. The role of dopamine in peripheral tissues is an area of current interest [64] . Our data is consistent with the observation that exposure to acrylamide inhibited the uptake of dopamine into striatal synaptic vesicles of rats [1] and [47] .
AA is a type-2 alkene, which is a soft electrophile that reacts preferentially with nucleophilic cysteine thiolate sites of proteins, as can be found in catalytic triads [48] . However, a low but consistent reaction with the N-terminal Val group of hemoglobin has also been observed for AA and this has been used as an effective bioassay for acrylamide exposure [23] , [24] and [58] . In vivo , AA is metabolized by CYP 2E1 to glycidamide (an epoxide), which is very reactive toward nucleophiles and has been implicated in adduct formation for hemoglobin and nucleic acids [23] and [53] .
Glycidamide is also a potential source of oxidative stress. AA is converted to glycidamide, with rates varying from organ to organ [37] . AA can also bind to glutathione [25] , [29] , [49] and [70] , which normally is at high intracellular concentrations between 0.1 mM and 15 mM [13] . While most cellular proteins would be protected by the high GSH, the intracellular concentrations can vary depending on the location, with the lowest intracellular ratio of GSH-GSSG measured in the rough endoplasmic reticulum [13] . Modulation of GSH levels by AA has been demonstrated in Syrian Hamster Embryo (SHE) cells and is an essential occurrence for cell transformation [39] .
Because AA effectively alkylates Cys residues, a low ratio of GSH:GSSG would promote alkylation of proteins, which would be influenced by active drinking of AA laced water. Both cysteine and glutathione were protective of AA induced malformation in frogs [63] . Reactivity toward Cys suggests vulnerabilities. If proteins with active site Cys residues were adducted, this could result in enzyme inactivation. The system would likely respond by making more mRNA for inactivated proteins. Interestingly, in our study, we identified increased levels of mRNAs for proteins that are known to be cysteine rich and that may contain Cys residues in their active sites, including cysteine-rich with EGF-like domains 2 (Creld2), glutamate-cysteine ligase (Gclc) catalytic subunit, and selenocysteine lyase (Scly), which were increased by 2.2 fold and 2.1 fold; by 2.0 and 1.4; and by 1.7 and 1.3 in male and female rats at night, respectively (Table 2 ). Creld2 has been proposed as a novel endoplasmic reticulum stress-inducible gene, associated with folding, processing and protein transport [60] . Scly is a selenoprotein involved in cancer with its main function being protection against an excess of ROS [6] and [68] . Gclc controls synthesis of GSH, the main antioxidant in the cell [9] , [14] and [42] .
It is useful to consider a model that includes adaptive/compensatory gene expression changes as protective action against damage by low concentrations of AA. With chronic exposure, the compensatory pathways may not fully contain cellular damage, unleashing a second line of defense, for example up-regulation of apoptotic processes. At this stage of cellular assault, one would expect to see additional pathways altered including those involving structural proteins within the cell, since caspases activated via apoptosis can cut spectrin and other cellular scaffold proteins [75] . Only at higher levels of cellular damage, caused by the conversion of AA to glycidamide, would covalent adducts be seen with proteins and/or DNA, unleashing DNA repair and disturbing cell cycle regulation and check point pathways [10] .
Dose-response relationships between the formation of tumors in thyroid tissues and genotoxic events suggest that other mechanisms in addition to genotoxicity might be involved in tumor formation in the thyroid at low concentrations of AA [15] , for example through stimulation of thyroid cell proliferation by an alternate mechanism. The evidence that hormones may be involved actually comes from studies using relatively high doses of AA, 10–50 mg/Kg bw/d. At these high concentrations of AA, serum LH levels were significantly increased, while FSH, testosterone and prolactin, were significantly decreased in chronically dosed male rats [7] . At lower doses of AA (2–15 mg/Kg bw/d range; 2–7 day exposure), a slight increase in T4 and a decrease of TSH levels were previously reported [36] , which are in agreement with hormonal changes seen in our study.
Rats are nocturnal animals and sleep during the day from 6 AM to 6 PM [30] and [71] . The sampling regimen was designed to sample rats at night when they were actively drinking water laced with AA and compare them to quiescent animals sampled in the morning. In each case, the sampling occurred several hours into each phase, at 10 PM or 10 AM. While all animals showed alterations in genes related to DNA damage, there were additional altered pathways that appeared mainly in the night groups, suggesting that these were stimulated by newly introduced AA. These pathways were related with reactive oxygen species, oxidative stress, detoxification systems, energy production and metabolism.
The commonality among all of the AA dosed animals suggests DNA damage, cell damage and cell death as the primary, long-term effects caused by chronic AA exposure. These findings are consistent with other studies that point to DNA damage and tumorigenesis [11] , [17] , [26] , [28] , [38] , [41] , [50] , [54] , [61] and [62] .
These data bear directly on the regulatory classification of AA as a genotoxic carcinogen. In regulatory schemes AA risk is quantitated as that of a genotoxic carcinogen [18] and [20] . However, at low and intermittent doses, tumorigenesis appears to be consecutive to AA targeting key cellular proteins and their functions leading to oxidative stress and to thyroid hormone changes, rather than genotoxicity.
Conflict of interest
The authors declare that there are no conflict of interest.
Transparency document
Transparency Document.
Acknowledgements
We thank Kenneth L Phillips, ILS Genomics, Morrisville, NC, for a second bioinformatics analysis of the data. This research was funded by a grant from SNF SAS, ZAC de Milieux, Andrézieux, France. Marvin Friedman is a paid consultant/advisor to SNF SAS and he helped design the study and helped with interpretation of results. LPT is a contractor for SNF-SAS.
Appendix A. Supplementary data
The following are Supplementary data to this article:
References
- [1] D.S. Barber, S. Stevens, R.M. LoPachin; Proteomic analysis of rat striatal synaptosomes during acrylamide intoxication at a low Dose rate; Toxicol. Sci., 100 (1) (2007), pp. 156–167 http://dx.doi.org/10.1093/toxsci/kfm210
- [2] F.A. Beland, P.W. Mellick, G.R. Olson, M.C. Mendoza, M.M. Marques, D.R. Doerge; Carcinogenicity of acrylamide in B6C3F(1) mice and F344/N rats from a 2-year drinking water exposure; Food Chem. Toxicol., 51 (2013), pp. 149–159 http://dx.doi.org/10.1016/j.fct.2012.09.017
- [3] N. Ben-Jonathan, L.A. Arbogast, J.F. Hyde; Neuroendocrine [corrected] regulation of prolactin release; Prog. Neurobiol., 33 (5–6) (1989), pp. 399–447
- [4] I. Bieche, C. Narjoz, T. Asselah, S. Vacher, P. Marcellin, R. Lidereau, P. Beaune, I. de Waziers; Reverse transcriptase-PCR quantification of mRNA levels from cytochrome (CYP) 1, CYP2 and CYP3 families in 22 different human tissues; Pharmacogenet. Genomics, 17 (9) (2007), pp. 731–742 http://dx.doi.org/10.1097/FPC.0b013e32810f2e58
- [5] J.F. Bowyer, J.R. Latendresse, R.R. Delongchamp, L. Muskhelishvili, A.R. Warbritton, M. Thomas, E. Tareke, L.P. McDaniel, D.R. Doerge; The effects of subchronic acrylamide exposure on gene expression, neurochemistry, hormones, and histopathology in the hypothalamus-pituitary-thyroid axis of male Fischer 344 rats; Toxicol. Appl. Pharmacol., 230 (2) (2008), pp. 208–215 http://dx.doi.org/10.1016/j.taap.2008.02.028
- [6] P. Brenneisen, H. Steinbrenner, H. Sies; Selenium, oxidative stress, and health aspects; Mol. Aspects Med., 26 (4–5) (2005), pp. 256–267 http://dx.doi.org/10.1016/j.mam.2005.07.004
- [7] L. Camacho, J.R. Latendresse, L. Muskhelishvili, R. Patton, J.F. Bowyer, M. Thomas, D.R. Doerge; Effects of acrylamide exposure on serum hormones, gene expression, cell proliferation, and histopathology in male reproductive tissues of Fischer 344 rats; Toxicol. Lett., 211 (2) (2012), pp. 135–143 http://dx.doi.org/10.1016/j.toxlet.2012.03.007
- [8] C.C. Capen, S.L. Martin; The effects of xenobiotics on the structure and function of thyroid follicular and C-cells; Toxicol. Pathol., 17 (2) (1989), pp. 266–293
- [9] C.N. Chen, H.M. Brown-Borg, S.G. Rakoczy, D.A. Ferrington, L.V. Thompson; Aging impairs the expression of the catalytic subunit of glutamate cysteine ligase in soleus muscle under stress; J Gerontol. A Biol. Sci. Med. Sci., 65 (2) (2010), pp. 129–137 http://dx.doi.org/10.1093/gerona/glp194
- [10] J.-H. Chen, T.-C. Tsou, I.-M. Chiu, C.-C. Chou; Proliferation inhibition, DNA damage, and cell-Cycle arrest of human astrocytoma cells after acrylamide exposure; Chem. Res. Toxicol., 23 (9) (2010), pp. 1449–1458 http://dx.doi.org/10.1021/tx1000893
- [11] F.C. Clement, R. Dip, H. Naegeli; Expression profile of human cells in culture exposed to glycidamide, a reactive metabolite of the heat-induced food carcinogen acrylamide; Toxicology, 240 (1–2) (2007), pp. 111–124 http://dx.doi.org/10.1016/j.tox.2007.07.019
- [12] Committee-on-Mutagenicity-of-Chemicals-in-Food (2009). Statement on the Genotoxicity of Acrylamide In (doi, London, England).
- [13] M. Deponte; Glutathione catalysis and the reaction mechanisms of glutathione-dependent enzymes; Biochim. Biophys. Acta, 1830 (5) (2013), pp. 3217–3266 http://dx.doi.org/10.1016/j.bbagen.2012.09.018
- [14] D.A. Dickinson, A.L. Levonen, D.R. Moellering, E.K. Arnold, H. Zhang, V.M. Darley-Usmar, H.J. Forman; Human glutamate cysteine ligase gene regulation through the electrophile response element; Free Radic. Biol. Med., 37 (8) (2004), pp. 1152–1159 http://dx.doi.org/10.1016/j.freeradbiomed.2004.06.011
- [15] M. Dourson, R. Hertzberg, B. Allen, L. Haber, A. Parker, O. Kroner, A. Maier, M. Kohrman; Evidence-based Dose-response assessment for thyroid tumorigenesis from acrylamide; Regul. Toxicol. Pharmacol., 52 (3) (2008), pp. 264–289 http://dx.doi.org/10.1016/j.yrtph.2008.08.004
- [16] EFSA (2015). Acrylamide in food is a public health concern. In http://www.efsa.europa.eu/en/press/news/150604
- [17] A. Ehlers, D. Lenze, H. Broll, J. Zagon, M. Hummel, A. Lampen; Dose dependent molecular effects of acrylamide and glycidamide in human cancer cell lines and human primary hepatocytes; Toxicol. Lett., 217 (2) (2013), pp. 111–120 http://dx.doi.org/10.1016/j.toxlet.2012.12.017
- [18] EPA (2008). Toxicological Review of Acrylamide. In (E. S. A. Board, Eds.) doi, Washington, DC.
- [19] L.S. Erdreich, M.A. Friedman; Epidemiologic evidence for assessing the carcinogenicity of acrylamide; Regul. Toxicol. Pharmacol., 39 (2) (2004), pp. 150–157 http://dx.doi.org/10.1016/j.yrtph.2003.12.004
- [20] EU-Existing-Chemical-Branch (2000). Risk Assessment of Acrylamide. In (E.C. Branch, Eds.) doi, Milan, IT.
- [21] X. Fan, X. Wang, H. Zhu, W. Wang, S. Zhang, Z. Wang; KIF2A overexpression and its association with clinicopathologic characteristics and unfavorable prognosis in colorectal cancer; Tumour Biol., 36 (11) (2015), pp. 8895–8902 http://dx.doi.org/10.1007/s13277-015-3603-z
- [22] FAO/WHO; Expert Committee on Food Additives; (2010) http://www.who.int/foodsafety/chem/summary72_rev.pdf
- [23] T.R. Fennell, R. Snyder, B. Hansen, M. Friedman; Dosimetry of acrylamide and glycidamide over the lifespan in a 2-Year bioassay of acrylamide in wistar han rats; Toxicol. Sci., 146 (2) (2015), pp. 386–394 http://dx.doi.org/10.1093/toxsci/kfv104
- [24] M. Friedman; Chemistry, biochemistry, and safety of acrylamide. A review; J. Agric. Food Chem., 51 (16) (2003), pp. 4504–4526 http://dx.doi.org/10.1021/jf030204+
- [25] M. Friedman; Biological effects of Maillard browning products that may affect acrylamide safety in food: biological effects of Maillard products; Adv. Exp. Med. Biol., 561 (2005), pp. 135–156 http://dx.doi.org/10.1007/0-387-24980-X_12
- [26] M.A. Friedman, L.H. Dulak, M.A. Stedham; A lifetime oncogenicity study in rats with acrylamide; Fundam. Appl. Toxicol., 27 (1) (1995), pp. 95–105
- [27] M.A. Friedman, E. Zeiger, D.E. Marroni, D.W. Sickles; Inhibition of rat testicular nuclear kinesins (krp2; KIFC5A) by acrylamide as a basis for establishing a genotoxicity threshold; J. Agric. Food Chem., 56 (15) (2008), pp. 6024–6030 http://dx.doi.org/10.1021/jf703746f
- [28] B.I. Ghanayem, L.P. McDaniel, M.I. Churchwell, N.C. Twaddle, R. Snyder, T.R. Fennell, D.R. Doerge; Role of CYP2E1 in the epoxidation of acrylamide to glycidamide and formation of DNA and hemoglobin adducts; Toxicol. Sci., 88 (2) (2005), pp. 311–318 http://dx.doi.org/10.1093/toxsci/kfi307
- [29] K. Hashimoto, W.N. Aldridge; Biochemical studies on acrylamide, a neurotoxic agent; Biochem. Pharmacol., 19 (9) (1970), pp. 2591–2604
- [30] G.M. Herrera, A.L. Meredith; Diurnal variation in urodynamics of rat; PloS one, 8 (2010), p. e12298 http://dx.doi.org/10.1371/journal.pone.0012298
- [31] R.N. Hill, T.M. Crisp, P.M. Hurley, S.L. Rosenthal, D.V. Singh; Risk assessment of thyroid follicular cell tumors; Environ. Health Perspect., 106 (8) (1998), pp. 447–457
- [32] L. Jiang, J. Cao, Y. An, C. Geng, S. Qu, L. Jiang, L. Zhong; Genotoxicity of acrylamide in human hepatoma G2 (HepG2) cells; Toxicol. In Vitro, 21 (8) (2007), pp. 1486–1492 http://dx.doi.org/10.1016/j.tiv.2007.06.011
- [33] K.A. Johnson, S.J. Gorzinski, K.M. Bodner, R.A. Campbell, C.H. Wolf, M.A. Friedman, R.W. Mast; Chronic toxicity and oncogenicity study on acrylamide incorporated in the drinking water of Fischer 344 rats; Toxicol. Appl. Pharmacol., 85 (2) (1986), pp. 154–168
- [34] L.V. Johnson, J.C. Blanks; Application of acrylamide as an embedding medium in studies of lectin and antibody binding in the vertebrate retina; Curr. Eye Res., 3 (7) (1984), pp. 969–974
- [35] A. Kauffmann, R. Gentleman, W. Huber; arrayQualityMetrics—a bioconductor package for quality assessment of microarray data; Bioinformatics, 25 (3) (2009), pp. 415–416 http://dx.doi.org/10.1093/bioinformatics/btn647
- [36] M.A. Khan, C.A. Davis, G.L. Foley, M.A. Friedman, L.G. Hansen; Changes in thyroid gland morphology after acute acrylamide exposure; Toxicol. Sci., 47 (2) (1999), pp. 151–157
- [37] T.H. Kim, S. Shin, K.B. Kim, W.S. Seo, J.C. Shin, J.H. Choi, K.Y. Weon, S.H. Joo, S.W. Jeong, B.S. Shin; Determination of acrylamide and glycidamide in various biological matrices by liquid chromatography-tandem mass spectrometry and its application to a pharmacokinetic study; Talanta, 131 (2015), pp. 46–54 http://dx.doi.org/10.1016/j.talanta.2014.07.042
- [38] J.E. Klaunig; Acrylamide carcinogenicity; J. Agric. Food Chem., 56 (15) (2008), pp. 5984–5988 http://dx.doi.org/10.1021/jf8004492
- [39] J.E. Klaunig, L.M. Kamendulis; Mechanisms of acrylamide induced rodent carcinogenesis; Adv. Exp. Med. Biol., 561 (2005), pp. 49–62 http://dx.doi.org/10.1007/0-387-24980-X_4
- [40] E. Kotelnikova, M.A. Shkrob, M.A. Pyatnitskiy, A. Ferlini, N. Daraselia; Novel approach to meta-analysis of microarray datasets reveals muscle remodeling-related drug targets and biomarkers in Duchenne muscular dystrophy; PLoS Comput. Biol., 2 (2012), p. e1002365 http://dx.doi.org/10.1371/journal.pcbi.1002365
- [41] N. Koyama, H. Sakamoto, M. Sakuraba, T. Koizumi, Y. Takashima, M. Hayashi, H. Matsufuji, K. Yamagata, S. Masuda, N. Kinae, M. Honma; Genotoxicity of acrylamide and glycidamide in human lymphoblastoid TK6 cells; Mutat. Res., 603 (2) (2006), pp. 151–158 http://dx.doi.org/10.1016/j.mrgentox.2005.11.006
- [42] D.M. Krzywanski, D.A. Dickinson, K.E. Iles, A.F. Wigley, C.C. Franklin, R.M. Liu, T.J. Kavanagh, H.J. Forman; Variable regulation of glutamate cysteine ligase subunit proteins affects glutathione biosynthesis in response to oxidative stress; Arch. Biochem. Biophys., 423 (1) (2004), pp. 116–125 http://dx.doi.org/10.1016/j.abb.2003.11.004
- [43] J.S. Lafferty, L.M. Kamendulis, J. Kaster, J. Jiang, J.E. Klaunig; Subchronic acrylamide treatment induces a tissue-specific increase in DNA synthesis in the rat; Toxicol. Lett., 154 (1–2) (2004), pp. 95–103 http://dx.doi.org/10.1016/j.toxlet.2004.07.008
- [44] L. Lipworth, J.S. Sonderman, R.E. Tarone, J.K. McLaughlin; Review of epidemiologic studies of dietary acrylamide intake and the risk of cancer; Eur. J. Cancer Prev., 21 (4) (2012), pp. 375–386 http://dx.doi.org/10.1097/CEJ.0b013e3283529b64
- [45] K.J. Livak, T.D. Schmittgen; Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method; Methods, 25 (4) (2001), pp. 402–408 http://dx.doi.org/10.1006/meth.2001.1262
- [46] R.M. LoPachin, T. Gavin; Molecular mechanism of acrylamide neurotoxicity: lessons learned from organic chemistry; Environ. Health Perspect., 120 (12) (2012), pp. 1650–1657 http://dx.doi.org/10.1289/ehp.1205432
- [47] R.M. LoPachin, D.S. Barber, D. He, S. Das; Acrylamide inhibits dopamine uptake in rat striatal synaptic vesicles; Toxicol. Sci., 89 (1) (2006), pp. 224–234 http://dx.doi.org/10.1093/toxsci/kfj005
- [48] R.M. Lopachin, T. Gavin, A. Decaprio, D.S. Barber; Application of the hard and soft, acids and bases (HSAB) theory to toxicant–target interactions; Chem. Res. Toxicol., 25 (2) (2012), pp. 239–351 http://dx.doi.org/10.1021/tx2003257
- [49] Y.S. Luo, T.Y. Long, L.C. Shen, S.L. Huang, S.Y. Chiang, K.Y. Wu; Synthesis, characterization and analysis of the acrylamide- and glycidamide-glutathione conjugates; Chem. Biol. Interact., 237 (2015), pp. 38–46 http://dx.doi.org/10.1016/j.cbi.2015.05.002
- [50] I. Maniere, T. Godard, D.R. Doerge, M.I. Churchwell, M. Guffroy, M. Laurentie, J.M. Poul; DNA damage and DNA adduct formation in rat tissues following oral administration of acrylamide; Mutat. Res., 580 (1–2) (2005), pp. 119–129 http://dx.doi.org/10.1016/j.mrgentox.2004.10.012
- [51] R.R. Maronpot, R.J.M.M. Thoolen, B. Hansen; Two-year carcinogenicity study of acrylamide in wistar han rats with In utero exposure; Exp. Toxicol. Pathol., 67 (2) (2015), pp. 189–195
- [52] R.R. Maronpot, E. Zeiger, E.E. McConnell, H. Kolenda-Roberts, H. Wall, M.A. Friedman; Induction of tunica vaginalis mesotheliomas in rats by xenobiotics; Crit. Rev. Toxicol., 39 (6) (2009), pp. 512–537 http://dx.doi.org/10.1080/10408440902969430
- [53] C.J. Martyniuk, A. Feswick, B. Fang, J.M. Koomen, D.S. Barber, T. Gavin, R.M. Lopachin; Protein targets of acrylamide adduct formation in cultured rat dopaminergic cells; Toxicol. Lett., 219 (3) (2013), pp. 279–287 http://dx.doi.org/10.1016/j.toxlet.2013.03.031
- [54] N. Mei, J. Hu, M.I. Churchwell, L. Guo, M.M. Moore, D.R. Doerge, T. Chen; Genotoxic effects of acrylamide and glycidamide in mouse lymphoma cells; Food Chem. Toxicol., 46 (2) (2008), pp. 628–636 http://dx.doi.org/10.1016/j.fct.2007.09.093
- [55] D.S. Mottram, B.L. Wedzicha, A.T. Dodson; Acrylamide is formed in the Maillard reaction; Nature, 419 (6906) (2002), pp. 448–449 http://dx.doi.org/10.1038/419448a
- [56] F. Neumann; Early indicators for carcinogenesis in sex-hormone-sensitive organs; Mutat. Res., 248 (2) (1991), pp. 341–356
- [57] S. Niwa; Kinesin superfamily proteins and the regulation of microtubule dynamics in morphogenesis; Anat. Sci. Int., 90 (1) (2015), pp. 1–6 http://dx.doi.org/10.1007/s12565-014-0259-5
- [58] NTP 2005 NTP-CERHR Monograph on the Potential Human Reproductive and Developmental Effects of Acrylamide.
- [59] A.E. Obr, D.P. Edwards; The biology of progesterone receptor in the normal mammary gland and in breast cancer; Mol. Cell. Endocrinol., 357 (1–2) (2012), pp. 4–17 http://dx.doi.org/10.1016/j.mce.2011.10.030
- [60] K. Oh-hashi, H. Koga, S. Ikeda, K. Shimada, Y. Hirata, K. Kiuchi; CRELD2 is a novel endoplasmic reticulum stress-inducible gene; Biochem. Biophys. Res. Commun., 387 (3) (2009), pp. 504–510 http://dx.doi.org/10.1016/j.bbrc.2009.07.047
- [61] M. Pingarilho, N.G. Oliveira, C. Martins, B.C. Gomes, A.S. Fernandes, V. Martins, A. Labilloy, J.P. de Lima, J. Rueff, J.F. Gaspar; Induction of sister chromatid exchange by acrylamide and glycidamide in human lymphocytes: role of polymorphisms in detoxification and DNA-repair genes in the genotoxicity of glycidamide; Mutat. Res., 752 (1–2) (2013), pp. 1–7 http://dx.doi.org/10.1016/j.mrgentox.2012.12.013
- [62] N. Puppel, Z. Tjaden, F. Fueller, D. Marko; DNA strand breaking capacity of acrylamide and glycidamide in mammalian cells; Mutat. Res., 580 (1–2) (2005), pp. 71–80 http://dx.doi.org/10.1016/j.mrgentox.2004.11.009
- [63] J.R. Rayburn, M. Friedman; -Cysteine, N -Acetyl--cysteine, and glutathione protect xenopus laevis embryos against acrylamide-Induced malformations and mortality in the frog embryo teratogenesis assay ; J. Agric. Food Chem., 58 (20) (2010), pp. 11172–11178 http://dx.doi.org/10.1021/jf1023998
- [64] B. Rubi, P. Maechler; Minireview: new roles for peripheral dopamine on metabolic control and tumor growth: let’s seek the balance; Endocrinology, 151 (12) (2010), pp. 5570–5581 2010-0745 http://dx.doi.org/10.1210/en. 2010-0745
- [65] A. Shipp, G. Lawrence, R. Gentry, T. McDonald, H. Bartow, J. Bounds, N. Macdonald, H. Clewell, B. Allen, C. Van Landingham; Acrylamide: review of toxicity data and dose-response analyses for cancer and noncancer effects; Crit. Rev. Toxicol., 36 (6–7) (2006), pp. 481–608 http://dx.doi.org/10.1080/10408440600851377
- [66] D.W. Sickles, A.O. Sperry, A. Testino, M. Friedman; Acrylamide effects on kinesin-related proteins of the mitotic/meiotic spindle; Toxicol. Appl. Pharmacol., 222 (1) (2007), pp. 111–121 http://dx.doi.org/10.1016/j.taap.2007.04.006
- [67] G.K. Smyth; Limma: linear models for microarray data; R. Gentleman, V.J. Carey, W. Huber, R.A. Irizarry, S. Dudoit (Eds.), Bioinformatics and Computational Biology Solutions Using R and Bioconductor, Statistics for Biology, Health, Springer, New York (2005), pp. 397–420 (doi)
- [68] H. Steinbrenner, H. Sies; Protection against reactive oxygen species by selenoproteins; Biochim. Biophys. Acta, 1790 (11) (2009), pp. 1478–1485 http://dx.doi.org/10.1016/j.bbagen.2009.02.014
- [69] A. Subramanian, P. Tamayo, V.K. Mootha, S. Mukherjee, B.L. Ebert, M.A. Gillette, A. Paulovich, S.L. Pomeroy, T.R. Golub, E.S. Lander, J.P. Mesirov; Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles; Proc. Natl. Acad. Sci. U.S.A., 102 (43) (2005), pp. 15545–15550 http://dx.doi.org/10.1073/pnas.0506580102
- [70] G.C. Tong, W.K. Cornwell, G.E. Means; Reactions of acrylamide with glutathione and serum albumin; Toxicol. Lett., 147 (2) (2004), pp. 127–131
- [71] M. Verwey, B. Robinson, S. Amir; Recording and analysis of circadian rhythms in running-wheel activity in rodents; J. Vis. Exp., 71 (2013) http://dx.doi.org/10.3791/50186
- [72] F.H. Yerlikaya, A. Toker, Y. Yener; Effects of acrylamide treatment on oxidant and antioxidant levels in rats; Kafkas Univ. Vet. Falk. Derg., 19 (2013), pp. 607–612
- [73] M.I. Yousef, F.M. El-Demerdash; Acrylamide-induced oxidative stress and biochemical perturbations in rats; Toxicology, 219 (1–3) (2006), pp. 133–141 http://dx.doi.org/10.1016/j.tox.2005.11.008
- [74] E. Zeiger, L. Recio, T.R. Fennell, J.K. Haseman, R.W. Snyder, M. Friedman; Investigation of the low-dose response in the in vivo induction of micronuclei and adducts by acrylamide; Toxicol. Sci., 107 (1) (2009), pp. 247–257 http://dx.doi.org/10.1093/toxsci/kfn214
- [75] X. Zhang, F. Chen, Z. Huang; Apoptosis induced by acrylamide is suppressed in a 21.5% fat diet through caspase-3-independent pathway in mice testis; Toxicol. Mech. Methods, 19 (3) (2009), pp. 219–224 http://dx.doi.org/10.1080/15376510802499048
Document information
Published on 02/05/17
Accepted on 02/05/17
Submitted on 02/05/17
Licence: Other
Share this document
Keywords
claim authorship
Are you one of the authors of this document?