Highlights
- Cytoplasmic DNA, comprising LINE-1-derived sequences, elicits IFN expression via the cGAS-STING pathway in SLX4-deficiency.
- Members of the Fanconi Anemia DNA repair pathway negatively regulate LINE-1 retrotransposition.
- Endogenous reverse transcriptase activities contribute to spontaneous and chemotherapy-induced inflammation.
Chronic inflammation favors tumorigenesis, negatively influencing patient prognosis. Yet, the underlying molecular mechanisms are poorly understood. Here, we show that increased endogenous retroelement-associated reverse transcriptase activity contributes to generate immunogenic cytoplasmic nucleic acids susceptible of triggering a pro-inflammatory response in the Fanconi Anemia (FA) cancer susceptibility syndrome. In addition, treatment of FA cells or of cells exposed to replication stress inducing drugs, with a reverse transcriptase inhibitor, decreases pro-inflammatory signals. Altogether our data suggest the involvement of endogenous reverse transcriptase activities in sustaining pervasive chronic inflammation, opening therapeutic perspectives for preventing its impact on tumorigenesis.
Abstract
Fanconi Anemia (FA) is a genetic disorder characterized by elevated cancer susceptibility and pro-inflammatory cytokine production. Using SLX4FANCP deficiency as a working model, we questioned the trigger for chronic inflammation in FA. We found that absence of SLX4 caused cytoplasmic DNA accumulation, including sequences deriving from active Long INterspersed Element-1 (LINE-1), triggering the cGAS-STING pathway to elicit interferon (IFN) expression. In agreement, absence of SLX4 leads to upregulated LINE-1 retrotransposition. Importantly, similar results were obtained with the FANCD2 upstream activator of SLX4. Furthermore, treatment of FA cells with the Tenofovir reverse transcriptase inhibitor (RTi), that prevents endogenous retrotransposition, decreased both accumulation of cytoplasmic DNA and pro-inflammatory signaling. Collectively, our data suggest a contribution of endogenous RT activities to the generation of immunogenic cytoplasmic nucleic acids responsible for inflammation in FA. The additional observation that RTi decreased pro-inflammatory cytokine production induced by DNA replication stress-inducing drugs further demonstrates the contribution of endogenous RTs to sustaining chronic inflammation. Altogether, our data open perspectives in the prevention of adverse effects of chronic inflammation in tumorigenesis.
Keywords
Inflammation ; DNA damage ; SLX4 complex ; endogenous retroelement ; Interferon ; Fanconi Anemia ; cGAS-STING ; Innate immune sensing ; Cytoplasmic DNA
1. Introduction
Fanconi Anemia (FA) is a rare autosomal recessive genetic disorder caused by bi-allelic mutations in any of the 17 identified FA complementation group genes that compose the FA DNA repair pathway (Wang and Smogorzewska, 2015 ). FA is characterized by elevated pro-inflammatory cytokine levels, heightened cancer susceptibility, severe developmental defects and progressive bone marrow failure (Bogliolo and Surralles, 2015 ). Recent work has shown that the SLX4FANCP effector of the FA DNA repair pathway, is directly involved in regulating pathogen-induced cytokine production (Laguette et al., 2014 ). Indeed, SLX4 assembles several proteins involved in DNA metabolism, into a complex (SLX4com) involved in the repair of double strand breaks (Matos and West, 2014 ), but also likely acts on incoming pathogen-associated nucleic acids to favor their clearance (Laguette et al., 2014 ) preventing innate immune sensing. Furthermore, spontaneous production of pro-inflammatory cytokines by FA cells (Du et al., 2014 , Dufour et al., 2003 and Laguette et al., 2014 ) also raises the hypothesis that this DNA repair pathway may be required for the processing of immunogenic endogenous nucleic acids.
Accordingly, besides their role in DNA repair, a direct role of DNA damage response (DDR) proteins in the control of spontaneous and pathogen-induced pro-inflammatory cytokine production has recently emerged (Bregnard et al., 2014 and Hartlova et al., 2015 ). It has been shown that DDR proteins can either detect non-canonical endogenous and pathogen-derived nucleic acids to trigger the production of pro-inflammatory cytokines, including antiviral interferons (IFN) (Ferguson et al., 2012 , Kondo et al., 2013 and Zhang et al., 2011 ), or promote their degradation and prevent recognition by the innate immune system (Laguette et al., 2014 , Stetson et al., 2008 and Yan et al., 2010 ). In agreement, persistent genomic lesions have been linked to elevated pro-inflammatory cytokine production (Bald et al., 2014 and Brzostek-Racine et al., 2011 ).
Recent reports also show that cancer cells present with cytoplasmic nucleic acids that trigger the production of type I IFN (Hartlova et al., 2015 , Koo et al., 2015 and Shen et al., 2015 ). Recognition of these nucleic acids relies on pattern recognition receptors that converge on adaptor proteins to ensure activation of transcription factors such as NF-κB and interferon regulatory factor (IRF) 3 and IRF7 to produce IFN and additional pro-inflammatory cytokines (Paludan, 2015 ). Detection of cytoplasmic DNA leads to the activation of the adaptor protein stimulator of interferon genes (STING) upon activation of sensor proteins like Cyclic GMP-AMP synthase (cGAS) or Gamma-interferon-inducible protein ifi-16 (IFI16) (Ishikawa and Barber, 2008 and Ishikawa et al., 2009 ), while extracellular nucleic acids are detected by Toll-Like Receptors (TLRs) that reside on intracellular organelles such as endosomes, leading to the activation of myeloid differentiation primary response gene 88 (MYD88) or TIR-domain-containing adaptor-inducing interferon-β (TRIF) (Rakoff-Nahoum and Medzhitov, 2009 ). While the origin of endogenous immunogenic nucleic acids is still debated, there are indications that they are generated in the nucleus (Hartlova et al., 2015 and Yang et al., 2007 ) and are, at least in part, comprised of DNA repair by-products (Rigby et al., 2014 and Shen et al., 2015 ). Interestingly, absence of DDR proteins can lead to upregulated retrotransposition (Stetson et al., 2008 ), which constitutes a potential source of immunogenic endogenous nucleic acids. Indeed, a subfamily of Long INterspersed Element-1 (LINE-1), retains the ability to retrotranspose autonomously and re-insert at distant genomic locations (Babushok and Kazazian, 2007 and Kazazian et al., 1988 ). In addition, de-repression of LINE-1 elements has been documented in a range of inflammatory and autoimmune diseases (Volkman and Stetson, 2014 ) associated with mutations in genes involved in the metabolism of nucleic acids, including genomic lesions and incoming pathogen-derived nucleic acids (Gasior et al., 2008 and Stetson et al., 2008 ). However, how upregulated LINE-1 activity may lead to induction and persistence of pro-inflammatory signals remains to be investigated.
Here, we wished to identify the trigger for spontaneous pro-inflammatory cytokine production in FA. We used SLX4 deficiency as a working model, because we previously identified a role for SLX4-associated nuclease activities in repressing HIV-mediated pro-inflammatory signaling (Laguette et al., 2014 ). We reveal that SLX4-deficient cells present cytoplasmic DNA that is recognized by the cGAS-STING pathway to activate IFN signaling. We further establish that these cytoplasmic nucleic acids are, at least in part, generated by upregulated LINE-1 activity. Accordingly, treatment with Tenofovir, a nucleoside analogue reverse-transcriptase (RT) inhibitor that can prevent endogenous RT activities, leads to decreased cytoplasmic LINE-1-derived nucleic acids and pro-inflammatory cytokine levels. Finally, we also show that treatment with Tenofovir prevents chemotherapy-induced inflammation. Altogether, by establishing a link between upregulated endogenous RT activities and spontaneous pro-inflammatory cytokine production in SLX4 deficiency, our findings open perspectives in preventing pervasive pro-inflammatory signaling in the FA cancer susceptibility syndrome and chemotherapy.
2. Materials and Methods
For detailed experimental procedures see Supplementary material.
2.1. Cell Lines and Plasmids
RA3331- and PD20-derived cell lines were kind gifts from Dr. Smogorzewska and Dr. Rosselli respectively. The A549 IFN-responsive cell line (Chen et al., 2010 ) was obtained from Dr. Goodbourn.
pCEP-UB_LRE3_GFPi (LINE-1 GFP), pAD2TE1 (LINE-1 NEO RT +) and pAD135 (LINE-1 NEO RT-) were gifts from Dr. Gilbert (Doucet et al., 2010 and Wei et al., 2001 ). pYX017 (LINE-1 LUC RT +) and pYX015 (LINE-1 LUC ORF1mut) were kindly provided by Dr. A n (Xie et al., 2011 ).
2.2. Cytoplasmic and Nuclear DNA Extraction and Analysis
All steps were performed at 4 °C except otherwise mentioned. Cytoplasmic and nuclear fractions were extracted from equal amounts of cells. Cells were lysed in low salt lysis buffer (100 mM NaCl; 10% glycerol; 0.1% Triton; 20 mM Tris [pH 7.5]; 2 mM MgCl2 ; 0.5 mM EDTA; 0.2 mM PMSF; 10 mM β-mercaptoethanol) for 20 min on wheel, followed by 10 min centrifugation at 4000 rpm. The supernatant, corresponding to the cytoplasmic fraction, was collected while the remaining pellet was further extracted with high salt lysis buffer (340 mM NaCl; 10% glycerol; 0.1% Triton; 20 mM Tris [pH 7.5]; 2 mM MgCl2 ; 0.5 mM EDTA; 0.2 mM PMSF; 10 mM β-mercaptoethanol) for 30 min on wheel, followed by 30 min centrifugation at 13,200 rpm. The supernatant, corresponding to the soluble nuclear fraction, was collected. Cytoplasmic and nuclear fractions were assessed by WB and further treated or not with RNase cocktail (RNase A 5 U/ml, RNase T1 200 U/ml; Ambion) for 30 min at 37 °C and Proteinase K (200 μg/ml; Ambion) for 2 h at 50 °C prior to DNA extraction using Phenol/Chloroform/Isoamyl pH 8 (12/12/1).
For qPCR analysis using LINE-1-specific primers, results were normalized on Cytochrome B levels and Genomic DNA contamination. Active LINE-1 sequences were identified using primers described by Boissinot et al. (2000) . To sequence cytosolic DNA from RA3331 cells, PCR products were cloned into the TOPO Blunt Vector (ThermoFisher Scientific). Analysis of sequences was performed using a consensus LINE-1 sequence (m80343 clone; NCBI).
For radiolabeling experiments, the DNA was dephosphorylated using rSAP (NEB) prior to labeling with γ32 P ATP for 30 min at 37 °C using the T4 Polynucleotide kinase (NEB). Subsequent S1 Nuclease, dsDNase and RNaseH (ThermoFisher Scientific) treatments were performed following the manufacturers protocol. Unbound radiolabeled nucleotides were removed using Illustra Microspin G-50 Columns (GE Healthcare) prior to resolution on 5% acrylamide, 0.5% Tris Borate Ethylamide gel and autoradiography.
Southern Blot analysis using a LINE-1-specific probe was performed following resolution on 1.3% agarose gels. The probe was generated using the pYX017 plasmid and labeled using the Prime-a-Gene Labeling System (Promega) according to the manufacturers protocol.
2.3. RNA Extraction and RT-qPCR
RNA was extracted using Trizol (Invitrogen) and treated with TURBO DNase (Ambion) according to the manufacturers' protocol. Reverse transcription (SuperSriptIII reverse transcriptase; Invitrogen) and qPCR (QIAGEN) using specific primers were performed using standard protocols.
2.4. Immunoprecipitation
293T cells were transfected with indicated plasmids using the calcium phosphate method. Forty-eight hours post-transfection, cells were lysed in 10 cell pellet volumes of buffer (20 mM Tris-HCl [pH 7.5], 0.5 mM EDTA, 150 mM NaCl, 10 mM KCl, 0.5% Triton, 1.5 mM MgCl2 , 10% glycerol, 10 mM; 0.2 mM PMSF; 10 mM β-mercaptoethanol) 30 min at 4 °C on wheel, followed by 30 min of centrifugation at 13,200 rpm. Whole-cell extracts were incubated with anti-FLAG beads (Sigma-Aldrich) and immunoprecipitates eluted using an excess of FLAG peptide followed by WB or DNA content analysis. For DNA-IP, DNA from input and eluates were extracted using Phenol/Chloroform/Isoamyl pH 8 (12/12/1). qPCR analysis was performed using specific primers and data normalized for input DNA. Disruption of protein/nucleic acid interactions was achieved by preparing whole cell extracts in the presence of 0.1 mg/ml Ethidium Bromide (Sigma-Aldrich) and 0.1 U/μl DNaseI (Sigma-Aldrich).
2.5. Retrotransposition Assay
After siRNA transfection or SLX4 overexpression, cells were transfected with LINE-1 reporter plasmids using the FuGene-6 transfection reagent (Promega). In cases where neomycin reporter plasmids were used, cells were grown in the presence of G418 (600 μg/ml) for 18 days prior to fixation with 3.2% paraformaldehyde and crystal violet staining. For dual luciferase retrotransposition assays, luminescence was measured 6 days post-transfection using the Dual-Luciferase Reporter Assay System (Promega), following the manufacturers instructions.
2.6. Analysis of Interferon Secretion
RA3331SLX4 cells were grown for 48 h in the presence of conditioned medium from RA3331 and RA3331SLX4 , pre-incubated or not with IFNα, IFNβ (pbl assay science) and IFNγ neutralizing antibody (Abcam) for 2 h. A549 cells were treated with conditioned medium from RA3331, RA3331SLX4 or RA3331SLX4ΔSAP for 24 h. IFN response was measured by RT-qPCR.
2.7. Immunofluorescence
Cells were seeded on fibronectin-coated coverslips (Sigma-Aldrich). Fixation was performed in 80% methanol followed by 1 h blocking in PBS, 0.1% Tween, 1% BSA. ssDNA was visualized using anti-ssDNA (Abcam) at 1:50 dilution in PBS, 0.1% Tween, 1% BSA followed by incubation with anti-mouse Alexa 488-conjugated secondary antibody (Life Technologies). When indicated, S1 Nuclease digestion was performed before ssDNA staining. Nuclei were stained with DAPI and images were collected on Leica DM6000 or Zeiss Apotome microscopes.
3. Results
3.1. Accumulation of Cytoplasmic ssDNA Leads to Interferon Production in SLX4 Deficiency
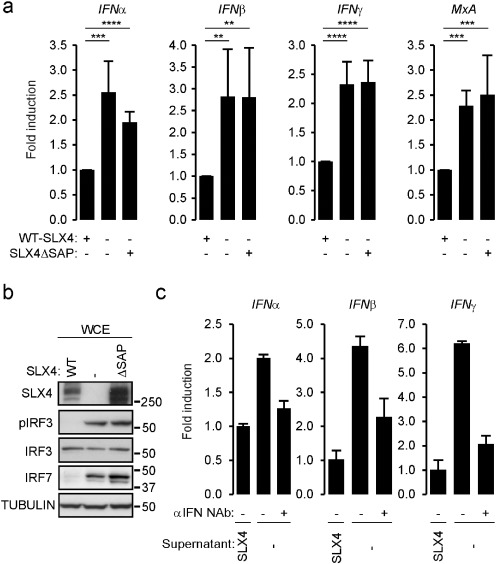
We previously reported (Laguette et al., 2014 ) that absence of SLX4 and of MUS81 both lead to upregulated type I IFN (IFNα and IFNβ) and IFN stimulated gene (MxA) mRNA expression. Here, we wished to determine whether the control of spontaneous pro-inflammatory cytokine production by SLX4 and MUS81 results from independent activities or a coordinated activity within the SLX4com. To this aim, we used the previously described cell lines derived from SLX4-deficient FA patient cells, complemented with either an empty vector (RA3331), a WT-SLX4 allele (RA3331SLX4 ), or a SLX4 allele harboring a deletion of the SAP domain (SLX4ΔSAP) responsible for interaction with MUS81-EME1 (RA3331SLX4ΔSAP ) (Kim et al., 2013 ). This SLX4 construct is impaired in its ability to interact with MUS81-EME1 but can still bind other SLX4 partners, including ERCC4 (Fig. S1A and (Kim et al., 2013 )). Accordingly, RA3331SLX4ΔSAP cells present elevated sensitivity to drugs that cause replication fork collapse, but not to DNA crosslinking agents such as Mitomycin C (MMC) (Kim et al., 2013 ). Measuring type I, type II IFN and MxA mRNA levels by RT-qPCR showed spontaneous production in RA3331 cells that was prevented by expression of SLX4, but not of SLX4ΔSAP (Fig. 1 A). Additional Interferon Stimulated Genes (ISGs) were also found to be upregulated in RA3331, and RA3331SLX4ΔSAP (Fig. S1B). Of note, WT SLX4 expression was more efficient at decreasing the expression of some tested ISGs, indicating a potential contribution of additional SLX4 domains in the control of pro-inflammatory signals. Consistently, analysis of protein levels of IRF7, IRF3 and of the phosphorylated active form of IRF3 (pIRF3) by Western Blot (WB) reveals an accumulation of IRF7 and pIRF3 in RA3331 and RA3331SLX4ΔSAP as compared to RA3331SLX4 (Fig. 1 B). To investigate whether upregulation of IFN mRNA expression results in bona fide secretion of cytokines, experiments were performed where RA3331SLX4 cells treated with conditioned media from RA3331 displayed increased type I and type II IFN mRNA levels as compared to cells treated with media collected from RA3331SLX4 (Fig. 1 C). In addition, pre-incubating conditioned medium from RA3331 with human neutralizing antibodies against IFNα, IFNβ and IFNγ prevented the measured increase of IFN mRNA (Fig. 1 C). This is further confirmed by cultivating the A549 IFN-responsive cell line in the presence of conditioned medium collected from RA3331, RA3331SLX4 , and RA3331SLX4ΔSAP showing that conditioned media from RA3331 and RA3331SLX4ΔSAP but not RA3331SLX4 induced upregulation of MxA mRNA levels (Fig. S1C). This, together with previous observation that MUS81 depletion leads to upregulation of pro-inflammatory cytokines (Laguette et al., 2014 ), strongly suggests that recruitment of MUS81-EME1 to SLX4 is required for SLX4com-mediated repression of pro-inflammatory pathways.
|
|
|
Fig. 1. The SAP MUS81-EME1-interacting domain of SLX4 is required for repressing inflammatory signals. (A) IFNα, IFNβ, IFNγ and MxA mRNA levels were measured by RT-qPCR in RA3331 cells complemented either with WT-SLX4 (RA3331SLX4 ) or SLX4ΔSAP (RA3331SLX4ΔSAP ). Graphs present means (± SD) of triplicate measurement of 6 independent experiments expressed as fold change in mRNA expression relative to RA3331SLX4 . ****: p < 0.001; ***: p < 0.005; **: p < 0.01. (B) WCE from RA3331, RA3331SLX4 and RA3331SLX4ΔSAP were analyzed by WB using indicated antibodies. (C) IFNα, IFNβ and IFNγ mRNA levels were measured in RA3331SLX4 treated with conditioned medium from RA3331SLX4 or RA3331 pre-incubated or not with neutralizing antibody to human IFNα, IFNβ and IFNγ. Graph represents mean (± SD) mRNA levels of one representative experiment relative to cells treated with conditioned medium from RA3331SLX4 . See also Fig. S1. |
Because the SLX4com processes HIV-derived nucleic acids (Laguette et al., 2014 ), preventing its innate immune sensing, and absence of SLX4 leads to activation of pattern recognition pathways (Fig. S1B), we hypothesized that SLX4-associated nuclease activities may be involved in processing endogenous nucleic acids to avoid spontaneous pro-inflammatory cytokine production. Consequently, absence of SLX4 should lead to accumulation of cytoplasmic immunogenic nucleic acids. To test this hypothesis, we performed immunofluorescence staining of RA3331, RA3331SLX4 and RA3331SLX4ΔSAP cells using a ssDNA-specific antibody (Fig. 2 A–B). Enrichment in cytoplasmic staining was observed in RA3331 and RA3331SLX4ΔSAP cells but not in RA3331SLX4 cells. Additionally, absence of MUS81 in mouse embryonic fibroblasts (MEFMUS81 −/− ) also led to accumulation of cytosolic ssDNA as compared to WT MEFs (Fig. S2A). This shows that lack of SLX4 results in accumulation of cytosolic ssDNA and highlights the requirement of MUS81 binding to SLX4 for cytosolic DNA clearance. Of note, treatment with the S1 Nuclease that preferentially digests ssDNA, caused a net decrease of the observed cytoplasmic staining (Fig. S2B).
|
|
|
Fig. 2. Cytoplasmic accumulation of ssDNA is responsible for pro-inflammatory signals in SLX4 deficiency. (A) Immunofluorescence analysis was performed on RA3331, RA3331SLX4 and RA3331SLX4ΔSAP using ssDNA-specific antibody and DAPI nuclear staining. Images are representative of at least 4 independent experiments. (B) Quantification of staining observed in A. (C) Left panel: experimental scheme. Briefly, cytoplasmic extraction was performed on RA3331, RA3331SLX4 and RA3331SLX4ΔSAP followed by RNase cocktail (RNase A and RNase T1) and proteinase K treatments prior to DNA extraction and radiolabeling. DNA extracted from RA3331 cells was subjected or not to a subsequent S1 Nuclease treatment prior to separation on acrylamide gel and autoradiography. Right panel: autoradiography of a representative experiment. (D) WCE from RA3331 expressing Luciferase- , MYD88- , or STING -targeting inducible shRNAs were analyzed by WB using indicated antibodies. (E) Graphs present RT-qPCR quantification of knock-down efficiency, measured using specific primers, as mean (± SD) mRNA expression relative to cells expressing Luciferase -targeting shRNA from 3 independent experiments. (F) MxA, RIG-I, TLR4 and TLR2 mRNA levels were measured in cells described in D. Graphs present means (± SD) of 3 independent experiments as fold increase mRNA expression relative to cells expressing a Luciferase targeting shRNA. (G) WCE from RA3331 expressing Luciferase , STING , cGAS or IFI16 -targeting inducible shRNAs were analyzed by WB using indicated antibodies. (H) MxA, STING, cGAS and IFI16 mRNA levels were measured in cells described in G. Graphs present means (± SD) of 3 independent experiments expressed as fold change in mRNA expression relative to cells expressing a Luciferase targeting shRNA. ****: p < 0.001; ***: p < 0.005; **: p < 0.01. See also Fig. S2. |
Since it has been previously reported that the cytoplasm of cancer cells contains a mix of ssDNA, dsDNA and DNA:RNA hybrids (Koo et al., 2015 and Shen et al., 2015 ), we questioned the nature of the nucleic acids present in the cytoplasm of SLX4-deficient cells. To this aim, γP32-labeling of cytoplasmic DNA from RA3331, RA3331SLX4 and RA3331SLX4ΔSAP cells was performed (Fig. 2 C, left panel). Cytoplasmic and nuclear extracts were analyzed by WB to check for nuclear contamination of cytoplasmic fraction (Fig. S2C). Autoradiography revealed increased presence of nucleic acids in the cytoplasmic fraction of RA3331 and RA3331SLX4ΔSAP as compared to RA3331SLX4 (Fig. 2 C, right panel). Importantly, treatment of the material obtained from RA3331 and RA3331SLX4ΔSAP cells with S1 Nuclease leads to reduction of the signal (Fig. 2 C and S2D) confirming the presence of ssDNA in the cytoplasm of SLX4-deficient cells. Similarly, treatment with dsDNAse or RNaseH, led to decreased levels of cytoplasmic nucleic acid in RA3331 and RA3331SLX4ΔSAP (Fig. S2E and S2F), indicating that the cytosol of SLX4-deficient cells contains dsDNA, ssDNA and DNA:RNA hybrids and that SLX4-MUS81 interaction is required to prevent their accumulation. Of note, it has been reported that immunogenic nucleic acids present in the cytoplasm of cancer cells are generated in the nucleus (Hartlova et al., 2015 and Yang et al., 2007 ). Interestingly, radiolabeling experiments of DNA extracted from the soluble nuclear fraction of RA3331, RA3331SLX4 and RA3331SLX4ΔSAP , followed by digestion with S1 Nuclease (Fig. S2G) shows the presence of ssDNA in the nuclear fraction of RA3331 and RA3331SLX4ΔSAP , but not RA3331SLX4 .
To investigate whether the DNA species present in the cytoplasm of RA3331 cells are responsible for IFN production, RA3331 cells were engineered to stably express an inducible short hairpin RNA (shRNA) targeting STING (TMEM173 ). To assess the contribution of extracellular nucleic acids to the signaling pathway triggered in the absence of SLX4, we also included shRNAs targeting MYD88 ( Fig. 2 D–F). As a control, RA3331 cells expressing an inducible shRNA targeting the Luciferase gene were also engineered. Following 6 days of shRNA induction, IRF3 phosphorylation status was assessed in whole cell extracts (WCE) (Fig. 2 D), while knockdown efficiency (Fig. 2 E), MxA and additional IFN-inducible genes mRNA levels (Fig. 2 F) were measured by RT-qPCR. Knockdown of STING but not MYD88 led to decreased pIRF3 and IFN signaling (Fig. 2 D–F). This indicates that cytoplasmic DNA present in SLX4 deficient cells is sensed through the STING pathway to induce IFN production. To investigate whether previously identified nucleic acid sensors of the STING pathway are required for pro-inflammatory signaling in SLX4 deficiency, we engineered cells where cGAS or IFI16 (Dempsey and Bowie, 2015 ) silencing is inducible and measured the levels of pIRF3 and phosphorylated active TBK1 kinase (pTBK1), known to be recruited by STING, by WB and of MxA by RT-qPCR (Fig. 2 G–H). Knockdown efficiency was assessed by RT-qPCR (Fig. 2 H). Silencing cGAS but not IFI16 led to decreased pro-inflammatory signaling (Fig. 2 G) and MxA induction (2H), highlighting a role for the cGAS-STING pathway in detecting cytosolic nucleic acids in SLX4-deficient cells. However, because of poor TRIF knockdown (not shown), we cannot exclude a contribution of endosomal nucleic acids in the activation of the inflammatory signal observed in SLX4-deficient cells.
3.2. The SLX4 Complex Interacts With the LINE-1 Retrotansposition Complex
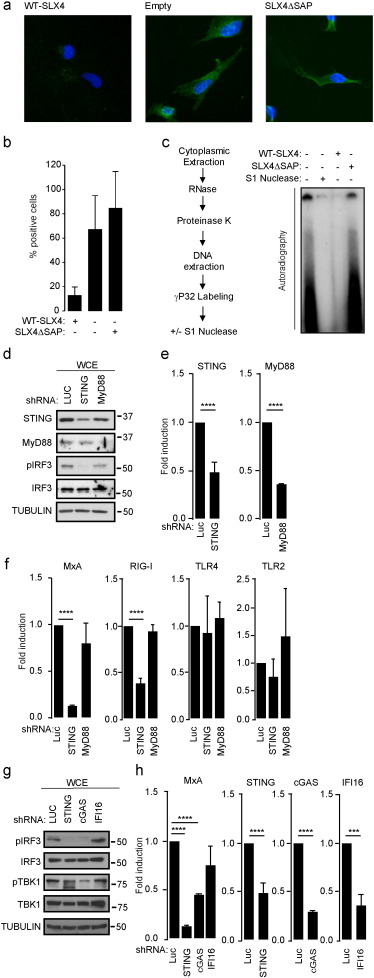
We have shown that absence of SLX4 leads to HIV-derived DNA accumulation (Laguette et al., 2014 ), suggesting that SLX4-associated endonucleases are involved in processing the latter. This is reminiscent of what has been reported for the TREX1 exonuclease (Yan et al., 2010 ). Because TREX1 inhibits LINE-1 retrotransposition and cell-intrinsic initiation of autoimmunity (Stetson et al., 2008 ), we hypothesized that absence of SLX4 may similarly lead to LINE-1 deregulation, which could account for subsequent cytoplasmic DNA accumulation. To test this hypothesis, DNA was extracted from the cytoplasmic fraction of RA3331 and RA3331SLX4 (Fig. S3A). LINE-1 DNA was either quantified by qPCR (Fig. 3 A) or visualized by Southern Blot (Fig. 3 B). We thereby observed an increase of LINE-1 DNA in the cytoplasmic fraction of RA3331 as compared to RA3331SLX4 (Fig. 3 A–B). Interestingly, this observation is concomitant to an increase of LINE-1 mRNA as measured by RT-qPCR (Fig. S3B), suggesting that this DNA accumulation is linked to increased LINE-1 activity. Indeed, qPCR performed using previously described primers (Boissinot et al., 2000 ) that allow the amplification of active LINE-1 (Fig. 3 C) and sequencing of obtained PCR products (Fig. S3C) confirmed the presence of DNA originating from the Ta-1 subfamily of active LINE-1 elements.
|
|
|
Fig. 3. The SLX4com regulates cytoplasmic accumulation of LINE-1 DNA. (A) LINE-1 DNA was measured by qPCR in the cytoplasmic fraction of RA3331 or RA3331SLX4 . #1: primers targeting the 5′ of ORF1 . #2: primers targeting the 3′ of ORF2 . (B) Cytoplasmic DNA extracted from RA3331 or RA3331SLX4 were analyzed by Southern Blot using LINE-1 specific probe. The right panel shows the molecular weight ladder. (C) Same as in A, except that primers allowing the amplification of active LINE-1 elements were used. #3 and #4: primer pairs targeting the 3′ UTR sequence of LINE-1 Ta-1 sub-family. (D) Constructs used in E-G and I. (E) FLAG/HA-WT-SLX4 or FLAG/HA-SLX4ΔSAP were FLAG-immunoprecipitated from 293T cells in the presence of a LINE-1-expressing plasmid. Bound material was peptide-eluted. Input and eluates were analyzed by WB using indicated antibodies. (F) FLAG-WT-MUS81 or FLAG-MUS81ΔWH were FLAG-immunoprecipitated from 293T cells in the presence of a LINE-1-expressing plasmid. Bound material was peptide-eluted. Input and eluates were analyzed as in E. (G) FLAG-WT-MUS81 was purified as in E in the presence or absence of 0.1 mg/ml EtBr and 0.1 U/μl DNaseI. Input and eluates were analyzed as in E. (H) LINE-1 construct used in I. Primers are indicated in red. (I) FLAG/HA-WT-SLX4, FLAG/HA-SLX4ΔSAP, FLAG-WT-MUS81 or FLAG-MUS81ΔWH were purified as in E prior to DNA extraction. Samples were analyzed by qPCR using primers indicated in H. See also Fig. S3. |
Because SLX4 regulates LINE-1-derived cytoplasmic DNA levels, we next asked whether the SLX4com interacts with the LINE-1 retrotransposition complex and whether this interaction is DNA-dependent. LINE-1 expresses a 6 kb bicistronic mRNA encoding two proteins that are essential for retrotransposition: the ORF1p RNA binding protein and the ORF2p protein that possess endonuclease and RT activities. During LINE-1 retrotransposition, ORF1p and ORF2p bind to LINE-1 RNA, forming a ribonucleoprotein particle competent for the reverse transcription of LINE-1 RNA into dsDNA that may integrate into genomic DNA. Using a previously described LINE-1 reporter construct (LINE-1-NEO) where the ORF1p protein is C-terminally T7-tagged (Fig. 3 D) (Doucet et al., 2010 ), we assessed the interaction between SLX4com and ORF1p. LINE-1-NEO was co-expressed with N-terminally FLAG- and HA-tagged SLX4 (F/H-SLX4) or SLX4ΔSAP (F/H-SLX4ΔSAP) in 293T cells (Fig. 3 E). WCE were subjected to FLAG-immunoprecipitation. Input and FLAG-eluted material were analyzed by WB. As shown in Fig. 3 E, ORF1p was efficiently recovered in F/H-SLX4, but not in F/H-SLX4ΔSAP immunoprecipitation. This shows that SLX4 interacts with ORF1p through its SAP domain. To explore a potential role of MUS81-EME1 in this interaction, FLAG-tagged MUS81 (F-MUS81) (Fig. 3 F) was co-expressed with the LINE-1-NEO reporter plasmid in 293T cells. WCE were subjected to FLAG-immunoprecipitation and the presence of ORF1p was analyzed by WB. Fig. 3 F shows that SLX4, EME1 and ORF1p were recovered in F-MUS81 immunoprecipitation. To evaluate the contribution of MUS81-DNA binding domain for interaction with ORF1p, we used the MUS81ΔWH mutant (Fadden et al., 2013 ) that harbors a deletion of its nucleic acid-binding domain (Fig. 3 D). While F-MUS81ΔWH and F-MUS81 displayed similar ability to interact with SLX4 and EME1, interaction of F-MUS81ΔWH with ORF1p was impaired (Fig. 3 F), showing that interaction of SLX4com with LINE-1 depends on MUS81 DNA-binding domain. Interestingly, when F-MUS81 immunoprecipitation was performed in the presence of DNaseI and Ethidium Bromide (EtBr) that disrupt nucleic acid/protein interactions, MUS81 and ORF1p interaction was lost (Fig. 3 G). Altogether, these experiments show that SLX4com binding to ORF1p is DNA-dependent and requires MUS81 DNA-binding domain.
We next wished to assess the binding of LINE-1-derived DNA to the SLX4com. To this aim, we used the LINE-1-GFP reporter plasmid (Fig. 3 H and (Coufal et al., 2009 )). In this construct, a green fluorescent protein (gfp ) gene is inserted in the 3′ untranslated region (UTR) of LINE-1 in antisense orientation relative to LINE-1 expression, under the control of an independent promoter. The sequence of the gfp is disrupted by an intronic sequence and GFP expression can only be measured when the retroelement has undergone a full cycle of retrotransposition, involving transcription, splicing, reverse transcription and integration. We designed qPCR primers that flank the intronic sequence to discriminate spliced (136 bp) from unspliced LINE-1-derived DNA (Fig. 3 H). This construct was co-expressed in 293T cells together with F/H-SLX4, F/H-SLX4ΔSAP, F-MUS81 or F-MUS81ΔWH prior to whole cell extraction and FLAG-immunopurification coupled to peptide elution. Efficient immunoprecipitation of target proteins was verified by WB (Fig. S3D). Eluted material was subjected to DNA extraction followed by qPCR. We observed that LINE-1 DNA was efficiently co-immunoprecipitated with F/H-SLX4 and F-MUS81, but not with SLX4ΔSAP and MUS81ΔWH (Fig. 3 I). We verified that the amplified LINE-1 fragment was indeed 136 bp and that similar levels of LINE-1 input material was used (Fig. S3E–F). Altogether, our observations strongly suggest that the SLX4com binds to LINE-1 reverse-transcribed DNA via interaction with the MUS81-EME1 endonuclease module.
3.3. The SLX4 Complex Negatively Regulates LINE-1 Retrotransposition
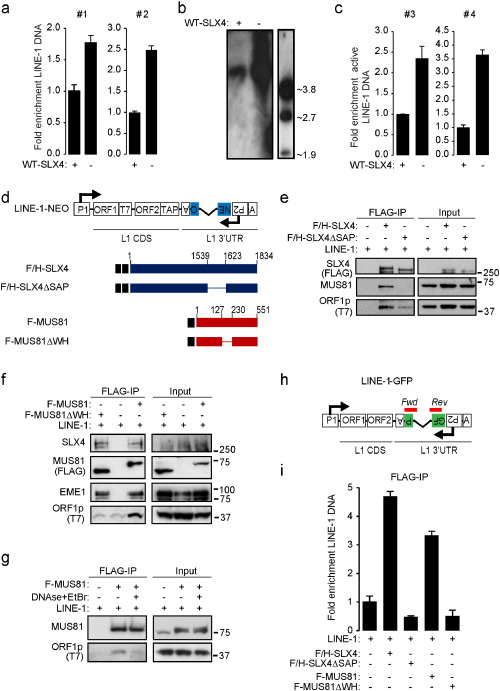
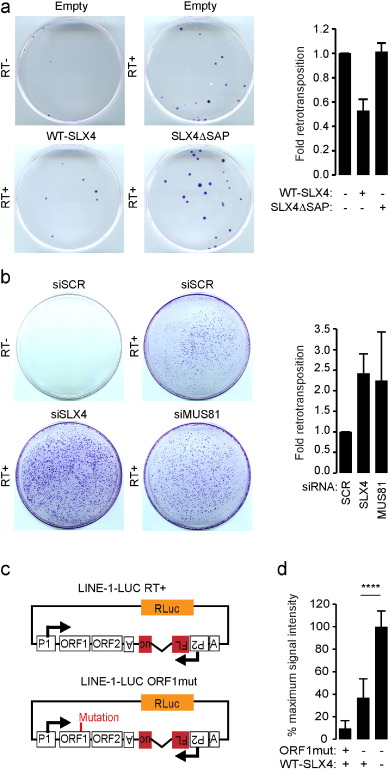
We next wished to investigate the consequences of SLX4com binding to LINE-1 retrotransposition complexes. To this aim, the LINE-1-NEO reporter construct (Fig. 3 D) was transfected in HeLa cells expressing empty vector, WT-SLX4 or SLX4ΔSAP. LINE-1-NEO is similar to the LINE-1-GFP construct, except that the gfp sequence is replaced by a neomycin resistance gene and neomycin resistance arises when the retroelement has undergone a full retrotransposition cycle. As negative control, a LINE-1 mutant construct unable to perform reverse transcription (RT-) was also included in the assay. Crystal violet staining of neomycin-resistant colonies showed that expression of WT-SLX4, but not of SLX4ΔSAP (Fig. S4A), decreases the number of colonies as compared to cells where empty vector was expressed ( Fig. 4 A). This shows that SLX4 is a negative regulator of LINE-1 retrotransposition, a function that requires its SAP domain. We next performed similar experiments except that we silenced SLX4 or MUS81 using small interfering RNA (siRNA) prior to transfection of the LINE-1-NEO reporter construct (Fig. 4 B and S4B–C). We observed that absence of SLX4 or MUS81 both lead to increased LINE-1 retrotransposition (Fig. 4 B). Finally, to confirm these observations, we performed retrotransposition assays in RA3331 cell lines. We used a previously described highly sensitive retrotransposition dual Luciferase reporter system (Xie et al., 2011 ) where a Firefly luciferase (FLuc ) gene, disrupted by an intronic sequence, is inserted in the 3′ untranslated region (UTR) of LINE-1 in antisense orientation relative to LINE-1 expression, under the control of an independent promoter and a Renilla luciferase (RLuc ) cassette is inserted in the same backbone to allow for normalization ( Fig. 4 C). As negative control, a mutant construct unable to perform retrotransposition (ORF1mut) was included in the assay (Fig. 4 C). These constructs were transfected in RA3331 and RA3331SLX4 cells and retrotransposition efficiency was quantified as the ratio of FLuc/RLuc. We thereby observed that complementing the RA3331 cells with WT-SLX4 caused a 2.5 fold decrease in the measured Luciferase activity (Fig. 4 D), reflecting decreased ability of LINE-1 to retrotranspose in the presence of WT-SLX4. Of note, to control whether sole expression of SLX4 could affect the expression of a neomycin resistance gene, we transfected a shRNA containing a neomycin resistance gene in HeLa cells expressing empty vector or WT-SLX4. After crystal violet staining, we observed that overexpression of SLX4 does not decrease the expression of neomycin resistance gene (Fig. S4D).
|
|
|
Fig. 4. The SLX4 complex negatively regulates LINE-1 retrotransposition. (A) LINE-1 retrotransposition assay was performed in HeLa cells overexpressing either WT-SLX4 or SLX4ΔSAP together with a LINE1-NEO reporter plasmid. Following neomycin selection, resistant clones were crystal violet stained. Left panel presents one representative experiment. Graph represents crystal violet staining quantified over 3 independent experiments relative to cells expressing an empty vector. (B) LINE-1 retrotransposition assay was performed as in A, except that HeLa cells were treated with scrambled siRNA or siRNAs targeting SLX4 or MUS81. Graphs represent quantification as in A. (C) LINE-1 constructs used in D. LINE-1-ORF1mut contains the JM111 mutation in the ORF1 sequence that abolishes retrotransposition. (D) LINE-1 retrotransposition assay performed in RA3331 or RA3331SLX4 transfected with plasmids described in C. Means (± SD) Luciferase activity of triplicate measurement from 4 independent experiments is presented as a percentage of maximum signal intensity. ****: p < 0.001. See also Fig. S4. |
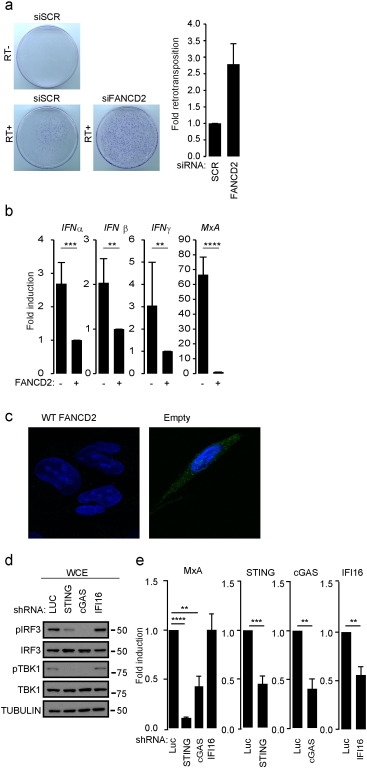
We next wanted to assess whether LINE-1 regulation is specific to the SLX4com or is a general function for the FA DNA repair pathway. To this aim, we tested the role of FANCD2 and FANCA in regulating LINE-1 retrotransposition. FANCD2 exerts a crucial role by orchestrating the activation of downstream effectors of the FA DNA repair pathway, including SLX4, while FANCA is a component of the FA core complex responsible of the activation of FANCD2-FANCI (Longerich et al., 2014 ). Retrotransposition assays using the LINE-1-NEO reporter plasmid in the presence or absence of siRNAs targeting FANCD2 or FANCA (Fig. 5 A, S5A–B) showed that their absence leads to increased LINE-1 retrotransposition. We next investigated whether absence of FANCD2 also leads to cytokine upregulation. To this aim, we measured IFNα, IFNβ, IFNγ and MxA mRNA levels by RT-qPCR in FANCD2-deficient or proficient cells. We observed that FANCD2-deficient cells, produced type I IFN, type II IFN and MxA mRNA as compared to their WT-FANCD2-complemented counterparts (Fig. 5 B). In agreement, immunofluorescence analysis of cytoplasmic ssDNA content, revealed the accumulation of ssDNA in the cytoplasm of FANCD2-deficient cells, as compared to their WT-FANCD2 complemented counterparts (Fig. 5 C, S5C). Importantly, as in SLX4-deficient cells, we observed that this spontaneous production of cytokines relies on the cGAS-STING pathway (Fig. 5 D–E). Altogether, our data demonstrate a correlation between absence of a functional FA DNA repair pathway, elevated pro-inflammatory cytokine production through activation of the cGAS-STING pathway (Fig. 2 and Fig. 5 ) and upregulated LINE-1 activity (Fig. 4 and Fig. 5 ).
|
|
|
Fig. 5. Absence of FANCD2 leads to upregulated STING-dependent IFN production. (A) Retrotransposition was measured and results presented as in Fig. 4 A except that cells were treated with scrambled siRNA or siRNA targeting FANCD2. (B) IFNα, IFNβ, IFNγ and MxA mRNA levels were measured in FANCD2 deficient cells and on their WT-FANCD2 complemented counterparts. Graphs present means (± SD) from triplicate measurement of 6 independent experiments expressed as fold mRNA expression relative to FANCD2 proficient cells. (C) Immunofluorescence was performed on FANCD2 deficient or proficient cells as in Fig. 2 A. (D) WCE from PD20 expressing Luciferase , STING , cGAS or IFI16 -targeting inducible shRNAs were analyzed by WB using indicated antibodies. (E) MxA, STING, cGAS and IFI16 mRNA levels were measured in cells described in D. Graphs present means (± SD) of 4 independent experiments expressed as fold change in mRNA expression relative to cells expressing Luciferase targeting shRNA. ****: p < 0.001; ***: p < 0.005; **: p < 0.01. See also Fig. S5. |
3.4. Endogenous Reverse Transcriptase Activities Contribute to Generate Immunogenic Nucleic Acid Species
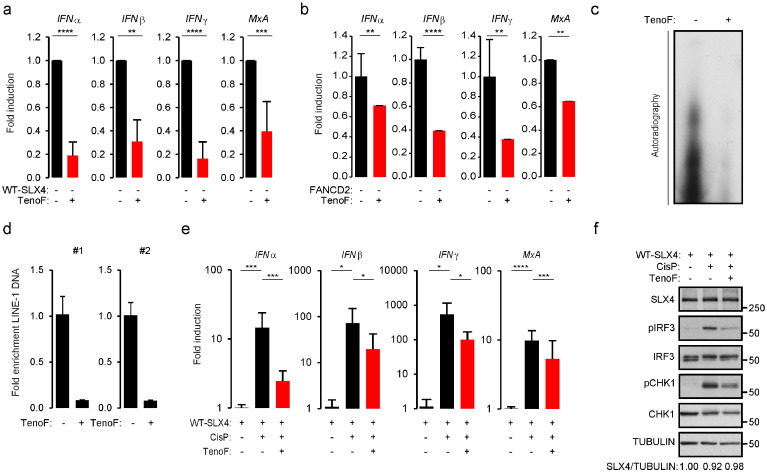
Our observations suggest a correlation between upregulated LINE-1 activity and increased cytokine production in FA. To further strengthen this correlation, we determined the effect of the Tenofovir (TenoF) nucleoside analogue RT inhibitor, known to prevent LINE-1 retrotransposition (Jones et al., 2008 ), on IFN production in FA. Treatment of RA3331 with TenoF, at doses known to inhibit LINE-1 retrotransposition (Fig. S6A), showed significant decrease of IFNα, IFNβ, IFNγ and MxA mRNA levels (Fig. 6 A). Similar results were obtained with FANCD2-deficient cells (Fig. 6 B). This indicates that inhibiting endogenous RT activities prevents spontaneous pro-inflammatory cytokine production in FA-derived cells. This further suggests that in FA, uncontrolled retrotransposition may be a source of immunogenic nucleic acids. To test this hypothesis, DNA was extracted from the cytoplasmic fraction of RA3331 cells treated or not with TenoF prior to radiolabeling (Fig. 6 C) or qPCR using LINE-1-specific primers (Fig. 6 D). We thus observed that such treatment caused a decrease of both total cytoplasmic and LINE-1-derived DNA.
|
|
|
Fig. 6. Upregulated retrotransposition leads to pro-inflammatory signaling. (A) IFNα, IFNβ, IFNγ and MxA mRNA levels were measured in RA3331 cells following 48 h treatment with 125 μM TenoF. Graphs present means (± SD) from triplicate measurement of 6 independent experiments expressed as fold change in mRNA expression relative to RA3331. ****: p < 0.001; ***: p < 0.005; **: p < 0.01. (B) IFNα, IFNβ, IFNγ and MxA mRNA levels were measured in FANCD2-deficient cells following 48 h treatment with 5 μM TenoF. Graphs present means (± SD) from triplicate measurement of 6 independent experiments expressed as fold change in mRNA expression relative to untreated cells. ****: p < 0.001; ***: p < 0.005; **: p < 0.01. (C) Cytoplasmic DNA was prepared from RA3331 cells following 8 h treatment with 125 μM TenoF, radiolabeled and analyzed by autoradiography. (D) DNA was prepared as described in C and analyzed by qPCR using primers described in Fig. 3 A. Graph presents data from a representative experiment. (E) IFNα, IFNβ, IFNγ and MxA mRNA levels were measured in RA3331SLX4 treated or not with 125 μM TenoF 24 h prior to treatment with 10 μM Cisplatin. Graphs present means (± SD) from 3 independent experiments as fold change in mRNA expression, relative to untreated cells. ****: p < 0.001; ***: p < 0.005; **: p < 0.01; *: p < 0.05. (F) WCE from cells treated as in E were analyzed by WB. See also Fig. S6. |
It has been shown that DNA damage-inducing agents, including the Cisplatin (CisP) DNA crosslinker, can induce LINE-1 activation (Rudin and Thompson, 2001 ). Thus, we tested the impact of DNA damage-inducing agents, such as CisP, MMC and hydroxyurea (HU), on the activation of pro-inflammatory pathways (Fig. 6 E–F and S6B–C). We observed that CisP treatment caused increased IFNα, IFNβ, IFNγ and MxA mRNA levels and pIRF3 accumulation, which was prevented by treatment of cells with TenoF 24 h prior to CisP treatment (Fig. 6 E–F). Of note, similar observations were made upon treatment with MMC and HU (Fig. S6B–C). To exclude that TenoF treatment would affect the ability of CisP, MMC and HU to cause DNA lesions, we assessed CHK1 phosphorylation status. Indeed, the CHK1 kinase becomes activated via phosphorylation upon replication stress, as induced by treatment with CisP, MMC and HU. We observed that TenoF treatment marginally affected CHK1 phosphorylation (Fig. 6 F and S6B–C). We also verified that TenoF alone did not significantly affect pCHK1 and SLX4 expression levels (Fig. S6D–E). This implies that TenoF inhibits activation of pro-inflammatory pathways triggered by tested DNA damage inducing drugs, independently from their ability to cause DNA lesions. Of note, we also observed that TenoF treatment weakly affected DNA crosslink- and cytokine-associated cell death of RA3331 cells (Fig. S6F–G). Thus, altogether, our observations strongly suggest that DNA damaging agents cause activation of pro-inflammatory pathways through endogenous reverse transcriptase activity including that of LINE-1.
4. Discussion
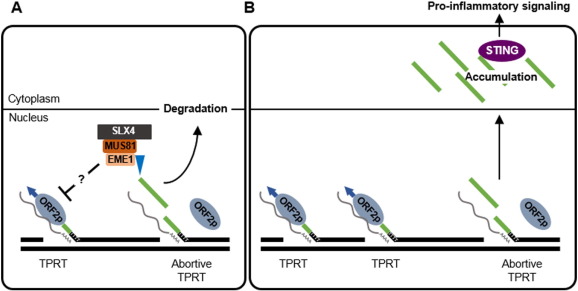
Chronic inflammation that fosters a favorable environment for tumor initiation can be generated by both cell-extrinsic and cell-intrinsic factors. The latter include inability to repair genomic lesions (Rodier et al., 2009 and Zheng et al., 2007 ). Indeed, recent work has provided evidence for the implication of DDR pathways in the regulation of spontaneous and pathogen-induced innate immune responses (Bregnard et al., 2014 ). However, there is as of today poor understanding of the underlying molecular mechanisms. Here we used FA as a model cancer susceptibility syndrome to investigate the origins of chronic inflammation. We show that elevated pro-inflammatory cytokine production in the absence of a functional FA DNA repair pathway results, at least in part, from upregulated LINE-1 activity. In this model, we observed that absence of SLX4 leads to upregulated LINE-1 retrotransposition, accumulation of immunogenic cytoplasmic DNA comprising active LINE-1-derived sequences and concomitant up-regulated pro-inflammatory cytokine production. This is further supported by the witnessed physical interaction between the SLX4com and the LINE-1 retrotransposition complex and decreased pro-inflammatory cytokine production upon RT inhibitor treatment. In addition, in SLX4-deficient cells, LINE-1 transcription and retrotransposition are increased and the nucleoplasm contains immunogenic nucleic acids that may subsequently be exported to the cytoplasm and recognized by the cGAS-STING pathway to induce pro-inflammatory cytokine production (Fig. 7 ). Requirement of cGAS for activation of pro-inflammatory pathways in these cells suggests that dsDNA species are involved in the signaling cascade. However, since we also observed the accumulation of ssDNA and DNA:RNA hybrids, one can speculate that such nucleic acid species may either be recognized by the same pathway, or by additional yet to be identified pathways. Moreover, additional transposable elements detected by the cGAS-STING pathway may induce IFN production in SLX4 deficiency (Zeng et al., 2014 ). Interestingly, it has also been reported that tumor-derived DNA can activate the STING pathway in innate immune cells within the tumor microenvironment (Woo et al., 2014 ) and that a STING-dependent mechanism is involved in the induction of an adaptive immune response following radiation (Deng et al., 2014 ). It would thus be interesting to investigate the influence of SLX4-deficient cells-derived DNA on adaptive immune responses.
|
|
|
Fig. 7. Inhibition of LINE-1 retrotransposition by the SLX4com prevents pro-inflammatory signaling. (A) The SLX4com inhibits LINE-1 retrotransposition by preventing accumulation of reverse transcribed LINE-1 DNA. LINE-1-derived DNA are likely subsequently degraded. (B) In the absence of the SLX4com, by-products of LINE-1 reverse-transcription accumulate in the cytoplasm. These are recognized by the cGAS-STING pathway to activate pro-inflammatory signaling. Grey lines represents: LINE-1 RNA; green lines: LINE-1 reverse-transcribed DNA. |
FA presents with progressive bone marrow failure, increased cancer susceptibility and elevated pro-inflammatory cytokine production. FA-associated bone marrow failure has been shown to, at least in part, result from exhaustion of chronic cytokine-stimulation of bone marrow-resident stem cells (Garaycoechea and Patel, 2014 ). While deregulated cytokine production contributes to these bone marrow-related hematological dysfunctions, it is possible that persistent secretion of IFN can also contribute to the advent of malignancies in FA. Here we show that SLX4 and its upstream activator FANCD2 directly repress LINE-1 retrotransposition and prevent accumulation of cytoplasmic nucleic acids. This suggests that the FA DNA repair pathway as a whole may be involved in regulating LINE-1 activity and subsequent pro-inflammatory cytokine production. This further implies that upregulated LINE-1 retrotransposition occurs in FA patients and that LINE-1 activity may play an important role in the advent of clinical manifestations of FA. This is particularly important because of the pivotal role of FANCD2 in the activation of the FA DNA repair pathway.
We also show that recruitment of MUS81-EME1 is required for SLX4com-mediated control of pro-inflammatory signaling. This suggests that, similar to what was previously reported for HIV biology (Laguette et al., 2014 ), the SLX4com likely represses LINE-1 retrotransposition through the endonucleolytic action of MUS81-EME1. Importantly, for the latter to acquire full endonuclease activity within the SLX4com, PLK1-dependent EME1 hyperphosphorylation is required, which occurs around the G2/M boundary. It would thus be interesting to investigate whether the control of endogenous retroelements mobility and the ability to repress pro-inflammatory cytokine production depend on the cell cycle status. Additional post-translational modifications, such as SUMOylation and PARylation, were recently shown to be required for SLX4 activity by ensuring its localization to sites of DNA damage (Gonzalez-Prieto et al., 2015 , Guervilly et al., 2015 and Ouyang et al., 2015 ). Requirement for such post-translational modifications in the recognition and processing of potential immunogenic nucleic acids remains to be investigated.
Interestingly, it has been previously reported that the bone marrow of FA patients is compromised because of accumulation of DNA damage in hematopoietic stem cells (Ceccaldi et al., 2012 ). Our findings imply that this may also be linked to upregulated LINE-1 activity. Indeed, LINE-1 mobility is mostly repressed in cells through 5′ UTR methylation (Woodcock et al., 1997 and Yu et al., 2001 ) and stem cells have a hypomethylated genome that allows enhanced LINE-1 activity (Wissing et al., 2012 ). Thus, absence of the FA pathway may also allow for pro-tumorigenic LINE-1 re-insertion events, especially in the light of the LINE-1 DNA deriving from the most active LINE-1 subfamily accumulating in the cytoplasm of SLX4-deficient cells. This raises the possibility that treatment with RT inhibitors, which inhibit the accumulation of immunogenic nucleic acids responsible for initiating pro-inflammatory cytokines production, may delay or prevent the onset of bone marrow failure in FA. In support, immunotherapy aiming at neutralizing a single pro-inflammatory cytokine (TNFα) has shown promising positive effects in patients (Mehta et al., 2012 and Miehsler et al., 2010 ). Accordingly, we observed that treatment with TenoF weakly decreased the cell death of SLX4-deficient cells following treatment with TNFα or DNA damaging drugs. Of note, bone marrow failure is a clinical trait shared by additional familial cancer susceptibility syndromes (Parikh and Bessler, 2012 ) that could also be linked to deregulated endogenous RT.
Finally, the key role played by chronic inflammation in the progression of cancer is highlighted by the fact that anti-inflammatory drugs when administered to patients can reduce the risk of progression to metastasis (Markowitz, 2007 ). One interesting finding from our work is that treatment with TenoF prevents inflammation caused by DNA-damaging agents such as those used in chemotherapy. This is of particular importance because it has been reported that treatment with chemotherapy may generate a pro-inflammatory environment that would select cells that are resistant to treatment (de Visser and Jonkers, 2009 and Koti et al., 2015 ). Our data confirm the previous observation that certain DNA-damage inducing drugs can cause LINE-1 reactivation (Rudin and Thompson, 2001 ) and enforce the contribution of endogenous RT activities to inflammation. Furthermore, we observed a weak but reproducible decrease of pCHK1 levels in cells treated with RT inhibitors in the presence of DNA-damaging drugs. This is in agreement with previous reports of LINE-1 activity poising a mutagenic threat to the genome because of their ability to integrate at distant genomic locations. Thus, altogether our in vitro observations suggest that combination of chemotherapy with RT inhibitors could prevent the establishment of an inflammatory tumor microenvironment that would favor chemoresistance and decrease the risk of genomic instability. Thus, our study may open perspectives for the treatment of FA patients but also for reducing adverse effects of chemotherapy.
Conflicts of Interest
We declare no conflict of interest.
Author Contributions
All authors performed experiments. CB, JG, MB and NL discussed and interpreted obtained results. CB and JG equally contributed to the design and performing of experiments. CB and JG assembled the figures. MB and NL wrote the manuscript.
Acknowledgments
We thank Dr. Smogorzewska for the gift of RA3331 cell lines, Dr. Rosselli for PD20 cells, Dr. N. Gilbert for LINE-1 constructs, and Dr. W. An for the LINE-1-LUC construct. We thank Drs. Dufourt, Cristofari and Gilbert for fruitful discussions. We thank members of the Molecular Basis of Cancer-Related Inflammation laboratory for critical reading of the manuscript. Work in NL laboratory was supported by the ERC (637763 ), ANRS (109560 ) and by “la Fondation ARC pour la Recherche sur le Cancer” (PJA20141201605 ). Work in MB laboratory was supported by ERC (250333 ). CB was supported by ERC (250333 ) followed by an ANRS fellowship (14065 ). JG and SD were supported by the ERC (637763 ).
Appendix A. Supplementary Data
Supplementary material.
References
- Babushok and Kazazian, 2007 D.V. Babushok, H.H. Kazazian Jr.; Progress in understanding the biology of the human mutagen LINE-1; Hum. Mutat., 28 (2007), pp. 527–539
- Bald et al., 2014 T. Bald, T. Quast, J. Landsberg, M. Rogava, N. Glodde, D. Lopez-Ramos, J. Kohlmeyer, S. Riesenberg, D. van den Boorn-Konijnenberg, C. Homig-Holzel, et al.; Ultraviolet-radiation-induced inflammation promotes angiotropism and metastasis in melanoma; Nature, 507 (2014), pp. 109–113
- Bogliolo and Surralles, 2015 M. Bogliolo, J. Surralles; Fanconi anemia: a model disease for studies on human genetics and advanced therapeutics; Curr. Opin. Genet. Dev., 33 (2015), pp. 32–40
- Boissinot et al., 2000 S. Boissinot, P. Chevret, A.V. Furano; L1 (LINE-1) retrotransposon evolution and amplification in recent human history; Mol. Biol. Evol., 17 (2000), pp. 915–928
- Bregnard et al., 2014 C. Bregnard, M. Benkirane, N. Laguette; DNA damage repair machinery and HIV escape from innate immune sensing; Front. Microbiol., 5 (2014), p. 176
- Brzostek-Racine et al., 2011 S. Brzostek-Racine, C. Gordon, S. Van Scoy, N.C. Reich; The DNA damage response induces IFN; J. Immunol., 187 (2011), pp. 5336–5345
- Ceccaldi et al., 2012 R. Ceccaldi, K. Parmar, E. Mouly, M. Delord, J.M. Kim, M. Regairaz, M. Pla, N. Vasquez, Q.S. Zhang, C. Pondarre, et al.; Bone marrow failure in Fanconi anemia is triggered by an exacerbated p53/p21 DNA damage response that impairs hematopoietic stem and progenitor cells; Cell Stem Cell, 11 (2012), pp. 36–49
- Chen et al., 2010 S. Chen, J.A. Short, D.F. Young, M.J. Killip, M. Schneider, S. Goodbourn, R.E. Randall; Heterocellular induction of interferon by negative-sense RNA viruses; Virology, 407 (2010), pp. 247–255
- Coufal et al., 2009 N.G. Coufal, J.L. Garcia-Perez, G.E. Peng, G.W. Yeo, Y. Mu, M.T. Lovci, M. Morell, K.S. O'Shea, J.V. Moran, F.H. Gage; L1 retrotransposition in human neural progenitor cells; Nature, 460 (2009), pp. 1127–1131
- de Visser and Jonkers, 2009 K.E. de Visser, J. Jonkers; Towards understanding the role of cancer-associated inflammation in chemoresistance; Curr. Pharm. Des., 15 (2009), pp. 1844–1853
- Dempsey and Bowie, 2015 A. Dempsey, A.G. Bowie; Innate immune recognition of DNA: a recent history; Virology, 479-480 (2015), pp. 146–152
- Deng et al., 2014 L. Deng, H. Liang, M. Xu, X. Yang, B. Burnette, A. Arina, X.D. Li, H. Mauceri, M. Beckett, T. Darga, et al.; STING-dependent cytosolic DNA sensing promotes radiation-induced type I interferon-dependent antitumor immunity in immunogenic tumors; Immunity, 41 (2014), pp. 843–852
- Doucet et al., 2010 A.J. Doucet, A.E. Hulme, E. Sahinovic, D.A. Kulpa, J.B. Moldovan, H.C. Kopera, J.N. Athanikar, M. Hasnaoui, A. Bucheton, J.V. Moran, et al.; Characterization of LINE-1 ribonucleoprotein particles; PLoS Genet., 6 (2010)
- Du et al., 2014 W. Du, O. Erden, Q. Pang; TNF-alpha signaling in Fanconi anemia; Blood Cells Mol. Dis., 52 (2014), pp. 2–11
- Dufour et al., 2003 C. Dufour, A. Corcione, J. Svahn, R. Haupt, V. Poggi, A.N. Bekassy, R. Scime, A. Pistorio, V. Pistoia; TNF-alpha and IFN-gamma are overexpressed in the bone marrow of Fanconi anemia patients and TNF-alpha suppresses erythropoiesis in vitro; Blood, 102 (2003), pp. 2053–2059
- Fadden et al., 2013 A.J. Fadden, S. Schalbetter, M. Bowles, R. Harris, J. Lally, A.M. Carr, N.Q. McDonald; A winged helix domain in human MUS81 binds DNA and modulates the endonuclease activity of MUS81 complexes; Nucleic Acids Res., 41 (2013), pp. 9741–9752
- Ferguson et al., 2012 B.J. Ferguson, D.S. Mansur, N.E. Peters, H. Ren, G.L. Smith; DNA-PK is a DNA sensor for IRF-3-dependent innate immunity; Elife, 1 (2012), Article e00047
- Garaycoechea and Patel, 2014 J.I. Garaycoechea, K.J. Patel; Why does the bone marrow fail in Fanconi anemia?; Blood, 123 (2014), pp. 26–34
- Gasior et al., 2008 S.L. Gasior, A.M. Roy-Engel, P.L. Deininger; ERCC1/XPF limits L1 retrotransposition; DNA Repair (Amst), 7 (2008), pp. 983–989
- Gonzalez-Prieto et al., 2015 R. Gonzalez-Prieto, S.A. Cuijpers, M.S. Luijsterburg, H. van Attikum, A.C. Vertegaal; SUMOylation and PARylation cooperate to recruit and stabilize SLX4 at DNA damage sites; EMBO Rep., 16 (2015), pp. 512–519
- Guervilly et al., 2015 J.H. Guervilly, A. Takedachi, V. Naim, S. Scaglione, C. Chawhan, Y. Lovera, E. Despras, I. Kuraoka, P. Kannouche, F. Rosselli, et al.; The SLX4 complex is a SUMO E3 ligase that impacts on replication stress outcome and genome stability; Mol. Cell, 57 (2015), pp. 123–137
- Hartlova et al., 2015 A. Hartlova, S.F. Erttmann, F.A. Raffi, A.M. Schmalz, U. Resch, S. Anugula, S. Lienenklaus, L.M. Nilsson, A. Kroger, J.A. Nilsson, et al.; DNA damage primes the type I interferon system via the cytosolic DNA sensor STING to promote anti-microbial innate immunity; Immunity, 42 (2015), pp. 332–343
- Ishikawa and Barber, 2008 H. Ishikawa, G.N. Barber; STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling; Nature, 455 (2008), pp. 674–678
- Ishikawa et al., 2009 H. Ishikawa, Z. Ma, G.N. Barber; STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity; Nature, 461 (2009), pp. 788–792
- Jones et al., 2008 R.B. Jones, K.E. Garrison, J.C. Wong, E.H. Duan, D.F. Nixon, M.A. Ostrowski; Nucleoside analogue reverse transcriptase inhibitors differentially inhibit human LINE-1 retrotransposition; PLoS One, 3 (2008), Article e1547
- Kazazian et al., 1988 H.H. Kazazian Jr., C. Wong, H. Youssoufian, A.F. Scott, D.G. Phillips, S.E. Antonarakis; Haemophilia a resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man; Nature, 332 (1988), pp. 164–166
- Kim et al., 2013 Y. Kim, G.S. Spitz, U. Veturi, F.P. Lach, A.D. Auerbach, A. Smogorzewska; Regulation of multiple DNA repair pathways by the Fanconi anemia protein SLX4; Blood, 121 (2013), pp. 54–63
- Kondo et al., 2013 T. Kondo, J. Kobayashi, T. Saitoh, K. Maruyama, K.J. Ishii, G.N. Barber, K. Komatsu, S. Akira, T. Kawai; DNA damage sensor MRE11 recognizes cytosolic double-stranded DNA and induces type I interferon by regulating STING trafficking; Proc. Natl. Acad. Sci. U. S. A., 110 (2013), pp. 2969–2974
- Koo et al., 2015 C.X. Koo, K. Kobiyama, Y.J. Shen, N. LeBert, S. Ahmad, M. Khatoo, T. Aoshi, S. Gasser, K.J. Ishii; RNA polymerase III regulates cytosolic RNA:DNA hybrids and intracellular microRNA expression; J. Biol. Chem., 290 (2015), pp. 7463–7473
- Koti et al., 2015 M. Koti, A. Siu, I. Clement, M. Bidarimath, G. Turashvili, A. Edwards, K. Rahimi, A.M. Masson, J.A. Squire; A distinct pre-existing inflammatory tumour microenvironment is associated with chemotherapy resistance in high-grade serous epithelial ovarian cancer; Br. J. Cancer, 112 (2015), pp. 1215–1222
- Laguette et al., 2014 N. Laguette, C. Bregnard, P. Hue, J. Basbous, A. Yatim, M. Larroque, F. Kirchhoff, A. Constantinou, B. Sobhian, M. Benkirane; Premature activation of the SLX4 complex by Vpr promotes G2/M arrest and escape from innate immune sensing; Cell, 156 (2014), pp. 134–145
- Longerich et al., 2014 S. Longerich, J. Li, Y. Xiong, P. Sung, G.M. Kupfer; Stress and DNA repair biology of the Fanconi anemia pathway; Blood, 124 (2014), pp. 2812–2819
- Markowitz, 2007 S.D. Markowitz; Aspirin and colon cancer–targeting prevention?; N. Engl. J. Med., 356 (2007), pp. 2195–2198
- Matos and West, 2014 J. Matos, S.C. West; Holliday junction resolution: regulation in space and time; DNA Repair (Amst) (2014)
- Mehta et al., 2012 P.A. Mehta, J. Svahn, S.M. Davies, Q. Pang, R. Harris, P. Ghezzi, T. Lanza, E. Ferretti, P. Barabino, R. Mueller, et al.; Etanercept treatment in Fanconi anaemia; combined US and Italian experience; Br. J. Haematol., 158 (2012), pp. 809–811
- Miehsler et al., 2010 W. Miehsler, G. Novacek, H. Wenzl, H. Vogelsang, P. Knoflach, A. Kaser, C. Dejaco, W. Petritsch, M. Kapitan, H. Maier, et al.; A decade of infliximab: the Austrian evidence based consensus on the safe use of infliximab in inflammatory bowel disease; J. Crohns Colitis, 4 (2010), pp. 221–256
- Ouyang et al., 2015 J. Ouyang, E. Garner, A. Hallet, H.D. Nguyen, K.A. Rickman, G. Gill, A. Smogorzewska, L. Zou; Noncovalent interactions with SUMO and ubiquitin orchestrate distinct functions of the SLX4 complex in genome maintenance; Mol. Cell, 57 (2015), pp. 108–122
- Paludan, 2015 S.R. Paludan; Activation and regulation of DNA-driven immune responses; Microbiol. Mol. Biol. Rev., 79 (2015), pp. 225–241
- Parikh and Bessler, 2012 S. Parikh, M. Bessler; Recent insights into inherited bone marrow failure syndromes; Curr. Opin. Pediatr., 24 (2012), pp. 23–32
- Rakoff-Nahoum and Medzhitov, 2009 S. Rakoff-Nahoum, R. Medzhitov; Toll-like receptors and cancer; Nat. Rev. Cancer, 9 (2009), pp. 57–63
- Rigby et al., 2014 R.E. Rigby, L.M. Webb, K.J. Mackenzie, Y. Li, A. Leitch, M.A. Reijns, R.J. Lundie, A. Revuelta, D.J. Davidson, S. Diebold, et al.; RNA:DNA hybrids are a novel molecular pattern sensed by TLR9; EMBO J., 33 (2014), pp. 542–558
- Rodier et al., 2009 F. Rodier, J.P. Coppe, C.K. Patil, W.A. Hoeijmakers, D.P. Munoz, S.R. Raza, A. Freund, E. Campeau, A.R. Davalos, J. Campisi; Persistent DNA damage signalling triggers senescence-associated inflammatory cytokine secretion; Nat. Cell Biol., 11 (2009), pp. 973–979
- Rudin and Thompson, 2001 C.M. Rudin, C.B. Thompson; Transcriptional activation of short interspersed elements by DNA-damaging agents; Genes Chromosom. Cancer, 30 (2001), pp. 64–71
- Shen et al., 2015 Y.J. Shen, N. Le Bert, A.A. Chitre, C.X. Koo, X.H. Nga, S.S. Ho, M. Khatoo, N.Y. Tan, K.J. Ishii, S. Gasser; Genome-derived cytosolic DNA mediates type I interferon-dependent rejection of B cell lymphoma cells; Cell Rep., 11 (2015), pp. 460–473
- Stetson et al., 2008 D.B. Stetson, J.S. Ko, T. Heidmann, R. Medzhitov; Trex1 prevents cell-intrinsic initiation of autoimmunity; Cell, 134 (2008), pp. 587–598
- Volkman and Stetson, 2014 H.E. Volkman, D.B. Stetson; The enemy within: endogenous retroelements and autoimmune disease; Nat. Immunol., 15 (2014), pp. 415–422
- Wang and Smogorzewska, 2015 A.T. Wang, A. Smogorzewska; SnapShot: Fanconi anemia and associated proteins; Cell, 160 (2015) (354–354 e351)
- Wei et al., 2001 W. Wei, N. Gilbert, S.L. Ooi, J.F. Lawler, E.M. Ostertag, H.H. Kazazian, J.D. Boeke, J.V. Moran; Human L1 retrotransposition: cis preference versus trans complementation; Mol. Cell. Biol., 21 (2001), pp. 1429–1439
- Wissing et al., 2012 S. Wissing, M. Munoz-Lopez, A. Macia, Z. Yang, M. Montano, W. Collins, J.L. Garcia-Perez, J.V. Moran, W.C. Greene; Reprogramming somatic cells into iPS cells activates LINE-1 retroelement mobility; Hum. Mol. Genet., 21 (2012), pp. 208–218
- Woo et al., 2014 S.R. Woo, M.B. Fuertes, L. Corrales, S. Spranger, M.J. Furdyna, M.Y. Leung, R. Duggan, Y. Wang, G.N. Barber, K.A. Fitzgerald, et al.; STING-dependent cytosolic DNA sensing mediates innate immune recognition of immunogenic tumors; Immunity, 41 (2014), pp. 830–842
- Woodcock et al., 1997 D.M. Woodcock, C.B. Lawler, M.E. Linsenmeyer, J.P. Doherty, W.D. Warren; Asymmetric methylation in the hypermethylated CpG promoter region of the human L1 retrotransposon; J. Biol. Chem., 272 (1997), pp. 7810–7816
- Xie et al., 2011 Y. Xie, J.M. Rosser, T.L. Thompson, J.D. Boeke, W. An; Characterization of L1 retrotransposition with high-throughput dual-luciferase assays; Nucleic Acids Res., 39 (2011), Article e16
- Yan et al., 2010 N. Yan, A.D. Regalado-Magdos, B. Stiggelbout, M.A. Lee-Kirsch, J. Lieberman; The cytosolic exonuclease TREX1 inhibits the innate immune response to human immunodeficiency virus type 1; Nat. Immunol., 11 (2010), pp. 1005–1013
- Yang et al., 2007 Y.G. Yang, T. Lindahl, D.E. Barnes; Trex1 exonuclease degrades ssDNA to prevent chronic checkpoint activation and autoimmune disease; Cell, 131 (2007), pp. 873–886
- Yu et al., 2001 F. Yu, N. Zingler, G. Schumann, W.H. Stratling; Methyl-CpG-binding protein 2 represses LINE-1 expression and retrotransposition but not Alu transcription; Nucleic Acids Res., 29 (2001), pp. 4493–4501
- Zeng et al., 2014 M. Zeng, Z. Hu, X. Shi, X. Li, X. Zhan, X.D. Li, J. Wang, J.H. Choi, K.W. Wang, T. Purrington, et al.; MAVS, cGAS, and endogenous retroviruses in T-independent B cell responses; Science, 346 (2014), pp. 1486–1492
- Zhang et al., 2011 X. Zhang, T.W. Brann, M. Zhou, J. Yang, R.M. Oguariri, K.B. Lidie, H. Imamichi, D.W. Huang, R.A. Lempicki, M.W. Baseler, et al.; Cutting edge: Ku70 is a novel cytosolic DNA sensor that induces type III rather than type I IFN; J. Immunol., 186 (2011), pp. 4541–4545
- Zheng et al., 2007 L. Zheng, H. Dai, M. Zhou, M. Li, P. Singh, J. Qiu, W. Tsark, Q. Huang, K. Kernstine, X. Zhang, et al.; Fen1 mutations result in autoimmunity, chronic inflammation and cancers; Nat. Med., 13 (2007), pp. 812–819
Document information
Published on 06/04/17
Licence: Other
Share this document
Keywords
claim authorship
Are you one of the authors of this document?