ABSTRACT: (1) Background: Accurately decoding motor imagery (MI) tasks is a prerequisite for creating a MI-based brain-computer interface (BCI). However, low signal-to-noise ratio and non-stationarity of EEG signals present a huge challenge for the classification of MI-EEG signals, restricting the extensive development of the BCI industry. (2) Methods: In this paper, we propose a novel deep learning model CTANet that integrates both channel and temporal attention mechanisms into a convolutional neural network to improve the classification accuracy of the MI-BCI systems. The model is constituted first by three serially connected temporal, spatial, and temporal convolution layers to extract features from the brain signals, with an efficient channel attention module inserted between the second and the third convolutional layers to highlight useful feature channels. Subsequently, the time segment for task decoding is partitioned into several time windows, and a variance layer is employed for computing the logarithmic variance of each window. Next, a multi-head attention mechanism is adopted to extract temporal dependency of features from different windows. Finally, a fully connected layer is used for classifying MI-EEG signals. (3) Results: The performance of the proposed model was evaluated on two publicly available BCI datasets and compared with the state-of-the-art methods. The experimental results show that for dataset BCIC-IV2a, our network achieved classification accuracies of 81.17% and 84.33% in inter-session and intra-session scenarios respectively, whereas for dataset OpenBMI, our network achieved classification accuracy of 73.06% and 77.59% in inter-session and intra-session scenarios respectively. (4) Conclusions: These results outperform state-of-the-art networks, indicating significant potential of the proposed model CTANet in MI decoding.
KEYWORDS: Brain-computer interface; motor imagery; deep learning; convolutional neural network; attention mechanism.
1 Introduction
Brain-Computer Interface (BCI) is a technology that enables direct interaction between the brain and external devices by decoding brain signals. BCI technology represents a potentially powerful option for communication and control in the interaction between the user and the BCI system [1]. Various kinds of neurophysiological patterns have been widely applied in electroencephalogram (EEG)-based BCI systems, such as Steady-State Visual Evoked Potential (SSVEP) [2], Event-Related Potential (ERP) [3], and Motor Imagery (MI) [4].
MI modulates the neural activity in the motor cortex of the brain by imagining the movement of specific body parts, without the need for external stimulation, providing users with a more flexible, convenient, and low-risk way of operation. EEG studies have shown that imagining the movement of different body parts leads to changes in the sensory-motor rhythm of the corresponding motor cortex areas. When the movement of a particular limb is imagined, the sensory-motor rhythm in the related area experiences power reduction, a phenomenon known as event-related desynchronization (ERD) [5]. At the same time, the sensory-motor rhythm power in the contralateral cortical area is enhanced, referred to as event-related synchronization (ERS) [6]. By analyzing the changes in these ERD/ERS patterns, it is possible to effectively distinguish between the imagined movements of different limbs and convert them into control signals for brain-machine interface systems. MI-based BCIs have been manifested as assistive tools for patients with paralysis to promote motor rehabilitation [7] and external equipment control [8]. However, in practical applications, achieving high-precision MI-EEG signal decoding still faces numerous challenges. Firstly, noise artifacts can interfere with signal analysis and processing, reducing data accuracy. Secondly, individual differences among users limit the applicability of universal decoding algorithms. Lastly, insufficient training data hinders the model's effective learning and generalization ability. Accordingly, despite the promising applications of MI-EEG, these challenges make its practical implementation far more difficult than it might appear [9].
Traditional machine learning (ML) methods for processing EEG data typically enhance classification performance by extracting neurophysiological features. First, the raw EEG signals are preprocessed to remove or reduce noise and interference, establishing a foundation for further analysis. Subsequently, algorithms are used to extract discriminative features from the preprocessed data, such as frequency, amplitude, and temporal changes. Classic algorithms such as support vector machines (SVM) [10] and linear discriminant analysis (LDA) [11] demonstrate strong capabilities and are widely used for EEG signal classification tasks. In terms of feature extraction, common spatial pattern (CSP) algorithm [12-13] has been shown to effectively enhance spatial features related to the task, further improving classification accuracy. Filter bank techniques are typically used to capture feature changes at varying frequency rhythms [14]. Continuous wavelet transform (CWT) is used to yield class-discriminative spatio-spectral-temporal representation of EEG signals within subject-specific sub-bands [15]. However, traditional feature extraction methods often rely on manually designed features, making it difficult to capture the complex spatiotemporal dynamics of EEG signals, and they have limited generalization capability.
In recent years, deep learning (DL) methods, particularly convolutional neural network (CNN), have demonstrated exceptional performance across various fields including image recognition, natural language processing, and EEG signal classification [16-18]. The merit of CNN lies in its outstanding ability to automatically learn hierarchical feature representations from raw data, thereby eliminating the need for manual feature extraction. Schirrmeister et al. [19] explored a range of CNN-based network designs that extract task-related information from raw EEG data. Lawhern et al. [20] designed a CNN architecture with a smaller number of parameters using deep and separable convolutions, achieving precise decoding of EEG for different BCI paradigms. Mane et al. [21] employed multi-view data representation and spatial filtering to extract spectral-spatial discriminative features. Saadatmand et al. [22] proposed a two-step multi-objective set-based fuzzy initialization evolutionary algorithm based on integer encoding, which can sequentially identify the optimal channels and the minimum feature set, thus reducing the combinatorial search complexity of MI. While these methods have surpassed classical ML techniques, they have not fully explored EEG features, and their improvements remain limited.
To address the afore-mentioned issues, some researchers have introduced attention mechanisms to enhance DL models, enabling them to focus on the most informative parts of the input data [23]. Attention mechanisms have been successfully applied in fields such as computer vision [24] and natural language processing [25], significantly improving performance. Zhang et al. [26] proposed a DL method that utilizes a squeeze-and-excitation (SE) module to learn the contribution of feature channels to MI tasks, which has evolved into an automatic channel selection strategy. Wang et al. [27] introduced an efficient channel attention (ECA) module within CNN, proposing a novel method for feature channel selection. In addition, Balendra et al. [28] proposed a Modified Vision Transformer model, which enhances the information flow and extracts diverse features by serially and parallelly feeding the initial block through consecutive Transformer blocks. Liu et al. [29] proposed the TCACNet model, which selects a small number of task-relevant time segments through the temporal attention mechanism and adaptively adjusts the attention on different electrode channels using the channel attention mechanism, thus addressing the temporal non-stationarity and uneven information distribution issues in EEG signals. Wu et al. [30] proposed the MSCTANN model, which leverages multi-scale and Channel-Temporal Attention Module (CTAM) to focus on key features, improving the accuracy of MI-EEG classification.
Currently, max pooling or average pooling strategies are the most commonly used techniques for reducing feature dimensions [19-20, 31]. Recently, Mane et al. [21] presented a variance layer that effectively extracts features from multiple time windows of the time series and obtained good performance on two MI datasets. Research has shown that long temporal dependencies between MI-relevant patterns at distinct stages of MI tasks are crucial for improving decoding performance. Ma et al. [32] effectively captured temporal dependencies between discriminative features in different time windows using a temporal attention module.
Although the above methods have achieved good performance, a highly recognized method for modeling the multi-level dependencies in both signal channels and time has not yet been established, making it possible to overlook the complex spatiotemporal structure inherent in EEG signals. Therefore, we propose a novel network for MI classification based on EEG signals, called channel and temporal attention-based DL network (CTANet). This DL framework can automatically select relevant feature channels from EEG signals, extract time-dependent features, and classify MI signals, achieving more accurate and robust task decoding compared to current state-of-the-art methods. The main contributions of this work are as follows:
1) We combine the ECA module, multi-head temporal attention module, and CNN to propose a network that efficiently utilizes both the spatial and temporal features of EEG data. During model training, the ECA module evaluates the relative contribution of each feature channel to the decoding accuracy of MI tasks, dynamically adjusting and assigning suitable channel weights. The multi-head temporal attention assigns weights to sequences within multiple time windows, capturing more advanced temporal features.
2) To effectively extract temporal information, we combined average pooling and variance layers to reduce the temporal dimension of the feature map. This approach not only effectively suppresses noise but also ensures the retention of key information, providing an efficient temporal dimension compression scheme for MI-EEG decoding.
3) We demonstrated that our proposed method outperforms state-of-the-art methods on two publicly available benchmark datasets. We also explored the impact of several key parameters on the model's classification performance.
2 Method
In this section, we first describe the general framework of the proposed network, followed by a detailed explanation of each module in the network. Finally, we outline the training strategy.
2.1 General Framework
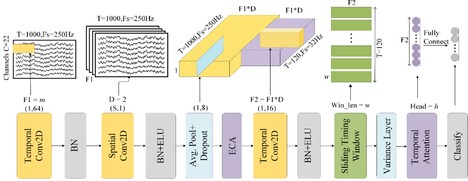
We propose a DL network that incorporates two types of attention mechanisms into a CNN-based network, as shown in Fig. 1. The detailed specifications of the proposed network are listed in Table 1. The network consists of five modules: 1) The convolutional module. It mainly comprises three convolutional layers, i.e., temporal, spatial and temporal convolutions layers, for feature extraction and a mean pooling layer to suppress noise and improve generalization; 2) The channel attention module (CAM). It is inserted before the third convolutional layer to automatically evaluate the importance of different feature channels arising from the first two convolutional layers and assign corresponding channel weights; 3) The deep feature extraction module. It follows the third convolutional layer, whose output is segmented into several time windows, with each time window passing through a variance layer to extract the most relevant temporal information, for reducing temporal dimensionality; 4) The temporal attention module (TAM). It aims to capture and fuse temporal dependencies across different windows; 5) The classification module. It is performed by a fully connected layer and a Softmax operation for transforming the feature signal into a class label. These modules are designed to extract EEG features from both spatial and temporal perspectives, automatically select discriminative feature channels, capture advanced temporal features in the MI-EEG signals, and finally classify the dimensionally reduced feature vectors.
| Layer | #Filters | Size | #Params | Output |
| Input | (1, S, 1000) | |||
| Conv2D | F1=m | (1, 64) | m*64 | (F1, S, 1001) |
| BatchNorm | m*2 | |||
| Deepconv2D | F2=D*m | (S, 1) | D*m*S | (F2, 1, 1001) |
| BatchNorm | F2*2 | |||
| ELU | ||||
| Avgpool2D | (1, 8) | (F2, 1, 125) | ||
| Dropout | ||||
| ECA | k | k | (F2, 1, 125) | |
| Conv2D | F2 | (1, 16) | F2*F2*16 | (F2, 1, 120) |
| BatchNorm | F2*2 | |||
| ELU | ||||
| Reshape | (F2, 120/w, w) | |||
| Variance | (F2, 120/w) | |||
| TAM | h | 120/w | h*(120/w) | (F2, 1) |
| Flatten | ||||
| Fully Connected | F2*N | N |
2.2 Convolution Module
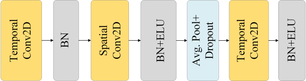
The convolutional module contains three convolutional layers, three batch normalization (BN) layers, two exponential linear unit (ELU) layers, a dropout layer and a pooling layer, as shown in Fig. 2. First, the first convolutional layer applies F1 filters of size (1, K1) for temporal convolution, where K1 is the filter size along the time axis and outputs F1 feature maps. To capture temporal information relevant to frequencies above 4 Hz, K1 is set to a quarter of the sampling rate. A batch normalization (BN) layer then standardizes the output of the first convolutional layer to maintain a stable distribution. Next, the spatial convolutional layer comprises F2 filters of size (S, 1), where S is the number of EEG channels. The number of filter groups is set to D (D=2 in the study), meaning that each feature channel has D filters. This layer outputs F2 feature maps and is then followed by a BN layer and an ELU for normalizing the output of spatial convolution, nonlinear activation respectively. Next, an average pooling layer of size (1, K2) is applied with K2 set to 8, for reducing the sampling rate to ~32 Hz. The average pooling layer is followed by a dropout layer for the reduction of overfitting. Finally, the third convolutional layer with F2 filters of size (1, K3), where K3 is set empirically to 16 to decode MI activity within 500 ms (for 32 Hz sampled data), aiming to further extract temporal features. Batch normalization and ELU activation are used for improving network stability.
In the first convolutional layer, we set the padding parameter to (0, K1/2) to ensure that the time dimension of the EEG signal remains unchanged after the convolutional operation. After the second convolutional layer, a pooling layer with a kernel size of (1, 8) is applied for reducing the length of time series to 1000/8=125. In the third convolutional layer, the padding parameter is set to (0, 5). The convolutional module outputs F2 time series, where the length of the time series is controlled by the three steps mentioned above, ensuring that the input signal with 1000 sampling points is compressed to a length of 120 after passing through the convolutional module.
2.3 Channel Attention Module (CAM)
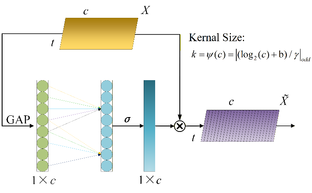
The first two convolution layers in the convolutional module generate numerous features, but their importance for task recognition varies considerably. Thereby, it is necessary to adjust their weights in order to better focus on the key features pertinent to the current task. We chose the ECA to enhance the network's attention to specific feature channels. The ECA module utilizes a one-dimensional (1D) convolution to capture global contextual information and adaptively adjusts weights across all channels. The overall structure of the module is illustrated in Fig. 3.
Fig. 3: The network architecture of ECA. ![]() is the input feature map with the size of
is the input feature map with the size of ![]() , where
, where ![]() and
and ![]() denote the number of feature channels and the number of time points respectively.
denote the number of feature channels and the number of time points respectively. ![]() and
and ![]() are two constants taking values of 2 and 1 respectively.
are two constants taking values of 2 and 1 respectively. ![]() denotes the Sigmoid function.
denotes the Sigmoid function.
Assume that the dimensions of the input feature map X are (N, C, S, T), where N, C, S and T represent the batch size, the number of channels, the size of spatial dimension and the size of temporal dimension respectively. The specific steps of the algorithm are as follows:
1) The global average pooling (GAP) is applied to the input feature map X, compressing it from dimensions (N, C, S, T) to (N, C, 1, 1). This step captures the global information of the entire feature map.
2) The 1D convolution is used to compute channel weights as
|
(1) |
where C1D indicates 1D convolution. The kernel size of the convolution (k) is adaptively determined according to the number of the input feature channels as
where ![]() represents the closest odd integer to t. We empirically set
represents the closest odd integer to t. We empirically set ![]() and b to 2 and 1 respectively. It is then mapped to the range [0, 1] using the Sigmoid function
and b to 2 and 1 respectively. It is then mapped to the range [0, 1] using the Sigmoid function ![]() . This step dynamically adjusts the weights of each channel based on its global information.
. This step dynamically adjusts the weights of each channel based on its global information.
3) Each channel of the input feature map is weighted according to its corresponding weight. This enhances the network’s focus on important features, and thus improves the model’s performance and effectiveness.
Choosing ECA as the CAM avoids dimensionality reduction while utilizing GAP to aggregate convolutional features. It aims to optimize feature extraction and representation capabilities through the capture of global information and dynamic weight adjustment.
2.4 Deep Feature Extraction Module
After the third convolution, the EEG data still contain significant intra-class variance and high noise content along the time dimension. Inspired by the Ref. [21], we use the variance layer to further extract temporal features. Capturing transient changes in EEG signals is crucial for decoding MI tasks. To capture MI-related patterns within these transient changes, we segment the ![]() time series
time series ![]() of length
of length ![]() output from the third convolution layer into N non-overlapping time windows of width w (w is 20 in the study), where
output from the third convolution layer into N non-overlapping time windows of width w (w is 20 in the study), where ![]() . The variance of ith (
. The variance of ith ( ![]() ) time series in nth (
) time series in nth ( ![]() ) window is as follows
) window is as follows
|
(3) |
where ![]() is the average of
is the average of ![]() in nth time window. The variances derived from all N time windows
in nth time window. The variances derived from all N time windows ![]() are then input to the temporal attention module for further feature extraction.
are then input to the temporal attention module for further feature extraction.
2.5 Temporal Attention Module (TAM)
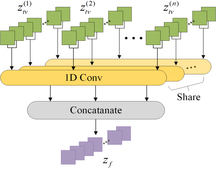
After the variance layer, the feature signals arising from all these time windows are mutually independent. A new temporal attention module was created in order to acquire the temporal dependencies between varying time windows, as depicted in Fig. 4. It employs the concept of multi-head attention, where multiple heads jointly focus on information from different time windows at various positions. 1D convolutions are used as parallel attention heads, with each convolution performed independently on each channel. The kernel size of each convolution is equal to the number of time windows.
Initially, the weights of these convolutions are standardized in the temporal dimension through a Softmax operation. Subsequently, the variance features from the nth window output from the variance layer ![]() are divided into h groups with each group comprising
are divided into h groups with each group comprising ![]() channels, where F2 is the number of feature maps in each window. Next, the features belonging to the same group are fed into the 1D convolutions. In order to achieve multi-head attention, the weights of the 1D convolutions are shared across different groups. Ultimately, the obtained features from all groups are linked together in order to produce the final output signal
channels, where F2 is the number of feature maps in each window. Next, the features belonging to the same group are fed into the 1D convolutions. In order to achieve multi-head attention, the weights of the 1D convolutions are shared across different groups. Ultimately, the obtained features from all groups are linked together in order to produce the final output signal ![]() for the output channel c, which is computed as
for the output channel c, which is computed as
|
(4) |
where ![]() stands for the weight of 1D convolution and ‘mod’ denotes the modulus operation. The output signal
stands for the weight of 1D convolution and ‘mod’ denotes the modulus operation. The output signal ![]() is then employed for the MI decoding.
is then employed for the MI decoding.
2.6 Classification Module
The classification module includes a fully connected layer and a Softmax layer. The feature ![]() output from the temporal attention module is first flattened into a 1D feature vector, which is then input to a fully connected layer to convey into a feature vector with the size equal to the number of classes. Finally, the dimensionally reduced feature vector is transformed into class label using the Softmax function. Regarding model optimization, the network’s loss function is computed by minimizing the cross-entropy loss as follows
output from the temporal attention module is first flattened into a 1D feature vector, which is then input to a fully connected layer to convey into a feature vector with the size equal to the number of classes. Finally, the dimensionally reduced feature vector is transformed into class label using the Softmax function. Regarding model optimization, the network’s loss function is computed by minimizing the cross-entropy loss as follows
|
(5) |
where ![]() represents the number of EEG categories,
represents the number of EEG categories, ![]() and
and ![]() are the ground truth and predicted label respectively.
are the ground truth and predicted label respectively. ![]() demotes the number of trials in a batch.
demotes the number of trials in a batch.
2.7 Model Training
The network is implemented using the PyTorch library in Python 3.7 on a GeForce 3080 GPU. The model training process is divided into two phases. In the first phase, early stopping is enabled, with a maximum training epoch set to 500 and a maximum patience of 50, meaning that if the validation accuracy does not improve for 50 consecutive epochs, the first phase of training will be terminated, and the current network weight parameters will be saved. In the second phase, the weight parameters from the first phase are inherited, and the number of training epochs is set to 200. All weights in the network are initialized using the Glorot initializer, with biases set to zero. The cross-entropy loss is used for network optimization. The Adam optimizer is used with default settings. The initial learning rate is set to 0.001, and if the training loss does not improve for 20 consecutive epochs, the learning rate will be reduced by a factor of 0.6. The dropout rate is set to 0.5.
3 Experimental Setup
In this section, we first describe two datasets used for evaluating our network. We then introduce the baseline methods employed in this experiment. Finally, we provide a detailed explanation of the methods used to evaluate the performance of the network in this experiment.
3.1 Evaluation Datasets
Two publicly available benchmark MI-BCI datasets, BCI Competition IV-2a (BCIC-IV2a) [31] and OpenBMI [33], were used in this study. They differ in the number of subjects, trials, electrode channels and mental tasks. The BCIC-IV2a dataset, as the standard dataset for BCI competitions, has been adopted in over 300 SCI papers and has become the gold standard for reproducible research. On the other hand, the OpenBMI dataset comes from three hospitals in South Korea, where there are differences in the equipment used. Among the 54 subjects, 53.7% of the subjects are BCI illiteracy, and the quality of the EEG signals is relatively poor, which better reflects the signal heterogeneity in real-world applications. The selection of these two datasets helps to comprehensively evaluate the performance and robustness of classification networks.
1) BCIC-IV2a. The dataset contains MI-EEG data from 9 healthy participants, each undergoing 2 sessions. Each session consists of 288 trials, comprising the four MI tasks of the left hand, right hand, foot, and tongue, with 72 trials per task. The EEG signals were recorded using 22 channels with a sampling rate of 250 Hz. In each trial, a fixed cross was displayed on the screen with a brief warning. At 2 seconds, an arrow indicating the direction to be imagined (left, right, up, or down) was displayed for 1.25 seconds in conjunction with the corresponding MI task. The arrow directed the participant to perform the designated MI task until the task reached its conclusion at 6 seconds. Following a brief intermission of 1.5 to 2.5 seconds, the trial was brought to a conclusion.
2) OpenBMI. In the dataset, 54 healthy subjects were instructed to perform two different MI tasks, one was the MI of left hand and the other the MI of right hand. Each subject participated in two sessions, with each session comprising 100 trials. The duration of each trial was four seconds. The MI-EEG signals were acquired with 62 electrodes at a sampling rate of 1000 Hz. According to the methodology established in the original work [33], 20 electrode channels (FC-5/3/1/1/2/4/6, C-5/3/1/z/2/4/6, and CP-5/3/1/z/2/4/6) were selected over the motor cortex area for EEG decoding.
The Braindecode [34] framework, which is supported by PyTorch, was employed for preprocessing the two datasets. For BCIC-IV2a, signals from the eye movement channels were removed, and the data segment from 0 to 4 seconds after the visual cue was extracted for subsequent analysis without applying the 4-40Hz filter. For OpenBMI, due to a large number of subjects and poor EEG quality, we not only extracted the 0-4 seconds data segment following the visual cue, but also applied a 4th-order Butterworth filter with a 4-40 Hz bandpass, with the sampling rate set to 250 Hz. After these preprocessing steps, the dimension of a single trial is (S, 1000), where S is the number of EEG channels.
3.2 Evaluation Baselines
We compared our network with the five baseline methods: one traditional ML method FBCSP14; two classical DL networks Deep ConvNet [19] and EEGNet8.2 [20]; and two state-of-the-art DL networks FBCNet [21] and LightConvNet [32].
1) FBCSP: FBCSP applies the classic CSP algorithm independently on each sub-band of the EEG signals and enhances classification discriminability by integrating multi-band features, reflecting the performance limit of the era of handcrafted features.
2) Deep ConvNet: Deep ConvNet was the first to introduce deep CNNs into MI-EEG analysis, establishing the foundational architecture for spatiotemporal convolutions. By using a hierarchical convolutional structure, it effectively captures event-related desynchronization features in motor imagery tasks. With its outstanding performance and adaptability, Deep ConvNet has become an important tool in the field of computer vision and has achieved significant success in many practical applications.
3) EEGNet: EEGNet is a lightweight model designed for BCI practical applications. It uses depthwise separable convolutions to construct the spatiotemporal filtering module, reducing the parameter scale by two orders of magnitude compared to traditional CNNs, while still achieving good classification performance.
4) FBCNet: FBCNet adopts multi-view data representation and captures discriminative features from multiple frequency bands of EEG signals. The model achieves state-of-the-art performance on several public MI-EEG datasets. In our proposed method, we use the same variance layer as in FBCNet to further reduce the dimensionality in the time domain and extract deep features.
5) LightConvNet: LightConvNet utilizes dynamic convolutions to implement time attention. By using the time attention mechanism, it focuses on the most representative time periods during feature extraction, thereby achieving excellent classification performance.
3.3 Experimental Methods
To evaluate the model's classification performance, we leveraged both inter-session and intra-session experimental scenarios to evaluate the performance of the proposed network. In the inter-session setup, we employed the data from the first session of each subject as training set, with 20% of the training data set aside as the validation set, and the data from the second session as the test set for the decoding experiment. In the intra-session setup, we used ten-fold cross-validation for the decoding experiment, with both the training and testing data drawn only from the first session to avoid the influence of inter-session differences on the decoding performance. We divided the first session into ten parts (maintaining class balance), using one part for testing each time, with the remaining nine parts used for training, and 20% of the training data set aside as the validation set. The final classification result was obtained by averaging the outcomes from the 10 evaluations. In the default configuration of CTANet, we set the number of filters in the first convolutional layer m to 32, the temporal window length w to 20, and the number of temporal attention heads h to 8.
Based on the BCIC-IV2a and OpenBMI datasets, we conducted classification experiments using the proposed model, CTANet. The decoding performance for each evaluation is represented by the average accuracy and F1-score (Accuracy ± Std and F1-score ± Std). First, we compared the proposed method, CTANet, with baseline methods. Paired t-tests at 95% confidence level are used for statistical analysis to compare the difference between two methods. We provided the trainable parameters of CTANet and four DL baseline methods, as well as the time required to complete one inter-session setup experiment, to analyze whether CTANet has practical significance. Additionally, we conducted ablation experiments on the two attention modules in the network to analyze their contributions to the overall performance. Finally, we performed experiments with different hyperparameter combinations to select the best-performing CTANet hyperparameter configuration.
4 Results
In this section, we first provide a performance comparison between the proposed method and five baseline methods, then introduce the results of an ablation experiment and finally describe the parameter sensitivity.
4.1 Comparison with baseline methods
The classification results of our method in both cross-session and within-session scenarios, compared with five baseline methods, are shown in Table 2. The paired t-tests at 95% confidence level are employed for comparing the mean accuracyof our model CTANet and that of each baseline method. Our proposed model achieves the best classification accuracies in each of the two scenarios on the two datasets. Specifically, in the inter-session scenario, our model yields the highest accuracies of 81.17% and 73.06% on the BCIC-IV2a and OpenBMI datasets, respectively. The two classical DL methods, Deep ConvNet and EEGNet, outperform the traditional machine learning method FBCSP, indicating that convolutional neural network (CNN)-based DL methods have stronger feature representation capabilities. Two state-of-the-art DL methods, FBCNet and LightConvNet, surpass the classic methods as they embed filter banks into their networks. Compared to the FBCNet, LightConvNet significantly enhances the accuracy and F1-score (p<0.05) by introducing a temporal attention mechanism, highlighting the important role of TAMin decoding MI-EEG signals. Compared to LightConvNet, our method further improves the accuracy by 1.68% and 2.63% on BCIC-IV-2a and OpenBMI respectively, by jointly exploiting channel and temporal attention mechanisms.
In the intra-session scenario, on the two datasets, Deep ConvNet and EEGNet have lower accuracy than FBCSP on BCIC-IV-2a, while Deep ConvNet also has lower accuracy than FBCSP on OpenBMI. The reason might be that FBCSP fully extracts the feature information in frequency domain by utilizing the filter bank technique, whereas both Deep ConvNet and EEGNet do not. Similar to the inter-session setting, FBCNet and LightConvNet significantly outperform Deep ConvNet and EEGNet in terms of classification accuracy and F1 score on both datasets (p<0.05). Furthermore, by incorporating a time attention mechanism, LightConvNet significantly outperforms FBCNet in both classification accuracy and F1 score (p<0.05). Compared to LightConvNet, our proposed method improves accuracy by 2.01% on BCIC-IV-2a and slightly increases accuracy by 0.07% on OpenBMI. This could be due to the higher quality and fewer categories in the dataset, allowing the excellent DL methods to yield similar decoding performance.
Table 2: Classification accuracies and standard deviations (mean±std, %) of our method and the compared methods. The best numerical values are highlighted in boldface, and * denotes the statistically significant difference between our method and one of the compared methods with p<0.05.
| Data set | Method | Inter-session | Intra-session | ||
| Accuracy | F1-score | Accuracy | F1-score | ||
| BCIC-IV-2a | FBCSP | 66.51±14.04* | 64.51±15.98* | 74.16±13.50* | 73.60±13.80* |
| Deep ConvNet | 70.87±12.90* | 70.59±13.00* | 71.10±11.51* | 70.42±11.36* | |
| EEGNet | 72.76±11.19* | 72.68±11.35* | 73.07±11.54* | 72.68±11.77* | |
| FBCNet | 75.19±13.10* | 74.58±13.66* | 79.25±13.21* | 78.90±13.48* | |
| LightConvNet | 79.48±12.01 | 79.00±12.58 | 82.32±12.72 | 82.03±12.92* | |
| CTANet | 81.17±7.47 | 81.06±7.56 | 84.33±7.13 | 84.18±7.09 | |
| OpenBMI | FBCSP | 59.46±14.73* | 53.57±23.20* | 64.44±17.12* | 63.59±17.77* |
| Deep ConvNet | 59.62±8.99* | 52.35±21.37* | 63.94±11.21* | 58.97±14.77* | |
| EEGNet | 59.72±8.62* | 56.27±17.80* | 66.98±12.29* | 66.54±12.48* | |
| FBCNet | 68.45±13.94* | 67.83±15.36* | 75.77±14.10 | 75.48±14.27 | |
| LightConvNet | 70.43±13.64* | 68.63±18.62* | 77.52±12.92 | 77.21±13.20 | |
| CTANet | 73.06±15.14 | 71.99±16.30 | 77.59±15.01 | 77.30±15.26 | |
Table 3: The number of trainable parameters (Trainable Params) and training time (Train Time) of CTANet and four DL baseline methods.
| Methods | Trainable Params | Train Time (s) (seconds) |
| Deep ConvNet | 282879 | 559.15 |
| EEGNet | 4028 | 351.85 |
| FBCNet | 6564 | 731.51 |
| LightConvNet | 6596 | 904.34 |
| CTANet | 69719 | 553.17 |
Model complexity and training efficiency are of significant guiding importance for the practical deployment of BCI. The number of trainable parameters and the time required to complete a full cross-session experiment on the BCIC-IV-2a dataset for CTANet and four DL baseline models are shown in Table 3. The hardware setting includes an Intel Core i5-12400F CPU and a NVIDIA GeForce RTX 3080 GPU. As shown in the table, EEGNet benefits from designs such as depthwise separable convolutions, achieving the best performance in terms of both trainable parameters and training time. Additionally, Deep ConvNet, EEGNet, and CTANet do not involve multi-band data, so even with significant differences in trainable parameters, the training time remains relatively fast. FBCNet and LightConvNet, although having fewer trainable parameters, require longer training times, possibly due to the additional time needed to process multi-band data. CTANet achieves the best classification performance with relatively little training time (second only to EEGNet), while moderately increasing computational resource consumption, demonstrating its potential in practical BCI applications. Although the training of CTANet takes more than 9 minutes, the testing time for a single trial is minimal (less than 0.2 seconds). Since this model can be trained beforehand, it is fully feasible for real-time online applications.
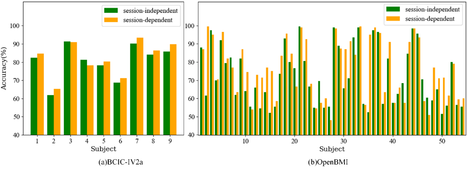
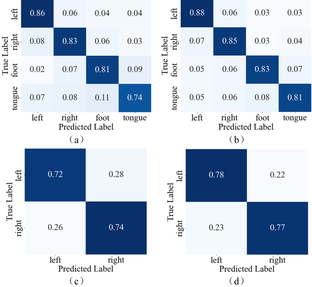
To further explore the effects of inter-session and intra-session, classification results for each subject on the two datasets are shown in Fig. 5. From the figure, it can be seen that accuracies of most subjects in the intra-session condition are higher than those in the inter-session condition, indicating that the features of EEG signals are less variable within the same session but exhibit greater variability across different sessions. Fig. 6 shows the confusion matrices of the CTANet model's classification results on two datasets. As shown in the figure, the average classification accuracy differences for each class between the two conditions are small in both datasets. Therefore, although the model performs better in the intra-session scenario, this scenario involves training and testing data from the same session, which is significantly different from real-world applications. In practical applications, BCI systems need to handle data from different sessions, and the inter-session scenario better reflects the model's generalization ability and robustness. Thereby, although our network is still affected by cross-session variability, it exhibits good decoding performance and strong generalization ability in the inter-session condition.
Fig. 5: Classification accuracy of each subject under inter- and intra-session conditions on (a) dataset BCIC-IV-2a and (b) dataset OpenBMI.
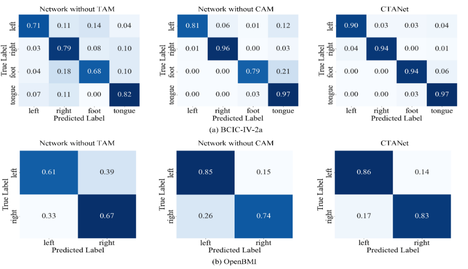
Fig. 6: Confusion matrix of the CTANet model's classification results on two datasets. (a) and (b) represent the confusion matrices of CTANet on the BCIC-IV2a dataset in the inter-session and intra-session scenarios, respectively. (c) and (d) represent the confusion matrices of CTANet on the OpenBMI dataset in the inter-session and intra-session scenarios, respectively.
4.2 Ablation experiment
Compared to the networks composed entirely of CNNs, the key improvement of our network is the incorporation of both CAM and TAM to learn global representations. We conducted an ablation study to validate their effectiveness. The ablation study investigates three sets of methods: 1) without the TAM; 2) without CAM; and 3) CTANet. All experiments were conducted under the same conditions and the results are shown in Table 4. The paired t-tests at 95% confidence level are employed for comparing the mean accury of CTANet and that of the model without TAM or CAM.The results indicate that TAM and CAM contribute to improvement in accuracy, but TAM contributes more than CAM. For BCIC-IV-2a, removing the TAM resulted in an accuracy reduction by 5.36% (p<0.05) for inter-session setting and 5.07% (p<0.01) for intra-session setting. For OpenBMI, removing the TAM led to an accuracy reduction by 2.05% (p<0.05) for inter-session setting and 5.45% (p<0.01) for intra-session setting.
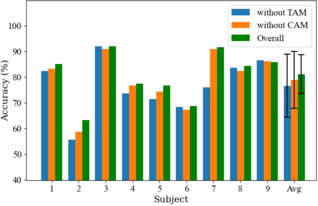
Taking BCIC-IV-2a as an example, the classification results for each subject in the inter-session setting are shown in Fig. 7. It is evident that when the TAM is removed, there is a substantial drop in accuracy for most subjects, with Subject 7 experiencing the greatest decline of 15.63%. When the CAM is removed, there is a relatively small decrease in accuracy for most subjects, indicating that while the CAM helps in extracting more discriminative features, features yielded by TAM contain more relevant classification information for MI decoding.
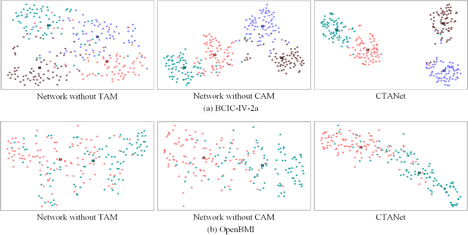
We further analyzed scatter plots of feature signals in the inter-session setting for two representative subjects: Subject 7 from BCIC-IV-2a and Subject 2 from OpenBMI. The 2D scatter plots are achieved by t-SNE [35] from high-dimensional feature signals at the classification layer. Fig. 8 displays the scatter plots obtained from the three sets of methods in the ablation experiment. The figure reveals that for the two datasets, removing either TAM or CAM results in unclear classification boundaries and less distinct class differences and the former behaves more pronounced. This explains that embedding the TAM can significantly enhance decoding performance of MI-BCI systems, whereas embedding the CAM also provides a noticeable improvement in decoding performance. In addition, the confusion matrices for Subject 7 from BCIC-IV-2a and Subject 7 from OpenBMI under the cross-session condition with the three classification methods are shown in Fig. 9. From the figure, it can be seen that the inclusion of CAM and TAM improves the decoding accuracy of CTANet across different class signals, especially TAM, which plays a more significant role in accurately decoding each category of motor imagery signals.
Table 4: Accuracies and standard deviations of the three sets of methods used for the ablation study. The asterisks * denotes the statistically significant difference between our method and one of the compared methods with p<0.05.
| Dataset | Method | Inter-session | Intra-session | ||
| Accuracy | F1-score | Accuracy | F1-score | ||
| BCIC-IV2a | Without TAM | 75.81±12.33* | 75.73±12.15* | 79.26±11.00* | 78.98±11.14* |
| Without CAM | 79.24±11.11 | 79.07±11.30 | 82.08±8.58* | 81.90±8.57* | |
| CTANet | 81.17±7.47 | 81.06±7.56 | 84.33±7.13 | 84.18±7.09 | |
| OpenBMI | Without TAM | 71.01±13.08* | 70.50±13.03* | 72.14±16.06* | 71.68±16.41* |
| Without CAM | 72.35±15.75 | 71.19±17.23 | 76.21±15.61* | 75.84±15.94* | |
| CTANet | 73.06±15.14 | 71.99±16.30 | 77.59±15.01 | 77.30±15.26 | |
Fig. 7: Classification accuracies of each subject yielded by the three sets of methods in the ablation study for inter-session condition on BCIC-IV-2a. Error bars indicate standard deviations.
Fig. 8: The scatter plots yielded by t-SNE from feature signals of the classification layer for the three sets of methods in the inter-session setting. Red, green, purple and brown colors represent MI of left hand, right hand, foot, and tongue respectively. 'x' represents the center point of the class, which is the mean of all scatter points from a class.
4.3 Parameter sensitivity
This section presents a comprehensive assessment of the influence of pivotal parameters on the efficacy of classification. These parameters comprise the number of filters (m), the length of time windows (w), and the number of attention heads (h) in the temporal attention module (TAM).
The number of filters in the first convolutional layer largely determines the total number of trainable parameters in the network. To explore its influence on classification performance and computational efficiency, we explored the classification results of different m values from the set {8, 16, 32, 64}. Table 5 presents classification accuracies for varying m values. It can be observed that the decoding performance of the proposed network exhibits fluctuations with respect to the values of m. The optimal performance is achieved when m is set to 32 when the two datasets are considered all-together.
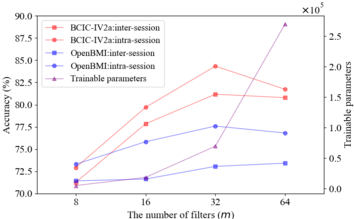
Additionally, we investigated the relationships between both the total number of trainable parameters and decoding accuracy and the number of filters on BCIC-IV-2a. As shown in Fig. 10, the parameter curve indicates that when the number of filters increased from 8 to 32, the number of trainable parameters grows slowly, and subsequently, it rises rapidly. The accuracy curves suggest that the four accuracies derived from the two scenarios and the two datasets scale approximately linearly when the number of filters increases from 8 to 32 and then drop rapidly except for the inter-session scenario on the OpenBMI, where the accuracy continues to rise slowly. It is evident that when the number of filters equals 32, the network achieves optimal classification performance while keeping the number of trainable parameters within a manageable range. Thereby, 32 filters are used in our network because the number of filters offers the best trade-off between the performance and the cost-effectiveness.
| Data set | m | Inter-session | Intra-session |
| BCIC-IV2a | 8 | 71.26±15.60 | 72.93±12.51 |
| 16 | 77.85±10.47 | 79.70±9.11 | |
| 32 | 81.17±7.47 | 84.33±7.13 | |
| 64 | 80.79±10.47 | 81.72±9.23 | |
| OpenBMI | 8 | 71.43±16.77 | 73.30±23.33 |
| 16 | 71.64±15.88 | 75.83±16.34 | |
| 32 | 73.06±15.14 | 77.59±15.01 | |
| 64 | 73.42±14.89 | 76.81±15.61 |
Fig. 10: The impact of the number of filters m on the network's classification accuracy and the number of trainable parameters.
In the subsequent experiments, we investigated the impact of window length (w) on the decoding performance of the model. The window lengths were selected in the set {10, 20, 30, 40, 60}, corresponding to the number of windows being the set {12, 6, 4, 3, 2}. Table 6 presents the accuracies and standard deviations achieved at varying lengths of time windows in the deep feature extraction module. Generally, reducing the window length and increasing the number of windows can enhance classification accuracy, as it helps to increase the amount of data used for model training and enables the model to learn varying MI information from different time points. However, after the improvement in accuracy reaches its peak, further reduction in window length may lead to diminished performance. This is because overly small window lengths partition the signal into narrow segments that may not contain sufficient MI information for effective model training. As shown in the table, when the window length is 10 (with 10 samples per window, corresponding to approximately 80 samples or 0.32 seconds in the original signal), the classification accuracies on the two datasets are lower than those yielded when the window length is 20 (with 20 samples per window, corresponding to approximately 160 samples or 0.64 seconds in the original signal). These results in the table indicate that a window length of 20 with 6 windows yields the best classification performance. This suggests that encoding the original MI-EEG signals with a resolution of 20 samples (640 ms) provides a good representation compared to lower (e.g., 320 ms) or higher (e.g., 960 ms) resolutions.
The number of heads (h) is also an important parameter in the multi-head attention mechanism, which allows features to be extracted from different perspectives. Thereby, we explored the effect of different h values from the set {2, 4, 8, 16} on the decoding performance of the model. The classification results are shown in Table 7. It can be seen that in each setting on each dataset, the performance first increases with the number of heads and then decreases. Specifically, in the inter-session setting on BCIC-IV-2a, the number of heads h has a significant impact on decoding performance, with the best classification performance achieved when h is set to 8. In contrast, on the OpenBMI dataset, the difference between different h values is relatively small in any setting.
Table 6: Accuracies and standard deviations achieved by the proposed model at different lengths of time windows (w) in the deep feature extraction module.
| Data set | w | Inter-session | Intra-session |
| BCIC-IV2a | 10 | 80.05±10.56 | 81.13±10.57 |
| 20 | 81.17±7.47 | 84.33±7.13 | |
| 30 | 78.86±11.41 | 81.17±10.33 | |
| 40 | 76.08±10.36 | 79.90±11.71 | |
| 60 | 75.50±11.44 | 79.91±11.04 | |
| OpenBMI | 10 | 72.31±15.56 | 76.79±15.66 |
| 20 | 73.06±15.14 | 77.59±15.01 | |
| 30 | 72.58±15.70 | 76.44±15.92 | |
| 40 | 72.45±15.69 | 75.90±15.69 | |
| 60 | 71.40±16.14 | 76.06±15.56 |
| Data set | h | Inter-session | Intra-session |
| BCIC-IV2a | 2 | 72.03±14.20 | 82.21±8.85 |
| 4 | 76.70±11.42 | 82.31±8.88 | |
| 8 | 81.17±7.47 | 84.33±7.13 | |
| 16 | 78.40±11.94 | 82.82±7.86 | |
| OpenBMI | 2 | 72.70±15.63 | 76.32±15.10 |
| 4 | 72.56±15.78 | 76.59±15.68 | |
| 8 | 73.06±15.14 | 77.59±15.01 | |
| 16 | 72.03±15.69 | 76.55±15.82 |
5 Discussion
The efficacy of a decoding algorithm directly determines whether the BCI system can be applied in real-world scenarios. We present a classification network CTANet that is both concise and efficient. This proposed model comprises five modules: 1) convolutional module; 2) CAM; 3) deep feature extraction module; 4) TAM; and 5) classification module. Compared to existing DL models consisting entirely of CNNs, CTANet can extract more discriminative features.
CTANet integrates both channel attention mechanism and temporal attention mechanism into the convolutional neural network. The channel attention mechanism enables the extraction of EEG features from a spatiotemporal perspective while automatically selecting relevant feature channels from the EEG signals, allowing the network to focus more on features crucial for the current task. The temporal attention mechanism leverages the multi-head attention to learn global temporal correlations of EEG signals, and thus overcomes the limitation of CNNs that only capture local temporal features. This approach further addresses the issue of long-term temporal dependencies and captures higher-level temporal features.
In this study, CTANet demonstrated superior performance over existing DL methods on both BCIC-IV-2a and OpenBMI. Ablation studies revealed that removing either the CAM or the TAM led to a decrease in accuracy, whereas t-SNE visualization indicated that such removals resulted in less clear classification boundaries and less distinct class differences. Compared to the CAM, the TAM was identified as the more effective component in the proposed network. In addition, intra-session implementations consistently achieved higher decoding accuracy compared to inter-session implementations, indicating that the model effectively utilizes intra-session data for learning and adjustment, and thus enhances decoding accuracy. However, the latter implementations are more practical for real-world applications. Finally, we explored the impact of several key parameters on network performance. Results showed that increasing the number of filters generally improved decoding performance, although this also increased model complexity and the number of trainable parameters. Thus, it is crucial to find a balance to optimize classification performance while avoiding resource wastage. Experiments with different window lengths indicated that while reducing window length and increasing the number of windows could enhance classification accuracy, excessively small windows might segment the signal into narrow subsections that lack sufficient information for training. The model's decoding performance fluctuated with the number of attention heads, with the best decoding performance achieved when that was set to eight.
Recently, attention mechanisms have been frequently employed to construct DL-based classification models, aiming to enhance the classification performance of BCI systems. Ref. [27] proposed an efficient channel attention (ECA) module for deep CNNs, which significantly reducesmodel complexity while bringing clear performance gain. Ref. [26] utilized a sparse squeeze-and-excitation module, namely a channel attention mechanism, for EEG channel selection of MI-BCIs. Ref. [28] proposed a scalogram set-based algorithm and a modifiedvision transformer model, i.e., a temporal attention mechanism, for classificationof EEG data. Ref. [29] presented a temporal and channel attention convolutional network (TCACNet) for MI-EEG classification. The temporal attention mechanism is used to identify the task-related time slices and handle the EEG signal’s temporal non-stationarity, whereas the channel attention mechanism aims toadaptively adjust the weight coefficients of each channel to make the network pay more attention to task-related channels. Ref. [30] integrates a multi-scale CNN module and a channel-temporal attention module (CTAM) for use in MI-based BCIs. The former is able to extract a large number of features, whereas the latter allows the model to focus attention on the most important features. The difference between the first three models and ours is that the former adopts only either channel or temporal attention mechanism, whereas the latter leverages two mechanisms. Although the last two models and ours make use of both channel and temporal attention mechanisms, the difference between them lies in completely different structures.
This study only contains offline analysis using publicly available datasets. The real-time implementation of the proposed CTANet for BCI applications should be considered. Take the hardware setting that includes an Intel Core i5-12400F CPU and an NVIDIA GeForce RTX 3080 GPU as an example. For the intra-session scenario, a user needs to first conduct a 10-minutes offline experiment for each MI task (The MI tasks appear randomly) to acquire enough EEG data, which are used for training the CTANet model for about 9 minutes, and finally the trained model is used for online experiment; For the inter-session scenario, a user can directly use the model trained by previously collected data for online experiment. In either scenario, the single-trial testing time (i.e. the output time of a command) is less than 0.2 seconds and thus CTANet is feasible for practical applications.
This study still has some limitations. First, it only considers spatiotemporal feature extraction in a single frequency band, while many studies have already considered the impact of multiple frequency bands [36-37]. Future research will attempt to combine multi-channel filter banks or Sinc filters to enhance network performance [36]. Second, although our model performs well in cross-session tasks, it has not yet been applied to cross-subject tasks [38]. Future work will attempt cross-subject tasks, starting with leave-one subject-out cross validation experiments on public datasets, and may later require fine-tuning the network parameters for cross-subject tasks. Moreover, while the proposed network has been tested within the MI paradigm, it has not yet been extended to applications such as emotional or SSVEP signal decoding. Future work will consider cross-task transfer learning [39]. Specifically, for emotional EEG decoding, emotional responses last longer and involve a wider range of brain areas, which may require adjusting the input time window and the spatial convolutional layer parameters of the network to capture more temporal features and a broader electrode distribution. For SSVEP decoding, which typically involves signals with higher signal-to-noise ratio and high temporal resolution, different pooling strategies may be required. Finally, CTANet was only evaluated on MI epoch classification without considering idle state discrimination. Future research will focus on real-time scenarios, emphasizing classification speed and performance to enable more natural and fluent BCI usage. Further evaluations will also include datasets with more degrees of freedom and explicit consideration of the idle state, which is of major importance for practical BCI applications. In addition, potential overfitting risk remains due to the relatively high model complexity compared to EEGNet.
6 Conclusion
This paper presents a concise and efficient classification network CTANet, which combines the strengths of CNNs and attention mechanisms. It employs both channel and temporal attention mechanisms to comprehensively capture relationships across all feature channels and analyze global temporal correlations of EEG signals, thereby enhancing the understanding and analysis of EEG data. CTANet also utilizes pooling and variance layers for temporal dimensionality reduction, enabling the extraction of more relevant information. The pooling layer addresses high noise levels, whereas the variance layer excels in capturing data variability. The concurrent use of both layers enhances the network’s applicability and robustness. Experiments conducted on two publicly available BCI datasets demonstrate that this method significantly outperforms the compared methods in decoding performance. Overall, our model provides great potential in advancing MI-EEG decoding.
Acknowledgement: Not applicable.
Funding Statement: This work was supported by the National Natural Science Foundation of China (#62066028).
Author Contributions:
“The authors confirm contribution to the paper as follows: Conceptualization, Shengliang Zou, Jianhua Wu; methodology, Shengliang Zou, Yongge Shi; software, Shengliang Zou, Qingguo Wei; validation, Shengliang Zou, Yongge Shi, Qingguo Wei, Jianhua Wu; formal analysis, Shengliang Zou; writing—original draft preparation, Shengliang Zou; writing—review and editing, Qingguo Wei; funding acquisition, Jianhua Wu. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are available from the Corresponding Author, JW, upon reasonable request.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Wolpaw JR, Birbaumer N, Heetderks WJ, et al. Brain-computer interface technology: a review of the first international meeting. IEEE Trans Rehabil Eng. 2000; 8:164-173.
2. Liu B, Chen X, Shi N, et al. Improving the performance of individually calibrated SSVEP-BCI by task-discriminant component analysis. IEEE Trans. Neural Syst. Rehabil. Eng. 2021; 29:1998-2007.
3. Santamaría-Vázquez E, Martínez-Cagigal V, Pérez-Velasco S, et al. Robust asynchronous control of ERP-based brain-computer interfaces using deep learning. Computer Methods and Programs in Biomedicine,. 2022; 215:Art. no. 106623.
4. Jin J, Sun S, Daly I, et al. A novel classification framework using the graph representations of electroencephalogram for motor imagery-based brain-computer interface. IEEE Trans. Neural Syst. Rehabil. Eng. 2022; 30:20-29.
5. Toro C, Deuschl G, Thatcher R, et al. Event-related desynchronization and movement-related cortical potentials on the ECoG and EEG. Electroencephalography and Clinical Neurophysiology/Evoked Potentials Section. 1994; 93:380-389.
6. Pfurtscheller G, Lopes da Silva , FH. Event-related EEG/MEG synchronization and desynchronization: basic principles. Clin Neurophysiol. 1999; 110:1842-1857.
7. Calderone A, Manuli A, Arcadi FA, et al. The Impact of Visualization on Stroke Rehabilitation in Adults: A Systematic Review of Randomized Controlled Trials on Guided and Motor Imagery. Biomedicines. 2025; 13:599-599.
8. Tang Z, Hong X, Lv S , et al. Hand rehabilitation exoskeleton system based on EEG spatiotemporal characteristics. Expert Systems with Applications. 2025; 270:126574-126574.
9. Lotte F, Bougrain L, Cichocki A, et al. A review of classification algorithms for EEG-based brain–computer interfaces: a 10-year update. J. Neural Eng. 2018; 15:Art. no. 031005.
10. Khalighi S, Sousa T, Oliveira D, et al. Efficient feature selection for sleep staging based on maximal overlap discrete wavelet transform and SVM. 2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Boston, MA, USA, 2011, pp. 3306-3309.
11. Kanagaluru V and Sasikala M. Two Class Motor Imagery EEG Signal Classification for BCI Using LDA and SVM. Traitement du Signal. 2024; 41:2743-2749.
12. Müller-Gerking J, Pfurtscheller G and Flyvbjerg H. Designing optimal spatial filters for single-trial EEG classification in a movement task. Clin. Neurophysiol. 1999; 110:787-798.
13. Ramoser H, Müller-Gerking J; Pfurtscheller G. Optimal spatial filtering of single trial EEG during imagined hand movement. IEEE Trans. Rehabil. Eng. 2000; 8:441-446.
14. Ang KK, Chin ZY, Zhang H, et al. Filter Bank Common Spatial Pattern (FBCSP) in brain-computer interface. 2008 IEEE International Joint Conference on Neural Networks (IEEE World Congress on Computational Intelligence), Hong Kong, China, 2008, pp. 2390-2397.
15. Kim J, Heo K, Shin D, et al. A learnable continuous wavelet-based multi-branch attentive convolutional neural network for spatio-spectral-temporal EEG signal decoding. Expert Systems with Applications. 2024; 251:Art. no. 123975.
16. Wang D, Yao J, Zhang Y. Human activities recognition from video images by using convolutional neural network. Journal of Intelligent & Fuzzy Systems. 2025; 48:931-942.
17. Malik S and Jain S. Deep Convolutional Neural Network for Knowledge-Infused Text Classification. New Generation Computing. 2024; 42:157-176.
18. Jirayucharoensak S, Pan-Ngum S, Israsena P. EEG-Based Emotion Recognition Using Deep Learning Network with Principal Component Based Covariate Shift Adaptation. The Scientific World Journal. 2014; 2014:Art. no. 627892. doi: 10.1155/2014/627892.
19. Schirrmeister R, Gemein L, Eggensperger K, et al. Deep learning with convolutional neural networks for EEG decoding and visualization. Hum. Brain Mapp. 2017; 38:5391-5420.
20. Lawhern VJ, Solon AJ, Waytowich NR, et al. EEGNet: A compact convolutional neural network for EEG-based brain-computer interfaces. J. Neural Eng. 2018; 15:Art. no. 056013.
21. Mane R, Chew E, Chua K, et al. FBCNet: A multi-view convolutional neural network for brain-computer interface. 2021, arXiv:2104.01233.
22. Saadatmand H and Akbrazadeh-T M-R. Multiobjective Evolutionary Sequential Channel/ Feature Selection for EEG Motor Imagery Analysis. IEEE Journal of Biomedical and Health Informatics. 2025; 29: 2546-2556.
23. Vaswani A, Shazeer N, Parmar N, et al. Attention is all you need. 31st International Conference on Neural Information Processing Systems (NIPS'17), Long Beach, CA, USA, 2017, pp. 1-11.
24. Woo S, Park J, Lee JY, et al. Cbam: Convolutional block attention module. 15th European Conference on Computer Vision (ECCV), Munich, Germany, 2018, vol.11211, pp. 3-19.
25. Zhao F, Quan B, Yang J, et al. Document Summarization using Word and Part-of-speech based on Attention Mechanism. Journal of Physics: Conference Series. 2019; 1168:Art. no. 032008.
26. Zhang H, Zhao X, Wu Z, et al. Motor imagery recognition with automatic EEG channel selection and deep learning. Journal of Neural Engineering. 2021; 18:Art. no. 016004.
27. Wang Q, Wu B, Zhu P, et al. ECA-Net: Efficient Channel Attention for Deep Convolutional Neural Networks. 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Seattle,USA, 2020, pp. 11531-11539.
28. Balendra, Negi PCBS, Sharma N, et al. Scalogram sets based motor imagery EEG classification using modified vision transformer: A comparative study on scalogram sets. Biomedical Signal Processing and Control. 2025; 104:Art. no. 107640.
29. Liu X, Shi R, Hui Q, et al. TCACNet: Temporal and channel attention convolutional network for motor imagery classification of EEG-based BCI. Information Processing & Management. 2022; 59:Art. no. 103001.
30. Wu R, Jin J, Daly I, et al. Classification of motor imagery based on multi-scale feature extraction and the channel-temporal attention module. IEEE Trans. Neural Syst. Rehabil. Eng. 2023; 31:3075-3085.
31. Tangermann M, Müller KR, Aertsen A, et al. Review of the BCI competition IV. Frontiers Neurosci. 6:Art. no. 55.
32. Ma X, Chen W, Pei Z, et al. A Temporal Dependency Learning CNN With Attention Mechanism for MI-EEG Decoding. IEEE Trans Neural Syst Rehabil. Eng. 2023; 31:3188-3200.
33. Lee M-H, Kwon O-Y, Kim Y-J, et al. EEG dataset and OpenBMI toolbox for three BCI paradigms: An investigation into BCI illiteracy. GigaScience. 2019; 8:Art. no. giz002.
34. https://github.com/braindecode/braindecode.
35. van der Maaten L and Hinton G. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008; 9:2579-2605.
36. Liu K, Yang M, Xing X, et.al. SincMSNet: a Sinc filter convolutional neural network for EEG motor imagery classification. J Neural Eng. 2023; 20:Art. no. 056024.
37. Fan Z, Xi X, Gao Y, et al. Joint Filter-Band-Combination and Multi-View CNN for Electroencephalogram Decoding. IEEE Trans Neural Syst. Rehabil. Eng. 2023; 31:2101-2110, 2023.
38. Sartipi S and Cetin M. Subject-Independent Deep Architecture for EEG-Based Motor Imagery Classification. IEEE Trans Neural Syst. Rehabil. Eng. 2024; 32:718-727, 2024.
39. Liang Z, Zheng Z, Chen W, et al. A novel deep transfer learning framework integrating general and domain-specific features for EEG-based brain–computer interface. Biomedical Signal Processing and Control. 2024; 95: part. B, Art. no. 10631.
Document information
Published on 05/02/26
Licence: CC BY-NC-SA license
Share this document
Keywords
claim authorship
Are you one of the authors of this document?