(Created page with " <!-- metadata commented in wiki content <div class="center" style="width: auto; margin-left: auto; margin-right: auto;"> <big>'''Proposing A Novel Deep Learning And Cluster...") |
m (Osman.alhussein moved page Draft Alhussein 343141266 to Alqazzaz et al 2026a) |
(No difference)
| |
Latest revision as of 22:17, 20 February 2026
Abstract:
Autism Spectrum Disorder (ASD) represents a complicated neurodevelopmental condition where early diagnosis is imperative for effective intervention. Behavior assessments, which are somewhat the norm, primarily slow down diagnosis. This thesis presents a hybrid deep learning approach using brain MRI data for the early detection of autism. This approach combines FCM and GMM clustering MRI data, the multi-channel deep learning ResNet architecture for image classification, and the Dwarf Mongoose Optimization Algorithm for feature enhancement, model refinement, and improving generalization. The proposed model achieved an accuracy of 93.72%, precision of 94.81%, recall of 93.14%, and an F1-score of 93.97% when evaluated on the ABIDE I dataset. These results demonstrate and give evidence of the model overcoming challenges related to previously insufficient models in overfitting and predictive performance. This research is geared towards helping automate feature extraction on highly dimensional MRI data having a great scope for ASD clinical diagnosis. This can benefit ASD clinical diagnosis and serve as a starting point for research in advanced ASD clinical diagnosis.
Keywords: Autism Spectrum Disorder, ResNet, FCM, MRI Image, Dwarf Mongoose Optimization.
Introduction
Among all neurodevelopmental disorders, Autism Spectrum Disorder (ASD) is one of the most complicated and involves the most intricate variations of communication, socialization, and behavior. It generally develops during the formative years, and early identification can meaningfully improve the quality of life of those affected. Unfortunately, identifying the disorder is a complicated endeavor, primarily due to the inaccuracies and insufficiency of objectivity of the traditional behavioral and psychological assessments [1].
Thanks to advancements in medical imaging tools, especially Magnetic Resonance Imaging (MRI) technology, it is possible to assess the functions and structures of the brain in detail. Advanced imaging of brain scans can be especially useful to understand those neuroanatomical differences that contribute to the diagnosis of pediatric developmental disorders. Because of the profound complexity, subtle differences in features, high dimensionality involved while evaluating thousands of images for a particular disorder and being unique to each brain makes formulation of an accurate assessment to be a challenging task [2].
Discovering new, intelligent technologies for the analysis of medical data has been a focus of research in recent years. Such methods provide patterns of brain data in higher accuracy, thus the flat pattern shape emphasized here can be detected at an earlier stage of development landmark of the BRAIN diseases. This is especially the case with autism spectrum disorder (ASD), since accurate and early diagnosis is so critical, owing to the potential for early and efficacious remediation that enormously improves cognitive and social functioning in afflicted children.
Problem Statement
Autism Spectrum Disorder (ASD) is a condition that affects how people interact, communicate, and behave. Getting help early can really make their lives better [3]. However, diagnosing the disorder is problematic as the conventional methods depend exclusively on the behavioral and psychological perspectives.
These approaches probably lack evaluative objectivity and precision, which results in various challenges and, subsequently, delays and errors in diagnosis [4]. Developing and understanding the structure and functions of the brain have always been greatly assisted by Magnetic Resonance Imaging (MRI) which has recently become widely popular. Brain anatomy is highly complex, and the degree of spatial resolution is often overshadowed by subtle structural variances, high dimensional data, and the intricate and unique nature of every individual’s brain patterns. This is, however, the right approach to interpreting and analyzing MRI data and calls for intelligent automated routines [5-6].
AI and deep learning algorithms perform image analysis in medicine and are rapidly changing the domain. With this in mind, one may combine enhanced nature-inspired algorithms like the Dwarf Mongoose Optimization (DMO) with convolutional deep neural networks like VGG-16 and ResNet to achieve more Asd precise fuzzy clustering with Fuzzy C-Means (FCM) and Gaussian Mixture Model (GMM) algorithm. This research primarily aims to implement a smart multifunctional framework to integrate the processes of segmentation, feature extraction, and classification of brain MRI images, intending to streamline the diagnostic process for autism and work towards automation. Such a framework attempts to improve the diagnostic methods that are currently used, aiming to improve the accuracy and reduce the reliance on subjective human judgment in the diagnostic process.
Significance and Necessity of the Study
Autism Spectrum Disorder (ASD) is a complex disorder and a serious condition. Its lifetime consequences can be mitigated through early detection [7]. Early intervention at critical developmental windows can reduce challenges in communication, behaviors, and social interactions. Unfortunately, conventional diagnostic methods that rely on behavioral and psychological assessments remain subjective and imprecise. Diagnosing ASD at an early stage remains difficult. Moreover, some fields of medicine have progressed, and more specifically, magnetic resonance imaging (MRI) has made it possible to obtain detailed anatomical pictures of a person’s brain.
High resolution does not guarantee diagnostic accuracy, however, because of the great variability between individuals and the nuanced and complex features of an autistic brain. Therefore, the use of integration of complex adaptive systems, artificial intelligence, machine learning and deep learning techniques that analyze the intersection of medical imaging and ASD is essential. MRI data requires copious amounts of intricate information, thus manual methods for image processing are becoming impractical. The combined Fuzzy C-Means (FCM) clustering with Gaussian Mixture Models GMM, VGG-16 Convolutional Neural Networks for feature extraction, and ResNet with Dwarf Mongoose Optimization heuristic proposed by us in this research provides steps toward more reliable and precise diagnosing of autism [8].
This integrated model attempts to increase efficiency and accuracy while ensuring it is practical for clinical use. In this respect, the goal of the research is to create an artificial intelligence-based system capable of sophisticated analysis of brain MRI scans. It overcomes the shortcomings associated with conventional diagnostic techniques, lowers cost and time of the diagnosis, and enhances healthcare processes for patients with autism [9].
Goals
Main Objective
To formulate and demonstrate a new approach for diagnosing autism from brain MRI scans using fuzzy clustering and Gaussian mixture models with feature extraction via deep neural network VGG-16 and final classification with ResNet.
Sub-Objectives
• Employing the VGG-16 neural network, investigate and extract relevant features from brain MRI images pertinent for diagnosing autism.
• Appraise the efficiency and precision with which fuzzy clustering and Gaussian mixture models devise categories within autism-related MRI data.
• To improve the precision and expediency with which autism is diagnosed, design and implement a hybrid model for the clustering and classification of MRI images.
• Measure accuracy, sensitivity, and overall operational efficiency in diagnosing autism using MRI images with the proposed technique relative to established methods to assess the value of the new approach.
• Explore and study particular brain regions containing structural features related to autism to help deepen the understanding of structural differences in the brains of autistic individuals.
Research questions
1. Segmentation-Related Research Questions:
To what degree does the hybrid approach of Fuzzy C-Means (FCM) and Gaussian Mixture Model (GMM) increase the accuracy and robustness of brain region segmentation in MRI scans?
What effect does the use of spatial and intensity information in the segmentation process have on the quality of the segmented brain regions?
2. Feature Extraction Research Questions:
How well can the VGG-16 Convolutional Neural Network (CNN) extract pertinent and distinctive features from the segmented brain regions regarding autism diagnosis?
What is the contribution of different layers of the VGG-16 network to the extraction of features pertaining to the abnormalities specific to autism?
3. Classification Research Questions:
How does the integration of ResNet with the Dwarf Mongoose Optimization Algorithm affect classification performance for autism diagnosis?
How does the presence of residual connections in ResNet impact its learning of distinguishing features for discerning autistic and typically developing individuals?
Research Hypotheses
1. Segmentation-Related Hypotheses
- H1: When it comes to segmenting brain regions on MRIs, hybrid Fuzzy C-Means (FCM) and Gaussian Mixture Model (GMM) surpass the standard methods in both accuracy and dependability.
- H2: The integration of segmentation dimensions of space and intensity results in improved brain region segmentation relative to autism.
2. Feature Extraction-Related Hypotheses
- H3: The VGG-16 Convolutional Neural Network performs feature extraction on autism-related brain region segments with exceptional efficiency.
- H4: Different layers of the VGG-16 net work add various structural abnormalities related to autism to the feature set, enhancing the net work’s discriminative power.
3. Classification-Related Hypotheses
- H5: Integration of ResNet and optimization techniques like the Dwarf Mongoose Optimization, increases classification accuracy between autistic and typically developing individuals.
- H6: The residual connections in ResNet enhance the learning of features within the MRI data which, in turn, increases diagnostic accuracy.
Innovation Aspect
This study outlines a new system to diagnose autism spectrum disorder (ASD) from MRI data by combining clustering and deep learning convolutional neural networks with other optimization strategies. The proposed research aims to accomplish this objective through various strategies.
1. Hybrid Feature Extraction Approach: The model captures complex patterns in MRI brain scans through Gaussian and fuzzy clustering techniques to reveal the brain structural differences underlying autism.
2. Use of Pre-Trained Deep CNNs: The pre-trained deep convolutional networks VGG16 and ResNet50 are used in a feature extraction role to automate classification of the datasets. VGG16 applies a deep network architecture to enhance representation of data at higher levels during training and provides quick classification to exceed the classification speed and accuracy of ResNet50 while overcoming the vanishing gradient problem.
3. Parameter Optimization Using the Dwarf Mongoose Optimization Algorithm (DMOA): Using DMOA as a nature-inspired meta-heuristic provides efficient hyperparameter tuning and optimization while minimizing the allocated computational resources. This approach maximizes model improvement and computational resource improvement relative to convergence. This increases computational resource speed during processing.
Research background
In the field of detection of autism spectrum disorder from MRI images and artificial intelligence and deep learning methods, many researchers have conducted huge work, and various research papers have been published. A majority of the research has been centered on enhancing the precision and efficiency of diagnosis using various image processing methods, cluster techniques, convolutional neural network, and algorithmic optimization methods. In this section, in order to give an overview of the outcomes and techniques employed in this field, a number of these studies and their findings are discussed with emphasis on the background and the importance of the current research
The heterogeneity of Autism spectrum disorder (ASD) effects on brain connectivity brings specific issues to machine learning diagnostic models and its detection with the help of MRI in particular. The neural network research that has been carried out is seemingly oriented on the study of brain connectomics, such as functional and structural connectomics of sparse MRI and fMRI that tend to ignore morphological relational variations among the brain areas. This is why there are few machine learning studies that use morphometric analysis of the brain networks to diagnose ASD and T1-weighted MRI. To solve this problem, Kaggle competition was launched to solve the problem of the creation of machine learning pipelines to diagnose neurodevelopmental diseases with a particular focus on ASD based on cortical morphometric networks of T1-weighted MRI. In addition to the competition, the users received a training dataset and their results were only tested on a public test dataset. The last evaluation was conducted on both the public and hidden test datasets in which the evaluations were done on the basis of accuracy, sensitivity, and specificity and the individual values were computed. Ranking of participants was on a one on one basis on each measure and on the aggregate ranking, it was determined by aggregating mean rankings of participants. The team with the highest rank reported accuracy of 70 percent and sensitivity and specificity of 72.5 percent and 67.5 percent respectively. The same measures were reported to be 63.8, 62.5, and 65 percent among the second team, 63.8, 62.5 and 65 percent among the third team reporting respectively [10].
ASDs are a variety of developmental disorders that affect the socio-interaction, communication and imagination use, and interaction with others. The number of ASDs cases that have been diagnosed in Malaysia appears to grow year by year. The diagnosis of ASD in the vast majority of cases is conducted on the basis of clinical examinations conducted by the doctor, including a detailed case history and analysis of behavior. Study [11] seeks to determine the biomarkers of ASD using the functional MRI neuroimaging method to facilitate the diagnosis of such patients. The Deep Learning methods used to distinguish between ASD and non-ASD patients include, in particular, some Convolutional Neural Networks (CNNs) that are quite effective at extracting features. The Overfitting and Underfitting measures used in the evaluation of the models are based on the ability of the models to classify the neural patterns using the preprocessed images. The brain image of autism patients is examined using ABIDE data. The CNN models, i. e. VGG-16 and ResNet-50, described the accuracies of 63.4 and 87.0, respectively. This approach is more precise in terms of identification of ASD that helps clinicians to measure the diagnosis with the help of diagnosed tests and not only basing on observable behavior analysis.
Autism spectrum disorders (ASD) are caused by a particular neurological healthy makeup of the individual that usually is manifested in communication difficulties, social interaction, and repetitive actions. There are rough estimates that ASD is present in one per 100 individuals in the world, although the disorder manifests differently in different individuals. Although there are patients who can live completely on their own, other ones need lifetime care and assistance. ASD may pose a severe barrier to academic and professional potential, and it puts a burden on the family that cares. ASD is normally diagnosed in the first stages of adolescence though its symptoms may be detected early in life. The article refers to reference [12] in the application of diagnostic imaging techniques, particularly MRI scans in terms of autism spectrum disorders. Different machine learning techniques have been conducted according to the comparison of MRIs of the children with autism and their normal counterparts, applying random forests, support vector machines, and convergent neural networks. Using a confusion matrix, random forest bias demonstrated the best accuracy of 100 percent therefore, expressed the best identification of ASD using MRI scans.
The study [13] was dedicated to the creation and verification of deep learning models that are used to diagnosis Autism Spectrum Disorder (ASD) through MRI and Apparent Diffusion Coefficient (ADC) images. There were 151 ASD children and 151 neurotypical controls of the same age (TD) with their information divided into training and validation sets. Also, there was another test group (ASD 20 children and TD 25 controls) where cMRI and diffusion-weighted imaging were conducted. DL models have been constructed to differentiate between ASD and TD based on cMRI and DWI data. There were five single-sequence models, one dominat-sequence, and one all-sequence model (ASM). To the models, the Attention Mechanisms were introduced in order to fine-tune feature detection. The greatest AUC of 0.824- 0.850 using SSM was obtained with both FLAIR and ADC in the validation and independent test sets. In the validation (0.873) and independent (0.876) sets, DSM performed better in AUC with unrestricted FLAIR and ADC(0.873). ASM also demonstrated a superior diagnostic value to SSMs in the validation (AUC = 0.838) and independent (AUC = 0.836) sets. The best attention mechanism model in terms of diagnostic performance with DSM, AUC, accuracy, sensitivity and specificity at of 0.898, 84.4%, 85.0% and 84.0% respectively. Consequently, this study emphasized the applicability of deep learning algorithms in the detection of the cases of autism spectrum disorder in the study of the normally developing participants based on cMRI and ADC images.
In [7] a CAD (computer-aided diagnosis) is created as a classification system of individuals with Autism Spectrum Disorder (ASD) that focuses on the detection of morphological anomalies of certain brain areas. The features are considered by their ability to distinguish an ASD or a typically developing (TD) individual using an ML model on the cortex. This approach has a prospect of improving CAD systems which are capable of diagnosing ASD early and customized. The suggested framework will be created of a total of five large parts: (i) to extract the cortical surfaces in structural MRI; (ii) to conduct cortical parcellation according to a standard atlas; (iii) to determine cortical features as Autism markers; (iv) to bias feature values on gender and age; and (v) to create neural atlases in Autism. ASD is then classified using artificial neural networks. Data in ABIDE I site is used to test the accuracy of the system and a mean balanced accuracy of 97 2 is obtained.
Because of the profound differences among individuals, the cognitive ability cannot be easily quantified as a component of the clinical diagnosis of Autism Spectrum Disorder (ASD). Unless therapies are used, early and precise ASD diagnosis facilitates easier development of effective management strategies. Study [14] proposes an ASD diagnosis procedure using sMRI via an outlier detection approach. A GAN is trained on only sMRI scans of healthy individuals to comprehend the complex relationships that constitute brain structural connectivity. Three neighboring slices are produced by the GAN generator from a batch of sMRI scans. The CNN classifier takes the output slices and passes them to determine whether the input image pertains to an ASD subject or not. With these methods, accuracy in patient ASD classification was 86%.
The dataset prepared by [15] contains two levels of ASD types (mild and severe) and rich features of age and gender compatible with machine learning for ASD diagnosis from MRI image scans. The primary focus of this research is to assess the impact of the disorder at various staging levels, exploring possible patterns associated with ASD on a global level. Moreover, sets of MRI and non-MRI features were formulated to improve the precision of computer-aided diagnostic systems. The findings indicate that machine learning techniques are effective in recognizing and classifying variances of ASD. The deep learning model proposed in [16] is named ASD-DiagNet. It demonstrates high accuracy reliably distinguishing the brain scans of patients with autism spectrum disorder (ASD) from those of neurotypical individuals. The combination of classical machine learning and deep learning methods allows for the extraction of ASD biomarkers from MRI datasets. This system, referred to as Auto-ASD-Network, utilizes deep learning and support vector machines for the classification of ASD scans and normal scans. These models aid in interpreting results generated by deep learning algorithms, thus enhancing explanatory and knowledge discovery processes for neuroscientists. Active diagnostic techniques focus on behavioral evaluations which are lengthy and subjective to the observer’s bias.
The authors in the article [17] outline a method of diagnosing autism using structural MRI brain images of the ABIDE-2 data and using well-trained machine learning algorithms. At 0.82 and 0.80 respectively, the accuracy of prediction used to quantify the diagnosis of autism spectrum disorder in the past was low in comparison to the prediction using supervised learning methods, random forests, and logistic regression. In addition, the VBM analysis of the identical dataset was conducted with the help of SPM8 to outline the structural variations in the brains of the autistic individuals and their neurotypical counterparts. This was analyzed on a control subject that produced the image of a brain using a glass brain.
The basis of ASD diagnosis is clinical judgment and is exceptionally demanding due to the absence of specific quantitative markers. Therefore, automatic classification of individuals on the ASD spectrum faces many challenges and leads to controversial results. Machine learning approaches are being embraced more and more as tools for diagnosis, with the possibility of increasing accuracy for distinguishing ASD from TD and revealing factors distinguishing ASD and providing insightful factors on the distinctions between autism and typically developing individuals. In study [18], numerous feature selection and classification methods were examined to distinguish ASD from TD employing data from a vast resting-state functional MRI (rs-fMRI) imaging database. Static functional connectivity and demographic data were combined with new features such as Haralick texture features and Kullback-Leibler divergence in attempts to identify the most global distinguishing features. This framework successfully identified a small amount of discriminative features and achieved greater accuracy than prior studies; the optimal classification accuracy determined was 0.725.
ASD is acknowledged as the fourth most prevalent neurodevelopmental disorder, with a prevalence of 1.4% in children. Diagnosis is pinpoint accurate, and depends heavily on the available specialists, which poses a problem within itself. That shortage has sparked interest in developing machine learning models that automatically integrate features from functional and structural MRIs (fMRI and sMRI) to detect central nervous system abnormalities associated with ASD. Study (Mellema et al., 2022) performed a hyperparameter tuning analysis on the twelve most popular and powerful machine learning algorithms. All models underwent training in a round-robin fashion with 15 feature sets consisting of fMRI and sMRI features and were optimized using model agnostic optimization. Diagnostic accuracy was higher in the deep learning models and those models generalized well to other MRIs. Our model achieved advanced diagnostic accuracy with an AUROC of 80% on the test data from the IMPAC dataset and 86%, 79%, and then 93%, 90% after supervised adaptation on ABIDE I and II datasets externally. The best models ascertained potential biomarkers for exact ASD diagnosis, confirming known and novel cerebellar biomarkers. This reliability along with the ability to claim the models could construct and derive a robust set of ASD biomarkers to be used raises the level of confidence in the understanding of the neural changes in autism.
ASD is distinguished as having difficulties with social behavior. The underlying mechanisms of ASD have been addressed by researchers in cognitive science, but the condition's biomarkers remain elusive. ASD may be diagnosed with the help of the anatomical structure of the corpus callosum and the intracranial brain volume, which are studied in relation to the ASD condition. Informed by these claims, study [19] seeks to develop a framework for automatic diagnosis of ASD using machine learning based on features extracted from the corpus callosum and the volume of the brain's intracranial cavity. The framework not only provides high accuracy of identification but additionally enables ease in the modeling of machine learning due to its feature selection for classification specific to ASD. Furthermore, in this study, the aim is to assess the capabilities of deep learning with neuroimaging data, thus a “transfer learning” technique is applied. For the purpose of classifying ASD, a VGG16 model that has already been trained on other tasks is used.
Methodology
In recent years, brain imaging techniques, particularly Magnetic Resonance Imaging (MRI), have developed immensely and have been held accountable for diagnosing and examining disorders such as autism. Autism, which is a neurologic disorder of complexity, should be diagnosed in time and accurately so that those afflicted with it are able to improve the quality of life and interventions should be effectively implemented. However, conventional MRI data analysis and processing methods are faced with issues such as low classification accuracy, poor feature extraction, and high complexity.
The conventional methods have mostly used the conventional image processing and modeling methods, which are usually incapable of deriving the natural and multifaceted patterns and features of brain images. Besides, the fact that cutting-edge deep learning models and optimization methods have never been fully implemented has made it so that such methods are less effective in producing high diagnostic accuracy. All these issues necessitate the establishment and formulation of a new and effective system of analyzing MRI data in the diagnosis of autism.
This study will present a new method in diagnosing autism in MRI images through new techniques of clustering, deep convolution neural networks, and advanced optimization algorithms. The novelty of this study lies in the extraction of features with the use of Gaussian models combined with fuzzy clustering, and the use of more sophisticated networks VGG16 and ResNet50 networks that can identify more complex patterns in the brain. Also, the aspect of using the Dwarf Mongoose Optimization Algorithm (DMOA) to optimize hyperparameters is not only good to improve the performance of the model, but also reduces the cost of computations. One of the greatest attainments has been the fact that the algorithms and networks make use of huge MRI data to demystify complex and obscured brain structures.
VGG16 extracted features, which provided refinement in the aid of representation optimization concerning input, and ResNet50’s sophisticated architecture to classified documents concerning speed and accuracy eliminates not only the problem concerning the vanishing gradient, but also aims to optimize speed and accuracy in classification. All to construct an advanced autism detection system from MRI images.
Stages of the Proposed Method
The purpose of the proposed study is to develop a hybrid model hybridising a convolutional neural network with the DM-ResNet classifier to increase the accuracy of brain MRI scan-based diagnosis of autism. The brain MRIs are first standardized and optimized to improve the quality of images. The final section of this step is to undergo the cortical area segmentation of the Fuzzy C-Means (FCM) and Gaussian Mixture Model (GMM) clustering algorithms. The two approaches are complementary measures to determine well the areas of the brain that are related to autism. The raw images are fused with the processed images to produce multichannel images which form the major input in identifying autism. Image feature extraction is the next step and is performed with the help of VGG-16 network which is a pre-trained network on ImageNet. The characteristics are then forwarded to the ResNet network which is firstly trained by employing the Dwarf mongoose Optimization Algorithm (DMOA) so that the hyperparameters may be modified to the best possible values, as established by DMOA. The polished ResNet is subsequently trained to identify the patterns of autism and tested with regard to its availability and robustness. DMOA has been implemented in this thesis to maximize the learning rate and batch size.
Figure 1 illustrates the entire process flow from preprocessing to final classification, which includes a diagram of the proposed method. Incorporating additional innovative machine learning methods strengthens the accuracy and dependability of image-based autism detection. In subsequent sections of this thesis, I will explain every step in the block diagram in detail.
Figure 1: Diagram of the proposed method
Figure 1: Diagram of the proposed method
Preprocessing of Brain MRI Images
In our proposed approach, we prioritize the preprocessing of brain MRI images, focusing on improving consistency and uniformity, while also enhancing the quality of the image. This stage starts with normalization, where all pixel intensities of images are adjusted through mapping so that their values are between 0 and 1. This process is facilitated through the use of statistical formulas which minimizes variability emanating from different imaging conditions while improving uniformity of the data.
Following normalization, noise and artifacts that may hamper correct analysis are removed. Sophisticated spatial filters are used for the task. Gaussian filters are used to reduce smooth noise, while median filters are used in removing salt-and-pepper noise. The filters come in handy in preserving edges and key details in the images.
Then non-brain tissues such as the skull and skin that contain redundant information for processing are removed. They do it using algorithms such as the Brain Extraction Tool (BET), which relies on intensity modeling and surface detection to separate brain tissues from other tissue types. The algorithms first detect brain edges and then use mathematical models to remove exterior tissues, resulting in a clean image of the brain for subsequent processing.
Once the non-brain tissues are removed, the next step is selecting the proper information channels. MRI images possess 256 channels with different slices of the brain displayed. There is little or no useful information in most channels, particularly the first and last ones, and they are usually simply black. For purposes of simplifying and making it efficient, only the channels containing significant and useful information should be retained. Through pixel intensity analysis and content inspection in each channel, 112 channels of information are selected and the remaining ones are discarded.
In aggregate, the preprocessing stage guarantees that the resulting images are of relevance, consistency, and high quality. Such datasets cleaned up as such are now prepared for subsequent stages such as feature extraction and segmentation with strong preference for retaining the most informative and critical material. This stage is extremely critical for the proper diagnosis of autism because it guarantees that only high-quality and meaningful data are forwarded for subsequent processing.
Feature Extraction Based on Clustering and the VGG16 Neural Network
Clustering based feature value extraction is identified to be a very important stage in processing image information as the main aim of which is to distinguish and remove significant features on the pictures by classifying the similar points into a cluster. It is also optimal in uncovering intricate patterns especially identifying places in the brain that contribute to ailment like autism during MRI. In this section the clustering algorithms, FCM and GMM, are used to extract brain image features. The extraction of the features based on clustering is explained as follows:
To begin with, when preparing data, a channel in two-dimensional MRI image is turned into a one-dimensional vector. This step makes the data straight forward and is prepared to be formed together. In this case not only is a two-dimensional (under normal conditions of size MxN that is, the number of pixels in the horizontal direction equal to the number of pixels in the vertical direction) image transformed into a one-dimensional (under normal conditions) pixel-intensity information (MxN) vector, but also the pixel intensity data is stored in the form of a one-dimensional vector instead of a two-dimensional array.
Once the pictures are transformed to a one-dimensional having vectors, the vectors are placed into a matrix of every patient. That is, given a MRI image of 112 channels, a MN× matrix is obtained. In the matrix, every column is associated with a one-dimensional vector on a single channel, whereas each row is related to the feature of the pixel on all the channels.
The second step is the application of the clustering algorithm e.g. FCM and GMM to divide the images and detect patterns of the pixels in various clusters.
- Clustering Using FCM
FCM algorithm is considered to be one of the most popular clustering algorithms. Unlike hard clustering algorithms such as K-means, FCM assigns relative membership of every point to every cluster. A pixel can be a member of more than one cluster simultaneously, a feature that is very desirable with images containing fuzzy data or boundary regions.
During the stages of executing FCM, the first step involves analyzing the image as well as the specific issue and then deciding the number of clusters (C). Cluster centers can be initiated randomly or can be done through methods like K-means. Then comes the step where each pixel’s membership in each cluster is assessed. Membership indicates the extent of belonging to different clusters, and is computed using a function describing the distance from the cluster centroids.
The calculation of the membership uij of pixel i in cluster j is based on the 3.1 equation:
|
|
(3.1) |
Here:
represents the membership of pixel i in cluster j,
is the distance of pixel i from the center of cluster j,
m is the fuzziness parameter, which ranges between 1.5 and 2,
and C is the number of clusters.
After the membership values were evaluated, the cluster center was updated using Equation (3-2):
|
|
(3.2) |
This process is repeated until the cluster centers reach a minimum and the algorithm converges.
• Clustering Using Gaussian Mixture Model (GMM)
The GMM algorithm is an unsupervised statistical model that represents the data as a mixture of several Gaussian distributions. In this method, the number of clusters KKK is initially determined, which is usually determined from the basis of parameters such as AIC or BIC. Once the number of clusters is established, the weights of the Gaussian distributions (weight, covariance, and mean of each cluster) are initialized randomly or using techniques such as K-means. The Expectation (E) step is then performed, where the probability that each pixel belongs to each cluster is computed. The probabilities are computed using the Gaussian distributions for each cluster. Secondly, in the Maximization (M) step, the parameters of the Gaussian model are updated optimally to fit the data best. Both these steps, i.e., the Expectation (E) and Maximization (M), are iteratively repeated until convergence of the algorithm is achieved and the model parameter changes become minimal. The probability of a pixel i being included in cluster k in the GMM model is expressed as Equation 3.3:
|
|
(3.3) |
Finally, the probability distribution obtained in the Expectation step is used to update the model parameters in the Maximization step.
• Combining the Clustering Results of FCM and GMM Algorithms
Merging the use of FCM and GMM algorithms together provides plenty of advantages which this thesis discusses. The FCM algorithm finds overlapping clusters and the GMM algorithm builds complex probabilistic structures. Together as complementary algorithms, they improve the understanding of the autism spectrum disorders brain pertaining to the illness. For example, the GMM also helps to locate brain areas of complex structures, of which excels at predicting and FCM overlaps with those structures or regions of uncertainty and diagnoses precisely. After both algorithms are implemented, the results of the clustering are transformed into images. As explained in the last paragraph, these images are combined with the original raw images to create a multi-channel image used for feature extraction and autism detection. The two are merged through an averaging method which incorporates the pixel membership grades from the FCM and the pixel cluster probabilities from the GMM.After the fusion, the raw images and the clustered images are used to create multi-channel images.This new multi-channel image, composed of both raw and clustered images, is used in deep learning models to improve and more precisely analyze the features of the extracted brains. The multi-channel approach allows deep learning models to improve brain feature processing and detection accuracy. Thus, clustering-based feature extraction based on FCM and GMM is significant in enhancing the accuracy and dependability of systems for identifying brain disorders such as autism. With these methods, classification of different brain regions that are associated with various disorders is possible, and the performance of deep learning–based systems is increased in this field.
• Feature Extraction from Multi-Channel Images Using the VGG16 Neural Network
in case multi-channel images are acquired at this stage, the images will be fed to a VGG-16 neural network to extract their features. VGG-16: It is a pretrained convolution neural network, which has been trained on over a million images in the ImageNet database to recognise images. The neural network architecture consists of 16 layers, 13 convolution layers and the remaining fully connected layers. VGG-16 has the task of learning deep and complex patterns of brain MRI scans.
In order to ensure the best performance of VGG-16 in relation to detecting autism using brain MRI scans the network requires to be fine-tuned. Fine tuning is a method where the weights of the network layers are changed so as to make the network learn to pick salient features of the new set.
At this stage, instead of using pre-trained weights originally trained for overall image classification, the weights are fine-tuned to be more suitable for brain MRI data and autism detection tasks.
Next, features are extracted exclusively from the VGG-16 model's convolutional layers. The convolutional layers extract spatial and structural information from the images efficiently, which is essential in the identification of brain regions associated with autism disorders. They are able to detect complex features like edges, texture, and specific brain structures, which are extremely important for the identification of autism-related brain abnormalities.
Since the fully connected layers are primarily used for processing and classification at the final stage, they are not feature-effective at extracting features from brain images and therefore are removed here. By removing the fully connected layers, the network focuses more on structural and spatial feature extraction from the convolutional layers, reducing the network's computational complexity. Therefore, such consequential features received from the convolutional layers are utilized in the subsequent stages, and pertinent and true information for finding brain areas associated with autism is obtained.
This approach ensures that the model employs meaningful and correct features to detect autism, thereby improving the model's accuracy and performance in identifying brain abnormalities for this disorder.
Classification Using the ResNet Neural Network Optimized with the Dwarf Mongoose Algorithm
In the last stage of our proposed approach, we do the labeling for extracted features by ResNet-50 with Dwarf Mongoose Optimization Algorithm (DMOA). The architecture is a 50-layer convolutional neural network with multiple residual blocks. The greatest advantage of ResNet lies in having the residual connections, which enables a better training of the network by overcoming this vanishing gradient problem. The structure of the network is formed by convolutional layers which are followed by batch normalization and pooling layers combined with fully connected layer for the classification. The most striking characteristic of ResNet is the appearance of residual connections, through which network can skip layers and support back-propagate gradients. This ensures that the network is able to learn deep networks while maintaining performance.
For hyperparameters, learning rate and batch size are two significant parameters that need to be adjusted in order to improve model performance. The learning rate specifies how much the weights of the model are adjusted at each training step. A large learning rate facilitates rapid model convergence but risks overshooting the solution, whereas a small learning rate brings slow model convergence but with a stable training process. Batch size determines the number of samples used to compute each weight update of the model. Using a small batch size can produce noisy gradients, while using a large batch size can stabilize training but can become memory intensive. To help find the optimal values for learning rate and batch size, the Dwarf Mongoose Optimization Algorithm is used. This nature-inspired algorithm can find the optimal pair of these hyperparameters with great efficiency by imitating the behavior of dwarf mongooses in the wild. Here, the tuning parameters are the learning rate and batch size while various tuning parameters are used for optimization. The objective function is the classification error of the ResNet-50 model.
The cost function is defined based on the equation 3.4:
|
|
(3.4) |
The Dwarf Mongoose Algorithm progressively fine-tunes the learning rate and batch size to determine the best pair to reduce the error within the set stopping criterion, which defaults to the maximum number of iterations. This ensures that within available computational time, the optimization routine will converge to a solution. Upon completion of the optimization routine, the trained network evaluates the performance of the model on the optimized hyperparameters, focusing on determining the accuracy and precision. This assists in gauging the effectiveness of the trained ResNet-50 model in detecting features of autism in brain MRIs and the model's performance on unseen instances of data. The main goal of this is to ensure that the model detects autism as accurately as possible while maintaining optimal performance.
Results and Discussion
Brain MRI scans have the potential to reveal the very first signs of autism. An early comprehension of this intricate neurodevelopmental disorder is paramount. The present research puts forward a hybrid deep learning framework that is a combination of advanced image preprocessing, an optimized VGG-16 for feature extraction, and an optimized ResNet-Dwarf Mongoose for classification. The proposed method contains the usage of multi-channel inputs along with both raw and segmented images which helps to expand the learning of the network regarding important features. The DMOA technique can be applied to the ResNet classifier for modification and optimization, and this will consequently improve its performance and accuracy leading the way to appropriate generalization of the classifier.
This section provide the result of the proposed technique, this on how good is it when use in recognizing the autism and wether better then what actually exist.
The specifications of the computer used in this study are presented in table 1.
Table 1: Specifications of the system used
| Operating System | Programming Language | RAM | Processor |
| Windows 11 Pro 64-bit | MATLAB 2024a | 64GB | Core i7 |
Dataset
ABIDE I dataset includes 1,112 MRI scans captured across 17 global sites. The data include structural MRI and resting-state fMRI for research on Autism Spectrum Disorder. Of these, 539 individuals have autism, and 573 are the healthy control group. Data were collected from participants who are 7 to 64 years old, and guidelines for data quality and confidentiality were followed per HIPAA standards. This database is a valuable wealth for the study of brain patterns in autism and was used in the present research to build the hybrid deep model.
Evaluation Metrics
To determine how well the suggested method classifies Autism Spectrum Disorder, it is crucial to apply some common classification metrics, which are composed of accuracy, precision, recall, and F1-score. This gives a comprehensive view of the classifier's capability to tell apart the two categories, healthy and autism, in the case of the model. The section below presents the relevant definitions of these measures.
• Accuracy: Accuracy measures the overall correctness of the classification and is evaluated as the ratio of correctly classified samples to the total number of samples. Equation 4.1 shows the calculation of this evaluation criterion.
• Precision: Precision calculates the ratio of correctly identified autism samples to the total number of samples predicted as autism. Equation 4.2 shows the calculation of this evaluation criterion.
(4.2)
• Recall: Recall measures the model's ability to correctly identify all autism samples. Equation 4.3 shows the calculation of this evaluation criterion.
(4.3)
• F1 Score: The F1 score is the harmonic mean of precision and recall, providing a balanced metric that accounts for both false positives and false negatives. Equation 4.4 shows the calculation of this evaluation criterion.
(4.4)
Analysis and Review of Results
In this section, the method implementation and the results at each step are presented. For the proposed method, the first step involves preprocessing the MRI images. For high-quality and homogeneous MRI images—which is needed for competent and dependable analysis—the absolute need for preprocessing is unquestionable. Preprocessed MRI images from the ABIDE I dataset undergo a sequence of preprocessing. First step is the normalization of image intensity values to standardize brightness and contrast levels across the whole dataset. Noise and artifacts are reduced by advanced filtering techniques, improving image quality. In addition, applying skull-stripping methods removes non-brain tissues, practically isolating brain regions in isolation for subsequent analysis.
Moreover, during these phases, the data dimensions are reduced by keeping only the informative channels from the original 256 channels. As observed, the first and last channel, which correspond to the top and bottom slices of the brain, respectively, are black and contain no informative content. From keen observation of the dataset, 112 meaningful information channels were discovered and kept while the remaining channels were left behind. This channel selection properly disregards much of the computational overhead at the expense of data integrity. Illustrations of randomly selected channels from preprocessed MRI scans of autistic and non-autistic samples are shown in Figures 2 and 3. These figures show how effective the preprocessing steps are in filtering the data to be prepared for future analysis.
Figure 2: Three randomly selected channels from a randomly chosen autistic sample from the processed dataset
Figure 3: Three randomly selected channels from a randomly chosen non- autistic sample from the processed dataset
Next, clustering-based extracted features are used to segment the MRI images. The size of the original image is 176×256×256, where the spatial size is 176×256 and the number of channels is 256. Because clustering is carried out along the channel axis, the spatial direction must be flattened first. For this purpose, each two-dimensional channel image is initially compressed into a vector with size 45056×1. These vectors are further arranged into 45056×112-sized matrices for each subject, which are the input data expressed in terms of 112 features. These matrices form the basis of GMM and FCM-based clustering.
The brain consists of several distinct regions, such as gray matter, white matter, and cerebrospinal fluid, each with different intensity distributions. Furthermore, neurological conditions such as autism can introduce additional heterogeneity. Meanwhile, subcortical structures (e.g., thalamus and hippocampus) require precise segmentation for accurate analysis of neural conditions. Additionally, MRI images include other regions such as black backgrounds, which contribute to the segmentation task. Therefore, clustering of brain MRI images requires a relatively large number of clusters due to the complexity of brain structures and the need for precise differentiation (in the proposed method, different clusters represent various parts of the MRI images). On the other hand, increasing the number of clusters may lead to covariance matrix multicollinearity issues in the GMM algorithm, as covariance matrices in Gaussian mixtures may become singular or nearly singular. Hence, after several trials, the number of clusters for clustering (in both GMM and FCM methods) was set to 20 to balance the complexity of brain MRI images and the covariance issue in GMM.
The clustering process assigns each pixel to a specific cluster based on its values across different channels, effectively performing unsupervised segmentation of the MRI images. This step captures unique patterns in the brain structure by grouping pixels with similar intensities across channels. After segmentation is completed, the clustered outputs are reconstructed into two-dimensional images. Then, the segmented outputs and the original 112-channel MRI images are resized to 224×224 pixels.
To reduce these images to 224×224, their height is increased and width reduced both through bicubic interpolation. Since the image size isn't upscaled extensively, this interpolation introduces no distortion or noise into images. The reduced images are then concatenated to create a three-dimensional representation of a feature map of 224×224×114 per sample. This feature set rich with raw and segmented data facilitates ensuing processing. Figures 4 and 5 illustrate the results of GMM and FCM algorithm clustering for random sets of autistic and non-autistic MRI images, comparing clustering performance visually.
Figure 4: GMM-and FCM- based feature from a randomly selected autistic sample from the dataset
Figure 4: GMM-and FCM- based feature from a randomly selected autistic sample from the dataset
Figure 5: GMM- and FCM- based feature from a randomly selected non- autistic sample from the dataset
Figure 5: GMM- and FCM- based feature from a randomly selected non- autistic sample from the dataset
The second step in the proposed method is employing the VGG16 network that has been widely recognized with great capability to extract information-rich features from complex image datasets. In its autism detection modification, the original classification layers of this network, including the fully connected layer with 1000 neurons and the SoftMax layer that was meant for COCO dataset classification, were removed. These three layers were substituted by three new layers: a fully connected layer with nF neurons for feature extraction, another fully connected layer with two neurons, and a SoftMax layer for binary classification. The middle feature layer is employed to extract expert features that are designed to be best suited for autism detection. The fine-tuned VGG16 network was fine-tuned using a small learning rate of 0.0001 for 100 training epochs. These conservative settings ensure weight adjustment in the pre-trained network is negligible and re-tuning it to detect features more topically associated with autism detection.
Following refinement, the extracted features were obtained by passing the input data into the network. The input images, initially of dimensions 224×224×114, were preprocessed to be compatible with the anticipated VGG16 input size of 224×224×3. To this end, the 114 channels were divided into three groups: the first 112 channels were made up of preprocessed MRI images, and the other two channels contained the GMM and FCM segmentation results. These channels were projected onto 112 224×224×3 sized images, of which the first channel of each image represented a particular processed MRI slice and the remaining two channels represented the fixed segmentation outputs. These images were passed through the modified network to provide 112 feature vectors of size nF. The nF value was set at 1344 for reshaping the 112 feature vectors to the final 224×224×3 size. This reconstructed feature representation preserved important spatial and contextual information for classification in the next stage via the ResNet architecture. The VGG16 architecture transformed is illustrated in Figure 6, with the changes made and feature extraction pipeline being emphasized.
Figure 6: Architecture of the modified VGG16 network
Figure 6: Architecture of the modified VGG16 network
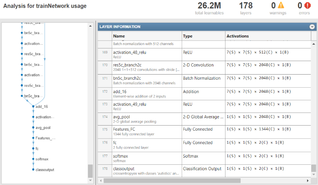
The final stage in the proposed method is dedicated to classifying MRI images using the ResNet architecture. ResNet, also known as a residual network, is a highly effective convolutional neural network structure that allows the design of deep architectures without facing the vanishing gradient problem. Instead of training the ResNet architecture from scratch, a pre-trained model was utilized named into part and retrained to serve as a ASD detection task. There are a number of benefits to using pre-trained networks such as the reduction in computational requirements, faster training times and using features that have been learned on large scale datasets. The redesign of ResNet was conducted in the same way as described for the VGG16 network.
ResNet’s original classification layers are discarded, and new fully connected layers for binary classification were added. The new layers are feature extraction, fully connected (two neurons), and SoftMax as can be seen in Figure 2 the classes are two only for binary classification. The resulting network architecture comprises 26.2 million learnable parameters, which are finely tuned toward the autism detection problem. Optimizing parameters including learning rate and batch normalization was required for ResNet pretraining. To find the values of these parameters, instead of using an old-fashioned trial-and-error method, we utilised a newly finding algorithm called the Dwarf Mongoose Optimization Algorithm. This feature-wise optimization process led to a successful and efficient fine-tuning of the ResNet, improving its generalization to new data. By taking advantage of the potent pre-trained ResNet architecture and its parameters fine-tuned by the Dwarf Mongoose Optimization Algorithm, the classification network has successfully realized a high-accuracy and reliable autism diagnosis figure 7.
Figure 7: Modified ResNet50 Architecture
The ResNet parameters were tuned out using Dwarf Mongoose Optimization Algorithm. The main parameters of the algorithm, population size, maximum number of iterations and upper and lower limits for decision variables are wisely set to balance computational burden with search space exploration. Both the size of population and rounds determine a trade-off between computational burden and the number of possible solutions that can be evaluated. For fine-tuning the network and computing results for each assessment (both being computationally intensive) we set these numbers to 4 and 8 respectively. This arrangement allowed fast execution combined with comprehensive coverage of the parameter space.
The bounds of the decision variables were established as to allow a complete search on the parameter values. The batch size ranges from 10 and 50 and the learning rate was set as 1×10-5 to 0.001. These limitations permitted the Dwarf Mongoose Optimization to investigate a wide space of possible solutions and at the same time optimize network performance. Furthermore, the number of training epochs for the CNN training under this optimization stage was 40. The number was relatively small to minimize computation load while guarantee an active exploration of candidate solutions.
Afterwards, the network was trained for the last time using all optimized parts to reach its optimum in accuracy as much as possible. Figure 8 presents the convergence curve of the Dwarf Mongoose Optimization, and clearly demonstrates an incremental value function. The best parameter values determined by this process are displayed in Table 2 Table. Employing this method largely brought the computational burden down while having no adverse impact on the final performance.
Figure 8: Convergence curve of RestNet50 parameter tuning for autism detection using the Dwarf Mongoose Optimization Algorithm
Table 2: Optimal parameter values obtained using the Dwarf Mongoose optimization algorithm
| Batch Size | Learning Rate | Hyperparameters |
| 38 | 0.0008 | Value |
the optimal hyperparameters using the DMO algorithm, the final fine-tuning of the designed ResNet was performed. For this stage, the number of epochs was set to 100, and the ADAM algorithm, known for its robust optimization capabilities, was employed as the training algorithm. These choices ensured that the network could effectively learn from the data while maintaining stability and efficiency during the training process. It should be noted that the dataset is divided into three categories of training, testing, and validation subsets randomly, with proportions of 70%, 20%, and 10%, respectively. Hence, 778 samples are dedicated to the training subset, 111 are kept for validation, and the remaining 223 ones are employed for assessing the results of the trained network. This split was selected to ensure a reliable and fair evaluation: the 70% training subset provides enough data for learning robust features, the 10% validation subset is used to monitor overfitting and guide model tuning during training, and the 20% test subset is kept completely unseen to report the final performance objectively.
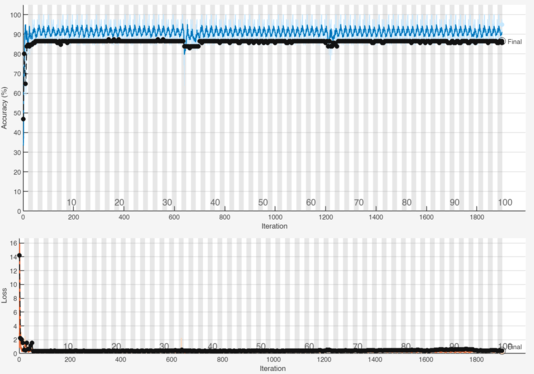
Figure 9 shows the training of the fine-tuned network. The results in this figure demonstrated that parameter tuning dramatically improved the performance of the model led to faster convergence and better accuracy. The optimal structure of the network promotes better generalization and robustness in autism detection. Later on in the further part of this chapter, the proposed approach is quantitatively as well as qualitatively assessed with metrics demonstrating a level of accuracy, robustness and effectiveness towards autism detection using MRI brain images.
Figure 9:Convergence curve of the fine-tuned ResNet50 for autism detection
Evaluation of Test and Training Sets Based on the Confusion Matrix
The confusion matrix is a good measure of classification models skill. It is shown in tabular form and it can give very detailed information on how the model discriminates between the different classes. In the present two-class (i.e., autism vs normal) study, confusion matrix contains 4 entries namely TP (True Positive), TN (True Negative), FP (False Positive) and FN (False Negative). TP denotes cases being correctly classified as autism, and TN denotes cases that are correctly recognized as normal. In contrast, FP) indicates normal cases misclassified as autism and FN is vice versa.
The metrics such as accuracy, precision, recall and F1-score mentioned above can be measured using this matrix. It should be noted that the confusion matrix is computed on the test set only, which contains 223 samples in total (102 autistic and 121 non-autistic). To give more details about classification performance, we also present in Figure 10 the confusion matrix of evaluating our method. This is a plot showing how each class relates to being predicted correctly/incorrectly and informs us directly about the capabilities of the model. As can be seen, all non-autistic cases are classified correctly. Moreover, 88 out of 102 cases of autistic samples are classified correctly, and just 14 autistic cases are mistakenly classified as non-autistic. The results from the confusion matrix indicate that the proposed method performs exceptionally well, achieving high accuracy and appropriate classification for both autism and normal cases.
Figure 10: Confusion matrix of the proposed model for the test dataset
Finally, the confusion matrix emphasizes the effectiveness of the proposed hybrid deep learning approach. By employing advanced preprocessing, feature extraction, and an optimized ResNet classifier, our method demonstrates superior classification performance compared to existing solutions. These results highlight the robustness and reliability of the model in accurately detecting autism from brain MRI images.
Evaluation of Training and Test Set Results Based on the Receiver Operating Characteristic Curve
The Receiver Operating Characteristic (ROC) curve is a tool that helps to visualize how well a binary classification system is working by varying its thresholds. The curve is plotted by using the True Positive Rate (sensitivity) and False Positive Rate. The true positive rate is a measure of the number of positive cases (such as autism) that are correctly identified in comparison to the total of positives. The False Positive Rate is the percentage of the normal instances that were incorrectly classified as positive. The ROC curve depicts the trade-off between sensitivity and specificity and it is one of the primary performance evaluation measures for binary classification models.
The Area Under the Curve (AUC) represented by the ROC curve can be observed on Figure 11, it is an important measure that can be extracted from the ROC curve and represents the degree of separation made between the two classes by the model. An AUC value of 1 represents ideal discriminative power and an AUC of 0 is equivalent to the performance that can be expected by random guessing. The ROC curve of the test set Figure 11 indicate that the proposed method can very well discriminate children with Autism and their typical peers since the Area Under the Curve (AUC) values are very close to optimal.
As shown in Figure 11, the ROC value for both classes (autism and non-autism) is equal to 0.9276.
Figure 11: ROC curve of the proposed model for the test dataset
Evaluation of the Test Dataset Based on the Introduced Assessment Metrics
In this part, performance of the proposed algorithm in autism detection is analyzed using few evaluation measures such as accuracy, precision, recall, and F1 score. To present a clear picture of the results, Figure 12 provides a summary of these metrics in the form of a bar chart. This visualization gives an idea of the worth of each metric and of the ease of understanding how the proposed model is doing on unseen data.
As we can see in Figure 12 that the accuracy is 93.72%, Precision is 94.81%, Recall is 93.14% and F1-Score is 93.97%. These high values demonstrate that our model has a good performance and represent the robustness and reliability of the proposed deep learning hybrid approach in identifying autism in unseen data.
Figure 12: Overall evaluation metrics of the the proposed hybrid model
Comparison of the Proposed Method with Previous Studies
In this section, performance of proposed hybrid deep learning approach is compared with two methods in the literature.
The first one, presented in [20], consisted of a series of CNN-based models for extracting features from neural images based on the ABIDE dataset. In this study, a neural network called ResNet-50 was used to classify neural patterns between ASD & TD group. The attained performance was an accuracy 87.0%. The primary goal of this evaluation was to have an objective tool for diagnosing autism that could supplement the conventional diagnostic methods that are based on behavioral assessments.
The 2nd approach suggested in [21], discussed convolutional and transfer learning techniques to process resting state fMRI (rs-fMRI) data that consists of 2000+ resting state fMRI data collected from patients with autism and normal, derived from the ABIDE I. The four-dimensional nature of rs-fMRI data (three spatial dimensions and the temporal dimension) makes it a promising procedure for the creation of diagnostic biologic markers of brain damage. The performance of the best CNN model was better than traditional ones that were previously reported on the ABIDE I dataset (81.56% accuracy in classification of autism). This research demonstrates that AI, IoT technologies and medical image analysis integration plays a role in autism detection and support improvement. In the results, Table 3, we will compare the accuracy of proposed method in this study with another research.
Table 3: Comparison of the proposed method with other studies
| References | Method | Dataset | Accuracy | ||
| Husna, R. N. S., et al | ResNet-50 | Autism Brain Imaging Data Exchange (ABIDE) | 87% | ||
| Ashraf, A., et al | CNN and transfer learning algorithms | Autism Brain Imaging Data Exchange (ABIDE) | 81.56% | ||
| Lamani, M. R., & Julian Benadit, P | PDNN | Autism Brain Imaging Data Exchange (ABIDE) | 90% | ||
| Lamani, M. R., Benadit, P. J., & Vaithinathan, K | CNN + Functional Connectivity Features | Autism Brain Imaging Data Exchange (ABIDE) | 92.2% | ||
| Singh, U., Shukla, S., & Gore, M. M |
|
Autism Brain Imaging Data Exchange (ABIDE) | 92.80% | ||
| Proposed method | FCM+GMM+ VGGnet+Resnet+DMO | Autism Brain Imaging Data Exchange (ABIDE) | 93.72% |
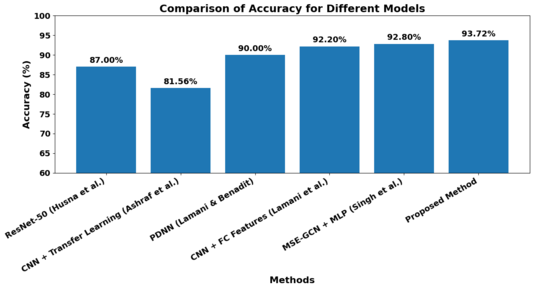
In the following, Figure 13 visualizes the data from Table 3 to provide a clearer understanding for the reader, illustrating the comparison of accuracy across different models.
As can be seen in Table 3 & figure 13, our approach combining sophisticated pre-processing, multi-channel image inputs and ResNet classifier fine-tuned using Dwarf Mongoose Optimization algorithm attained test accuracy of 93.72%. This progress demonstrates the power of our integrated strategy in optimizing between automation, efficiency and classification performance, clearly offering a more scalable solution for autism detection.
Conclusion
Autism Spectrum Disorder (ASD) is a heterogeneous neurodevelopment disorder, has profound consequences and review its colonization requires a diagnosis that should be performed early and effectively in order to allow the intervention. Conventional diagnosis depends on behavioral evaluations and this fact is intrinsically linked to the late diagnosis. Recent advances in neuroimaging, especially brain MRI, now bring with them the promise of objective diagnosis which would enable us to measure both the structures and functions of the brain related to autism. However, current computer-based methods for MRI analysis may fare poorly in high-dimensional data or lack reliable feature extraction and generalization.
To overcome these above mentioned limitation, a novel hybrid deep learning model in terms of improved pre-processing, multi-channel image input and ResNet classifier by employing DMA is presented in this thesis. The amalgamation provides improved accuracy and determinism in an autism detection scenario. The proposed method achieves better results on ABIDE I dataset. Upon evaluation, the model obtained 93.72% of accuracy, 94.81% of precision, 93.14% of recall and an F1-Score as high as 93.97%, which demonstrates its strong capability with respect to learning and classifying complex patterns. This outcome illustrates the efficiency of the proposed solution to mitigate drawbacks such as overfitting, and lack of prediction performance, that are major disadvantages of existing approaches.
The new models simply involves a number of useful features. Indeed, the combination of clustering-based segmentation methods (e.g. FCM and GMM) with full-trained deep learning framework is a true representation of brain structure. Multi-channel images increase the heterogeneity of features and hyperparameter fine-tune in the ResNet feature classifier using the Dwarf Mongoose Optimization Algorithm aids metrics performance. In addition, this method makes hardly no use of manual feature engineering and scales well in the context of high dimensional MRI data providing a good and scalable solution for autism detection. The presented thesis not only represents a major improvement of computational tools for autism diagnosis but also paves the way to new research and, ultimately, clinical applications.
Future Research Directions
Although the hybrid deep learning method we proposed for detecting autism from MRI images is indeed an interesting approach, it remains a proof of concept at this stage and needs further development and extension by others researchers before it will be improved further. One possible trajectory of research is examining other deep learning models like transformers or vision-based generative models that may result in higher accuracy of feature extraction and classification performance. Adding temporal information from fMRI or longitudinal designs would then lead to more subtle developmental trends of autism being detected, thus, making the diagnostic system dynamic and broad in terms of scope.
Transfer learning based on pre-trained models for medical imaging tasks is another interesting approach. The fine-tuning of the pre-trained models on autism classification can contribute to the improvement of generalization and performance, especially with small training sets. In a broader sense, multimodal data, for example, MRI-imaging and genetic or EEG records, may assist in providing a more accurate explanation and prediction of Autism Spectrum Disorder.
On the contrary, ethical issues and enhancement of model interpretability should be ranked among the topmost priorities in future research. The methods proposed can also be applied and expanded as global methods in further studies with different datasets of age groups and demographic attributes. Besides, the technologies will not only enhance the diagnosis of autism but also have a significant impact on the area of AI-based medical imaging in general.
References:
Nogay, H. S., & Adeli, H. (2023). Diagnostic of autism spectrum disorder based on structural brain MRI images using, grid search optimization, and convolutional neural networks. Biomedical Signal Processing and Control, 79, 104234
Prasad, V., Sriramakrishnan, G. V., & Diana Jeba Jingle, I. (2023). Autism spectrum disorder detection using brain MRI image enabled deep learning with hybrid sewing training optimization. Signal, Image and Video Processing, 17(8), 4001-4008.
Tang, L., Mostafa, S., Liao, B., & Wu, F. X. (2019). A network clustering based feature selection strategy for classifying autism spectrum disorder. BMC medical genomics, 12, 1-10.
Baadel, S. (2019). A machine learning clustering technique for autism screening and other Applications (Doctoral dissertation, University of Huddersfield).
Alves, C. L., Toutain, T. G. D. O., de Carvalho Aguiar, P., Pineda, A. M., Roster, K., Thielemann, C., ... & Rodrigues, F. A. (2023). Diagnosis of autism spectrum disorder based on functional brain networks and machine learning. Scientific reports, 13(1), 8072.
Ahmed, M. R., Ahammed, M. S., Niu, S., & Zhang, Y. (2020, March). Deep learning approached features for ASD classification using SVM. In 2020 IEEE international conference on artificial intelligence and information systems (ICAIIS) (pp. 287-290). IEEE.
Ali, M. T., ElNakieb, Y., Elnakib, A., Shalaby, A., Mahmoud, A., Ghazal, M., ... & El-Baz, A. (2022). The role of structure MRI in diagnosing autism. Diagnostics, 12(1), 165.
Moridian, P., Ghassemi, N., Jafari, M., Salloum-Asfar, S., Sadeghi, D., Khodatars, M., ... & Acharya, U. R. (2022). Automatic autism spectrum disorder detection using artificial intelligence methods with MRI neuroimaging: A review. Frontiers in Molecular Neuroscience, 15, 999605.
Katuwal, G. J. (2017). Machine learning based autism detection using brain imaging. Rochester Institute of Technology.
Bilgen, I., Guvercin, G., & Rekik, I. (2020). Machine learning methods for brain network classification: application to autism diagnosis using cortical morphological networks. Journal of neuroscience methods, 343, 108799.
Saputra, D. C. E., Maulana, Y., Win, T. A., Phann, R., & Caesarendra, W. (2023). Implementation of Machine Learning and Deep Learning Models Based on Structural MRI for Identification Autism Spectrum Disorder. J. Ilm. Tek. Elektro Komput. Dan Inform. JITEKI, 9, 307-318.
Guo, X., Wang, J., Wang, X., Liu, W., Yu, H., Xu, L., ... & Chen, Y. (2022). Diagnosing autism spectrum disorder in children using conventional MRI and apparent diffusion coefficient based deep learning algorithms. European Radiology, 1-10.
Devika, K., Mahapatra, D., Subramanian, R., & Oruganti, V. R. M. (2022). Outlier-based autism detection using longitudinal structural MRI. IEEE Access, 10, 27794-27808.
Duan, Y., Zhao, W., Luo, C., Liu, X., Jiang, H., Tang, Y., ... & Yao, D. (2022). Identifying and predicting autism spectrum disorder based on multi-site structural MRI with machine learning. Frontiers in human neuroscience, 15, 765517.
Eslami, T., Raiker, J. S., & Saeed, F. (2021). Explainable and scalable machine learning algorithms for detection of autism spectrum disorder using fMRI data. In Neural engineering techniques for autism spectrum disorder (pp. 39-54). Academic Press.
Mahadevaswamy, U. B., Ravikumar, R., Mahadev, R., Rao, K. V., & Anurag, K. S. (2021). Autism detection using machine learning. International Journal of Bioinformatics Research and Applications, 17(4), 375-387.
Karampasi, A., Kakkos, I., Miloulis, S. T., Zorzos, I., Dimitrakopoulos, G. N., Gkiatis, K., ... & Matsopoulos, G. (2020, December). A machine learning fMRI approach in the diagnosis of autism. In 2020 IEEE international conference on big data (big data) (pp. 3628-3631). IEEE.
Sharif, H., & Khan, R. A. (2022). A novel machine learning based framework for detection of autism spectrum disorder (ASD). Applied Artificial Intelligence, 36(1), 2004655
Husna, R. N. S., Syafeeza, A. R., Hamid, N. A., Wong, Y. C., & Raihan, R. A. (2021). Functional magnetic resonance imaging for autism spectrum disorder detection using deep learning. Jurnal Teknologi, 83(3), 45-52.
Ashraf, A., Qingjie, Z., Bangyal, W. H. K., & Iqbal, M. (2023). Analysis of brain imaging data for the detection of early age autism spectrum disorder using transfer learning approaches for internet of things. IEEE Transactions on Consumer Electronics, 70(1), 4478-4489.
Document information
Published on 20/02/26
Licence: CC BY-NC-SA license
Share this document
Keywords
claim authorship
Are you one of the authors of this document?